Search Count: 53
All
Selected
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2025-12-10 Classification: PROTEIN FIBRIL |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2025-12-10 Classification: PROTEIN FIBRIL |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2025-12-10 Classification: PROTEIN FIBRIL |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2025-12-10 Classification: PROTEIN FIBRIL |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2025-12-10 Classification: PROTEIN FIBRIL |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:2.20 Å Release Date: 2025-12-10 Classification: PROTEIN FIBRIL |
 |
Organism: Citrus sinensis, Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2025-08-20 Classification: PLANT PROTEIN Ligands: A1IRJ, MN, CL |
 |
Organism: Citrus sinensis, Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-08-20 Classification: PLANT PROTEIN Ligands: A1IRN, MN, CL |
 |
X-Ray Crystal Structure Of The Cspyl1 5M (V112L, T135L, F137I, T153I, V168A)-Icb-Hab1 Ternary Complex
Organism: Citrus sinensis, Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2025-08-20 Classification: PLANT PROTEIN Ligands: A1IRN, MN, CL |
 |
Organism: Solanum lycopersicum
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2025-07-02 Classification: PLANT PROTEIN Ligands: HC4, DMS |
 |
Organism: Solanum lycopersicum
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-07-02 Classification: PLANT PROTEIN Ligands: FER, DMS |
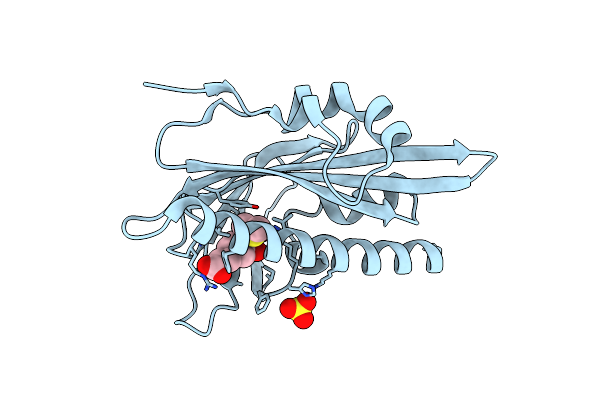 |
Organism: Solanum lycopersicum
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2025-07-02 Classification: PLANT PROTEIN Ligands: DHC, DMS, SO4 |
