Search Count: 131
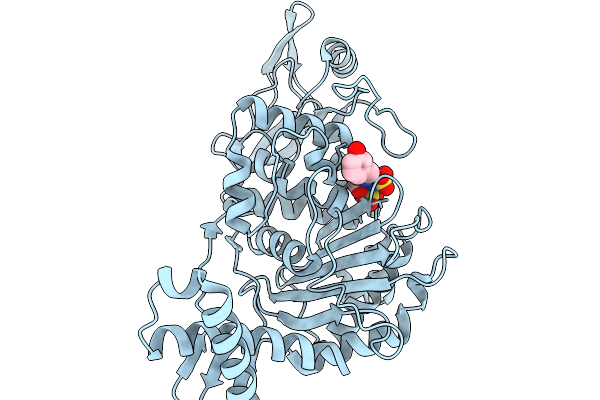 |
Organism: Streptococcus pneumoniae
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: A1JNW, SO3, CL |
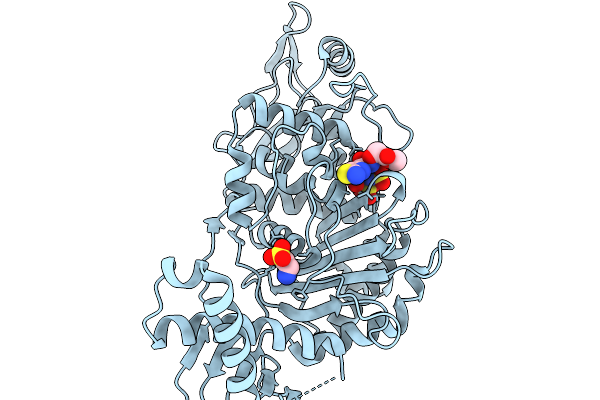 |
Organism: Streptococcus pneumoniae
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: A1JNT, SO3, CL, TAU |
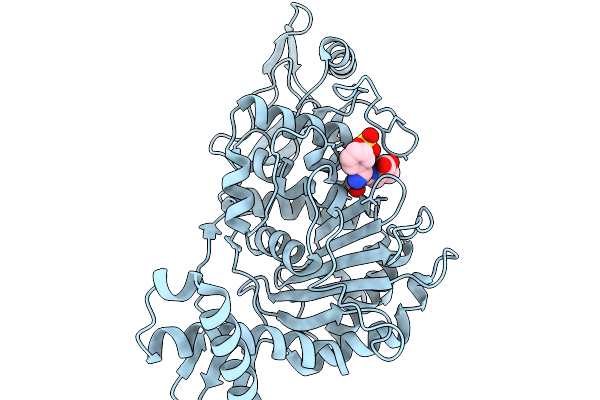 |
Penicillin-Binding Protein 1B (Pbp-1B) In Complex With Cephalexin - Streptococcus Pneumoniae R6
Organism: Streptococcus pneumoniae r6
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2026-02-18 Classification: TRANSFERASE Ligands: A1JXD, CL, SO3 |
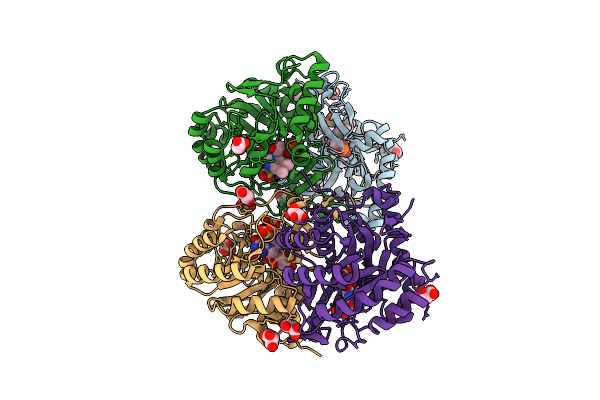 |
Crystal Structure Of The C131A Mutant Of Trypanosoma Brucei Dhodh In Fmn-Reduced, Ligand-Free Form
Organism: Trypanosoma brucei brucei treu927
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-11-26 Classification: FLAVOPROTEIN Ligands: FNR, MLI, SO3 |
 |
Crystal Structure Of Wild-Type Trypanosoma Brucei Dhodh In Fmn-Reduced, Ligand-Free Form
Organism: Trypanosoma brucei brucei treu927
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-11-26 Classification: FLAVOPROTEIN Ligands: FNR, MLI, SO2, SO3 |
 |
Organism: Pyrococcus horikoshii ot3
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2025-03-12 Classification: TRANSFERASE Ligands: SO4, SO3, EDO, PEG |
 |
Crystal Structure Of Nep1 In Complex With Adenosine From Pyrococcus Horikoshii Ot3
Organism: Pyrococcus horikoshii ot3
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2025-03-12 Classification: TRANSFERASE Ligands: CL, ADN, SO4, SO3, EDO, PEG |
 |
Structure Of Mycobacterium Thermoresistibile Nrdi(Reduced) Determined At 1.1 Angstrom Resolution
