Search Count: 85
All
Selected
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2026-01-14 Classification: LIPID BINDING PROTEIN Ligands: GDP, GOL |
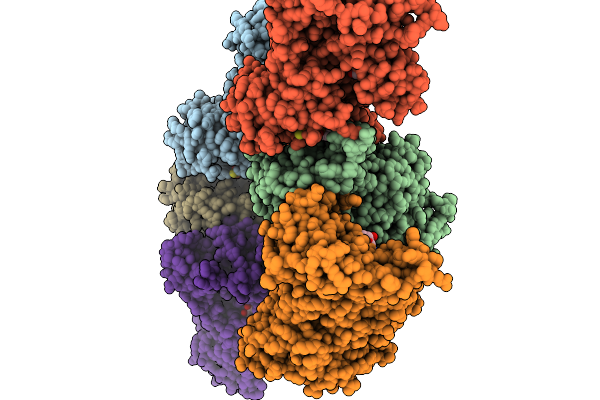 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2026-01-14 Classification: LIPID BINDING PROTEIN Ligands: GOL, GSP, MG |
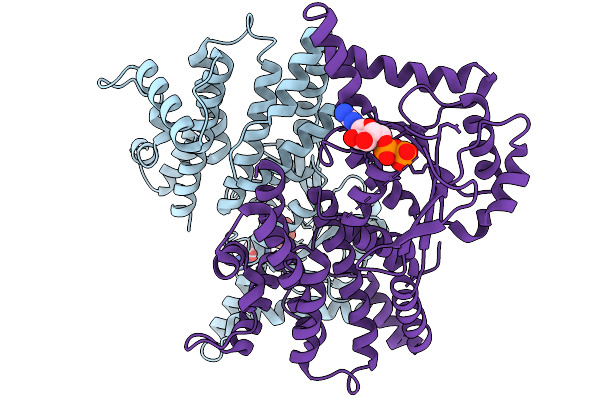 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:3.31 Å Release Date: 2026-01-14 Classification: LIPID BINDING PROTEIN Ligands: GDP |
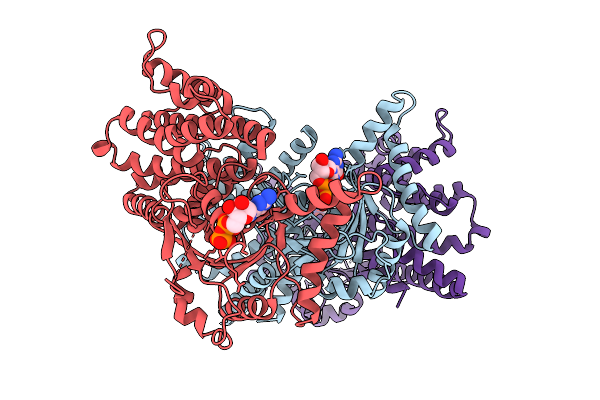 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-01-14 Classification: LIPID BINDING PROTEIN Ligands: GDP |
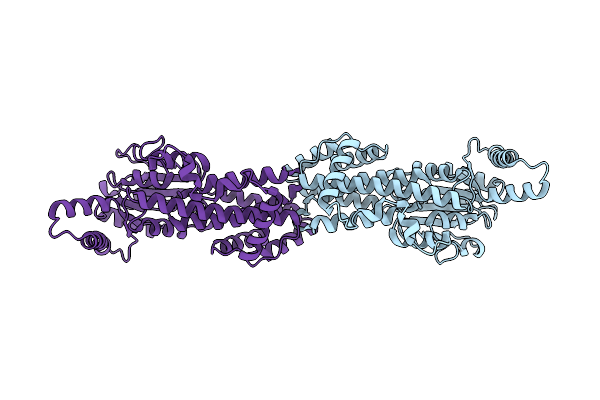 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: LIPID BINDING PROTEIN |
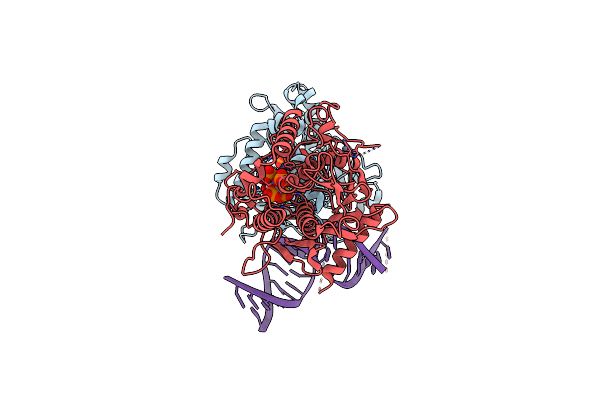 |
Cryo-Em Structure Of Human Adar1 In Complex With Dsrna Derived From Human Gli1 Gene