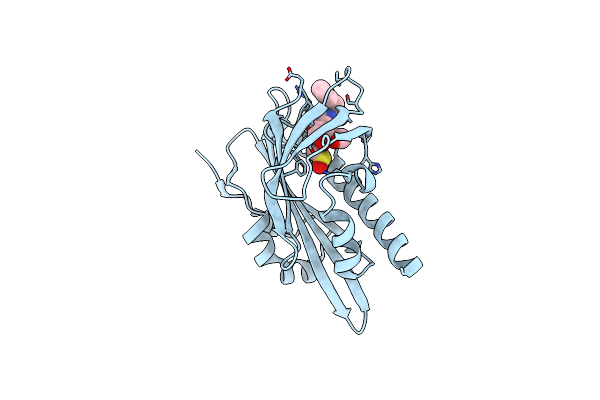 |
Organism: Solanum lycopersicum
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-07-02 Classification: PLANT PROTEIN Ligands: A1I70, DMS |
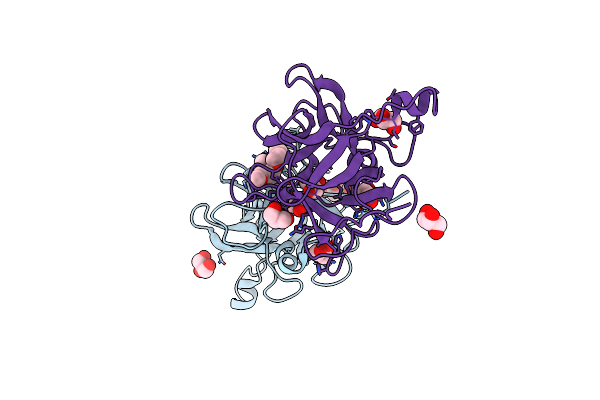 |
Organism: Solanum tuberosum
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2025-05-14 Classification: PLANT PROTEIN Ligands: GOL, ETE, MAN |
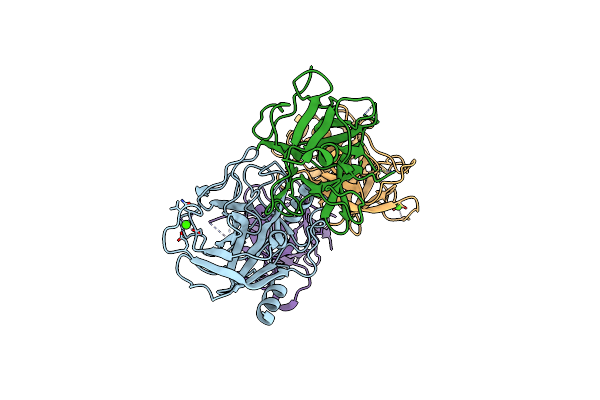 |
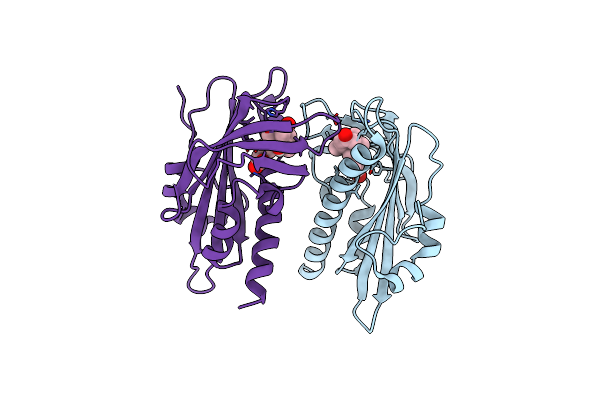
Complex Between The Porcine Trypsin And M271 A Kunitz-Sti From Solanum Tuberosum
Organism: Solanum tuberosum, Sus scrofa
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2025-05-14 Classification: HYDROLASE Ligands: CA |
 |
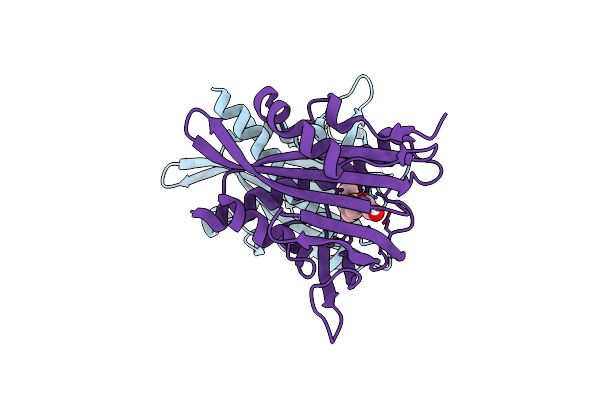
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2024-10-16 Classification: PLANT PROTEIN Ligands: A8S |
 |
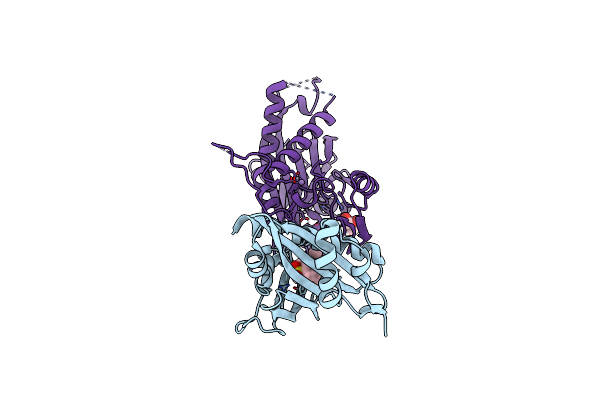
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2024-10-16 Classification: PLANT PROTEIN Ligands: P6G |
 |
Organism: Citrus sinensis, Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2023-03-22 Classification: PLANT PROTEIN Ligands: OE3, MN, CL, GOL |
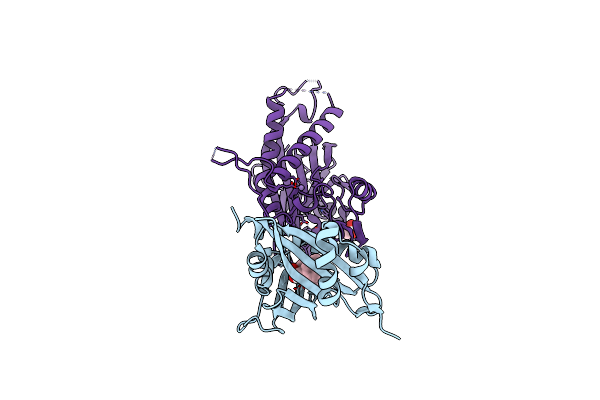 |
X-Ray Crystal Structure Of The Cspyl1(V112L, T135L,F137I, T153I, V168A)-Sb-Hab1 Ternary Complex
Organism: Citrus sinensis, Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2023-03-22 Classification: PLANT PROTEIN Ligands: QPZ, MN, CL, GOL |
 |
X-Ray Crystal Structure Of The Cspyl1(V112L, T135L,F137I, T153I, V168A)-Isb7-Hab1 Ternary Complex
Organism: Citrus sinensis, Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2023-03-22 Classification: PLANT PROTEIN Ligands: S5N, GOL, MN, CL |

