Search Count: 16
All
Selected
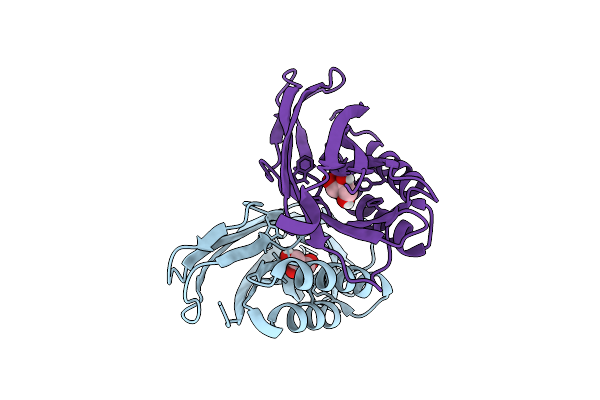 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-11-26 Classification: CELL ADHESION Ligands: GOL |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2025-11-26 Classification: CELL ADHESION Ligands: GOL, FMT, MG |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2025-11-26 Classification: CELL ADHESION Ligands: GOL |
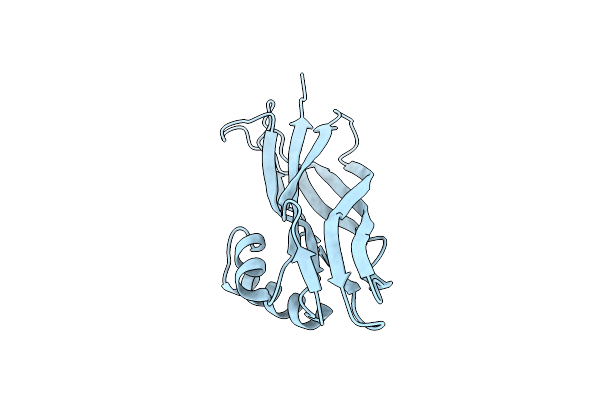 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2025-06-11 Classification: CELL ADHESION |
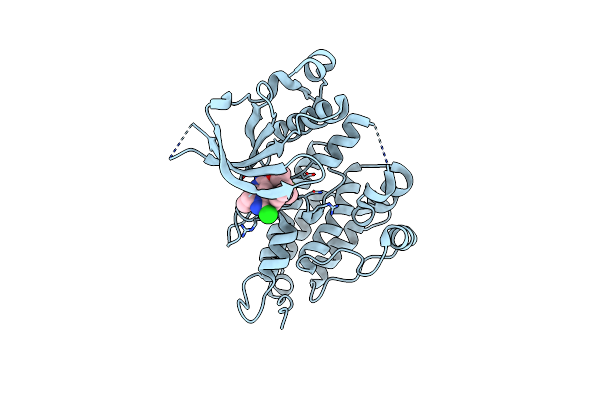 |
Structure Of Human Anaplastic Lymphoma Kinase (Alk) Harboring The G1202R/L1196M Compound Mutation In Complex With Nvl-655
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2024-09-18 Classification: ONCOPROTEIN Ligands: A1IJ8 |
