Search Count: 55
All
Selected
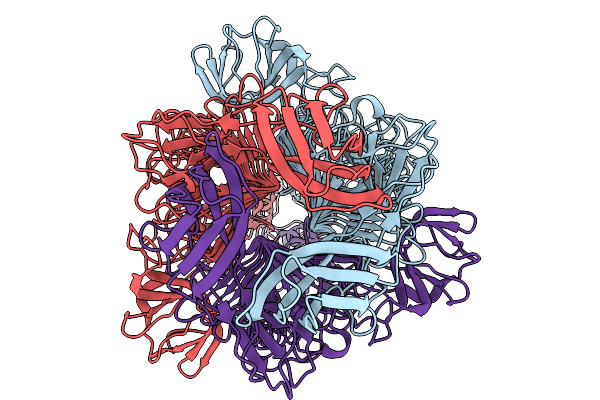 |
Organism: Klebsiella phage k64-1
Method: ELECTRON MICROSCOPY Resolution:2.24 Å Release Date: 2026-03-25 Classification: ANTIMICROBIAL PROTEIN |
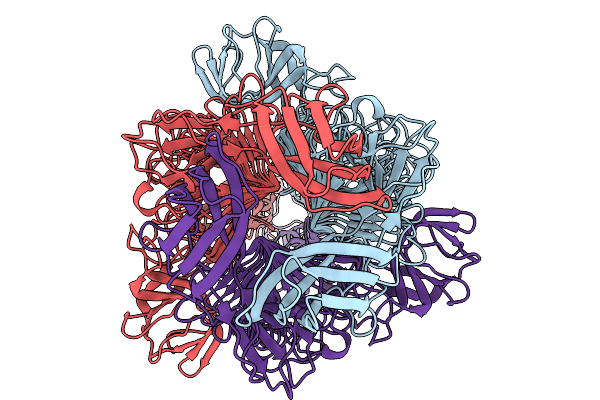 |
Organism: Klebsiella phage k64-1
Method: ELECTRON MICROSCOPY Resolution:2.18 Å Release Date: 2026-03-25 Classification: ANTIMICROBIAL PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2025-09-17 Classification: CELL CYCLE Ligands: A1JBJ |
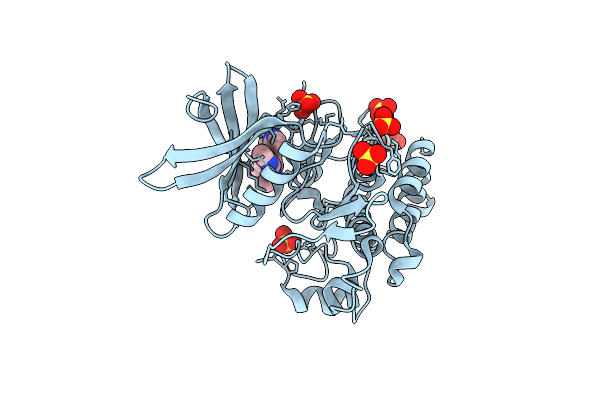 |
Crystal Structure Of Human Alk2 Kinase Domain With R206H Mutation In Complex With Rk783
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2025-06-25 Classification: SIGNALING PROTEIN Ligands: A1L4C, SO4 |
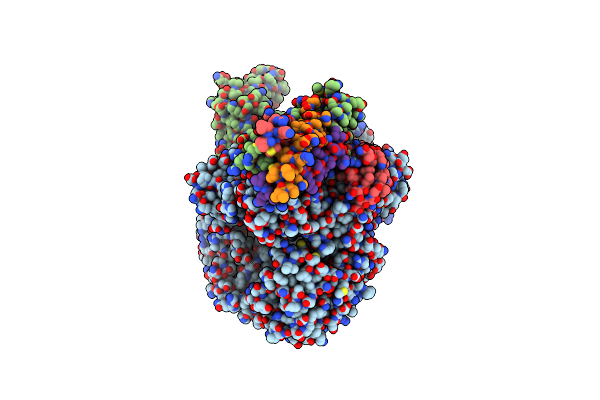 |
Cryo-Em Structure Of The Rna-Dependent Rna Polymerase Complex From Marburg Virus
Organism: Marburg virus - musoke, kenya, 1980, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2025-04-09 Classification: VIRAL PROTEIN Ligands: ZN |
 |
Cryo-Em Structure Of The Rna-Dependent Rna Polymerase Complex In A Compact Conformation From Ebola Virus
Organism: Escherichia coli (strain k12), Ebola virus - eckron (zaire, 1976)
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2025-04-09 Classification: VIRAL PROTEIN Ligands: ZN |
 |
Cryo-Em Structure Of The Rna-Dependent Rna Polymerase Complex From Marburg Virus
Organism: Marburg virus - musoke, kenya, 1980, Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2025-04-09 Classification: VIRAL PROTEIN Ligands: ZN |
 |
Organism: Homo sapiens, Mus musculus, Escherichia coli, Danio rerio, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-01-22 Classification: SIGNALING PROTEIN |
 |
Organism: Homo sapiens, Mus musculus, Escherichia coli, Danio rerio, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-01-22 Classification: SIGNALING PROTEIN/IMMUNE SYSTEM |