Organism: Escherichia coli, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-03-26 Classification: HYDROLASE/RNA Ligands: IHP, ZN |
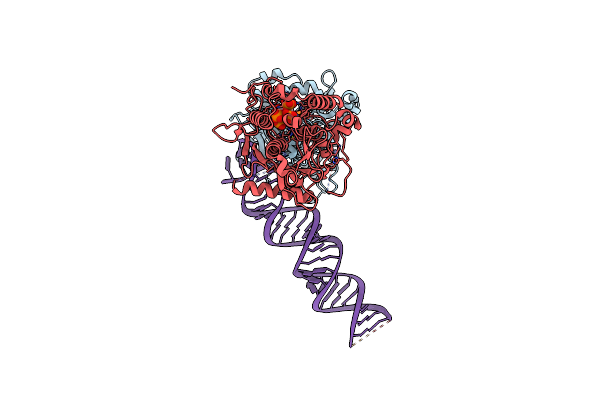 |
Cryo-Em Structure Of Human Adar1 In Complex With Dsrna Derived From Ht2C Gene
Organism: Escherichia coli, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-03-26 Classification: HYDROLASE/RNA Ligands: IHP, ZN |
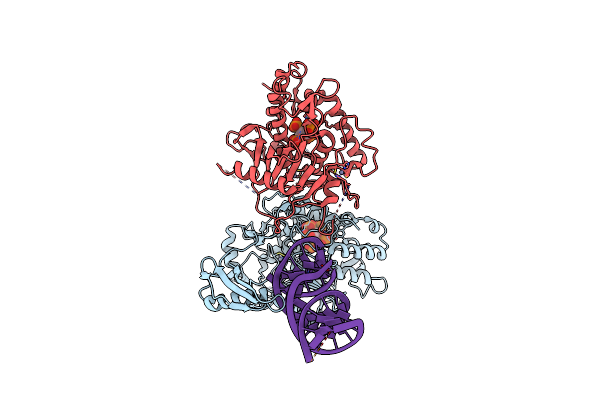 |
Cryo-Em Structure Of Human Adar1 In Complex With Dsrna Derived From Ht2C Gene In The Pre-Editing State
Organism: Escherichia coli, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-03-26 Classification: HYDROLASE/RNA Ligands: IHP, ZN |
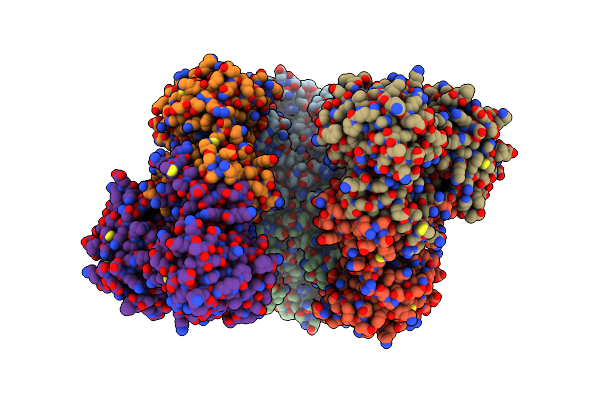 |
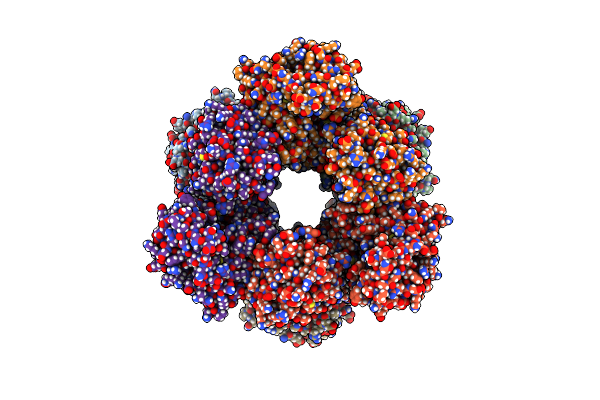
The Cryo-Em Structure Of Recombinantly Expressed Hugdh In Complex With Udp-4-Keto-Xylose