Organism: Mycolicibacterium thermoresistibile atcc 19527
Method: X-RAY DIFFRACTION Resolution:1.21 Å Release Date: 2024-10-09 Classification: FLAVOPROTEIN Ligands: FMN, SO3, PEO, PER, CL |
 |
Organism: Pectobacterium carotovorum
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2024-09-04 Classification: HYDROLASE Ligands: FES, NA, MG, SO3, GOL, SO4, CL |
 |
Crystal Structure Of Dimt1 From The Thermophilic Archaeon, Pyrococcus Horikoshii
Organism: Pyrococcus horikoshii ot3
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2024-08-28 Classification: TRANSFERASE Ligands: ZN, SO3, GOL, PEG, EDO, ACT, ARG, EPE |
 |
Crystal Structure Of Dimt1 In Complex With 5'-Methylthioadenosine And Adenosine From Pyrococcus Horikoshii
Organism: Pyrococcus horikoshii ot3
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2024-08-28 Classification: TRANSFERASE Ligands: MTA, ADN, ZN, CL, SO3, GOL, EDO, ACT, ARG, PEG |
 |
Crystal Structure Of Dimt1 In Complex With 5'-Methylthioadenosine From Pyrococcus Horikoshii (Formi)
Organism: Pyrococcus horikoshii ot3
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2024-08-28 Classification: TRANSFERASE Ligands: MTA, ZN, SO3, PEG, CL, GOL, EDO |
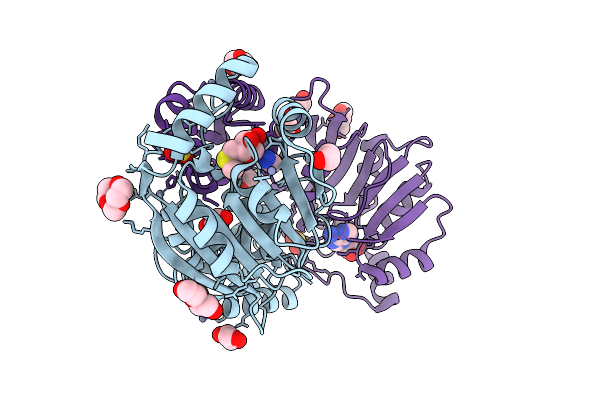 |
Crystal Structure Of Dimt1 In Complex With 5'-Methylthioadenosine From Pyrococcus Horikoshii (Formii)
Organism: Pyrococcus horikoshii ot3
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2024-08-28 Classification: TRANSFERASE Ligands: MTA, ZN, SO3, GOL, EDO, PEG, PGE, ACT |
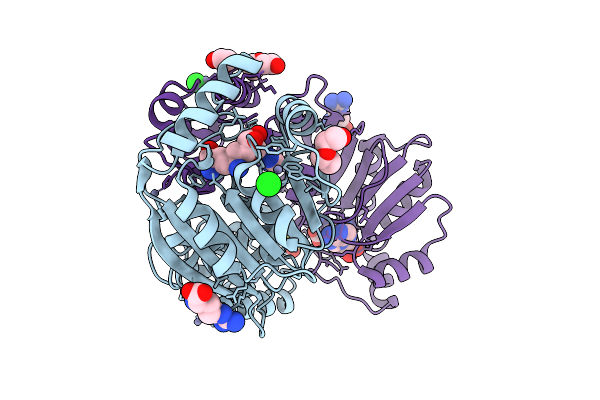 |
Crystal Structure Of Dimt1 In Complex With Adenosylornithine (Sfg) From Pyrococcus Horikoshii