 |
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-04-24 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-03-20 Classification: HYDROLASE Ligands: ZN, CA, NAG, CLR, OLA, SPL |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-03-20 Classification: HYDROLASE Ligands: ZN, CA, NAG, CLR, OLA, SPL |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-03-20 Classification: HYDROLASE Ligands: NAG, CLR, ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-03-20 Classification: HYDROLASE Ligands: NAG, CLR, ZN |
 |
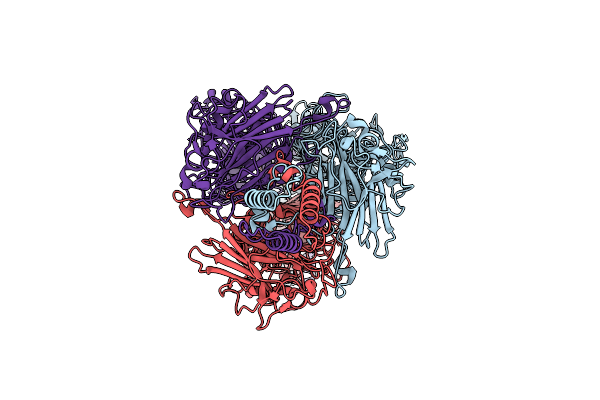
Cryo-Em Structure Of Depo32, A Klebsiella Phage Depolymerase Targets The K2 Serotype K. Pneumoniae
Organism: Klebsiella phage gh-k3
Method: ELECTRON MICROSCOPY Resolution:2.32 Å Release Date: 2023-08-30 Classification: LYASE |
 |
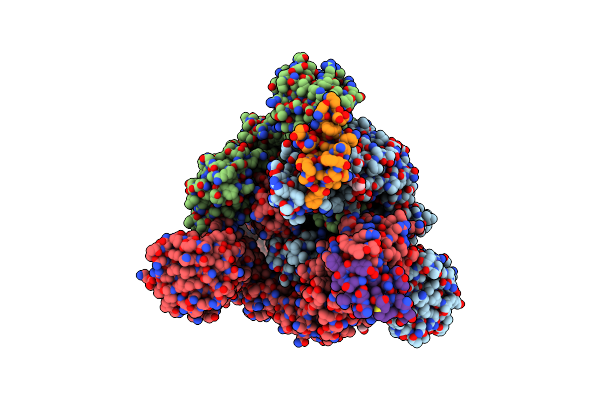
Cryo-Em Structure Of Depo32, A Klebsiella Phage Depolymerase Targets The K2 Serotype K. Pneumoniae
Organism: Klebsiella phage gh-k3
Method: ELECTRON MICROSCOPY Resolution:2.46 Å Release Date: 2023-08-30 Classification: HYDROLASE |
 |
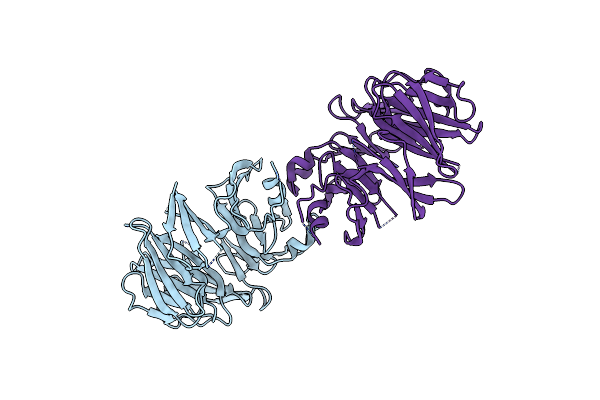
Conformation 1 Of Sars-Cov-2 Omicron Ba.1 Variant Spike Protein Complexed With Mo1 Fab
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-05-10 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Conformation 2 Of Sars-Cov-2 Omicron Ba.1 Variant Spike Protein Complexed With Mo1 Fab
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-05-10 Classification: VIRAL PROTEIN Ligands: NAG |
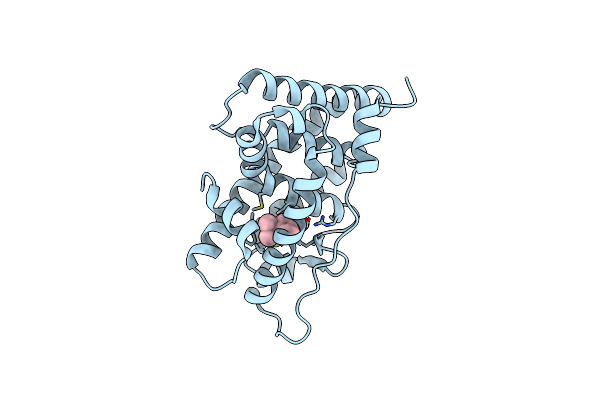 |
Crystal Structure Of Human Estrogen-Related Receptor Gamma Ligand Binding Domain Complex With Bpa-Monof
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2020-05-13 Classification: TRANSCRIPTION Ligands: CW6 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2019-01-09 Classification: CYTOSOLIC PROTEIN |