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: OXIDOREDUCTASE/INHIBITOR Ligands: A1BCU |
 |
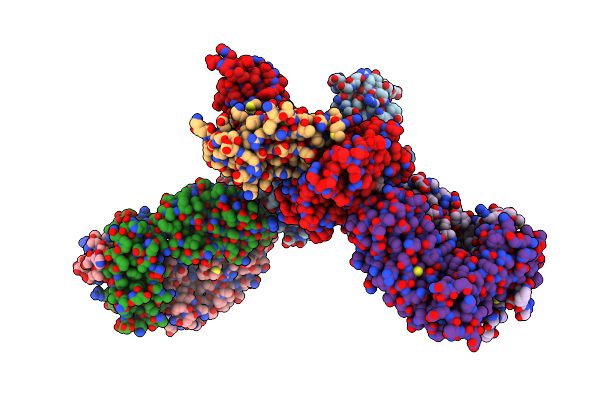
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: OXIDOREDUCTASE |
 |
Tcr In Complex With Hla-E*01:03 Bound To Hbv Envelope 371-379 Index Peptide
Organism: Homo sapiens, Hepatitis b virus
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2024-10-16 Classification: IMMUNE SYSTEM |
 |
Organism: Homo sapiens, Hepatitis b virus
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2024-10-16 Classification: IMMUNE SYSTEM |
 |
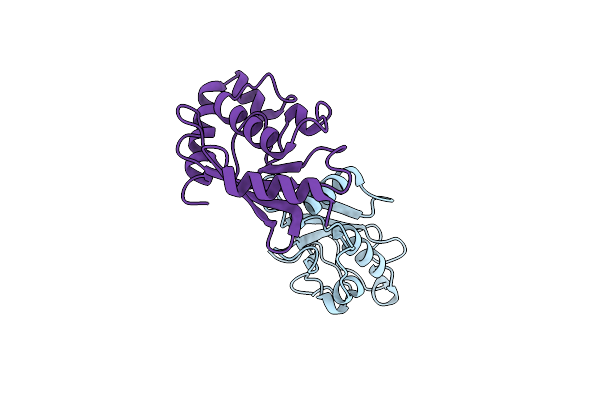
Organism: Homo sapiens, Hepatitis b virus
Method: X-RAY DIFFRACTION Resolution:2.61 Å Release Date: 2024-10-16 Classification: IMMUNE SYSTEM |
 |
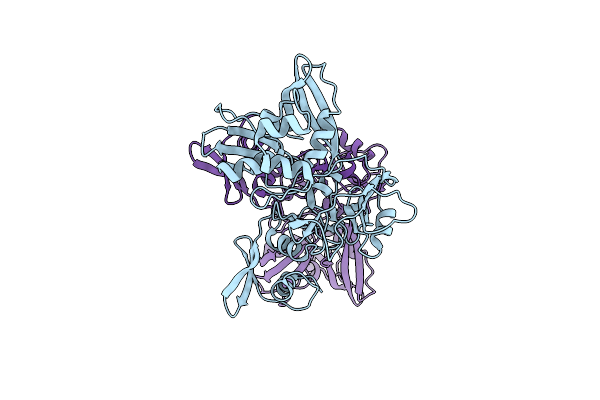