Organism: Pyrococcus horikoshii ot3
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2024-08-28 Classification: TRANSFERASE Ligands: SFG, ZN, CL, PEG, ARG, SO3, EDO |
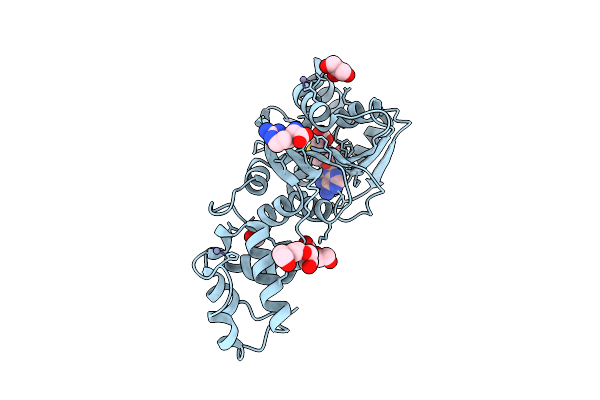 |
Crystal Structure Of Dimt1 In Complex With S-Adenosyl-L-Homocysteine (Sah) From Pyrococcus Horikoshii
Organism: Pyrococcus horikoshii ot3
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2024-08-28 Classification: TRANSFERASE Ligands: SAH, ZN, SO3, EDO, GOL, PEG, PGE, ARG |
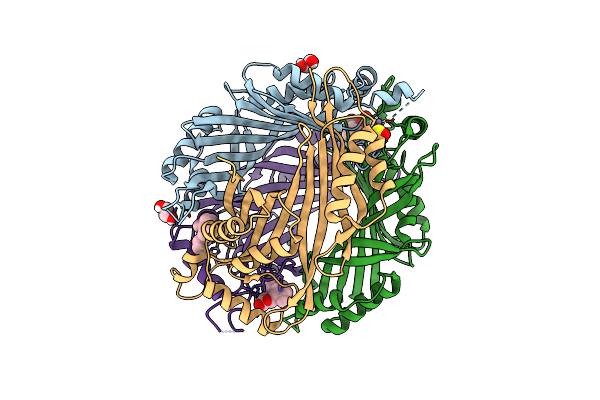
 |
Organism: Pyrococcus horikoshii ot3
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2024-08-28 Classification: TRANSFERASE Ligands: ZN, SO3, EDO, ARG, GOL, PEG |
 |
Organism: Burkholderia pseudomallei
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2024-07-03 Classification: HYDROLASE/INHIBITOR Ligands: MN, EDO, SO3, ZS9 |
 |
A Tethered Niacin-Derived Pincer Complex With A Nickel-Carbon Bond In Lactate Racemase R98A/R100A Variant Modeled With Separated Sulfite And Npn
Organism: Lactiplantibacillus plantarum wcfs1
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2023-01-25 Classification: ISOMERASE Ligands: EDO, SO3, 4EY, CA, MG, CL, NI |
 |
Toxoplasma Gondii Prolyl-Trna Synthetase (Tgprs) In Complex With Inhibitor L97 And L-Proline At 1.97 A Resolution
Organism: Toxoplasma gondii
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2022-09-07 Classification: LIGASE/LIGASE INHIBITOR Ligands: 1XK, PRO, GOL, CL, NA, SO3, ETA |
 |
Organism: Sus scrofa
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2022-07-06 Classification: ISOMERASE Ligands: CA, SO4, SO3 |