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2024-05-08 Classification: LIGASE |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.91 Å Release Date: 2023-06-14 Classification: RNA BINDING PROTEIN,VIRAL PROTEIN,LYASE |
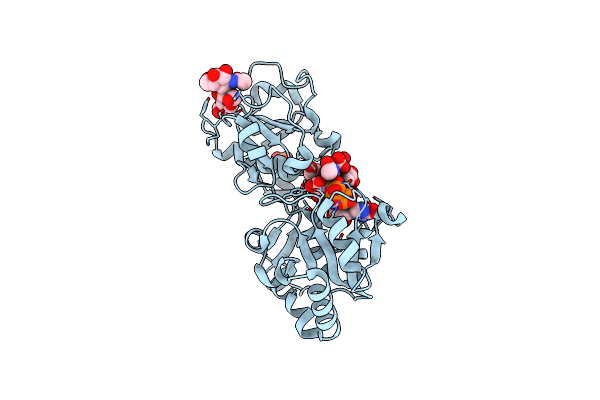 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2023-05-31 Classification: TRANSFERASE Ligands: 6CK, GDP, SO4 |
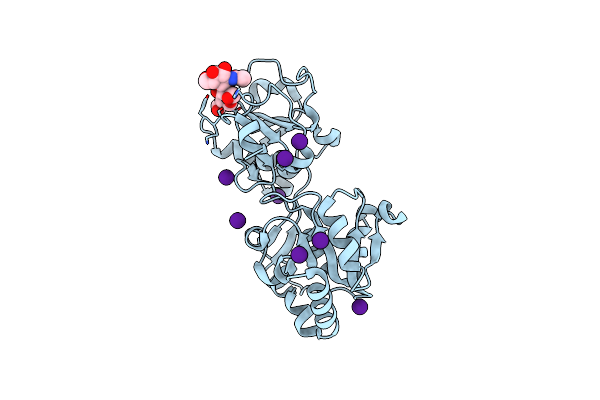 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2023-05-24 Classification: TRANSFERASE Ligands: IOD, CS |
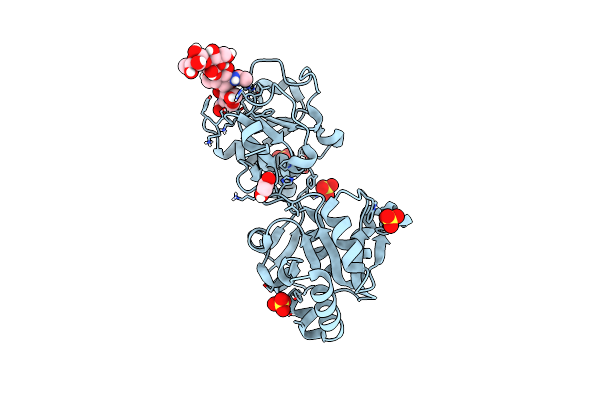 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.09 Å Release Date: 2023-05-24 Classification: TRANSFERASE Ligands: SO4, EDO |
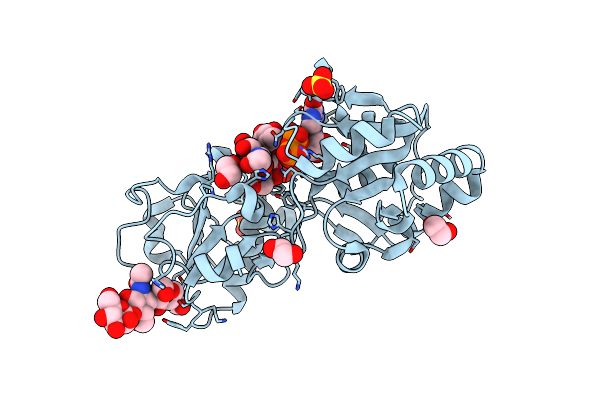 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2023-05-24 Classification: TRANSFERASE Ligands: GDP, SO4, EDO |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2023-05-24 Classification: TRANSFERASE Ligands: GDP, SO4, EDO |