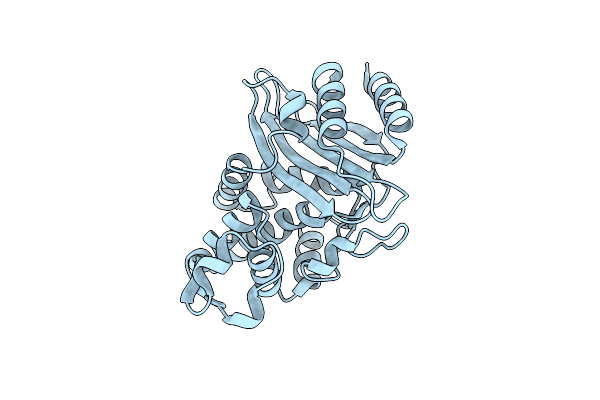 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2020-04-22 Classification: HYDROLASE |
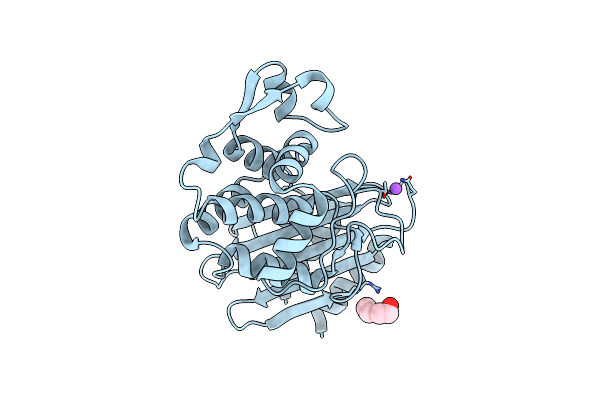 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2020-04-22 Classification: HYDROLASE Ligands: PEG, NA |
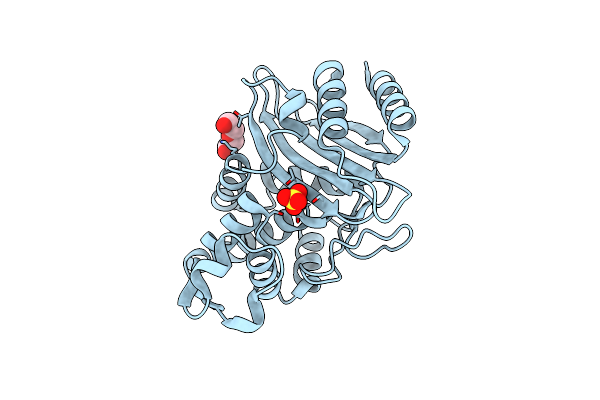 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2020-04-22 Classification: HYDROLASE Ligands: SO4, PEG |
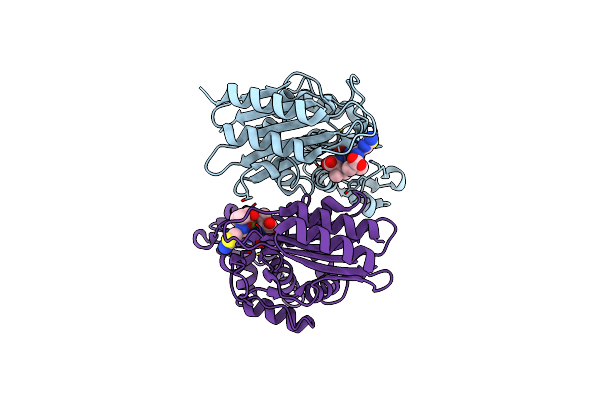 |
Crystal Structure Of Ctx-M-14 E166A/D240G Beta-Lactamase In Complex With Ceftazidime
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2020-04-22 Classification: HYDROLASE Ligands: CAZ |
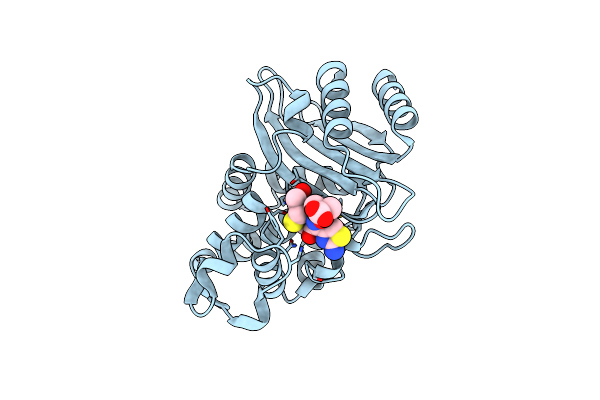 |
Crystal Structure Of Ctx-M-14 E166A/P167S/D240G Beta-Lactamase In Complex With Ceftazidime-1
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2020-04-22 Classification: HYDROLASE Ligands: CAZ |
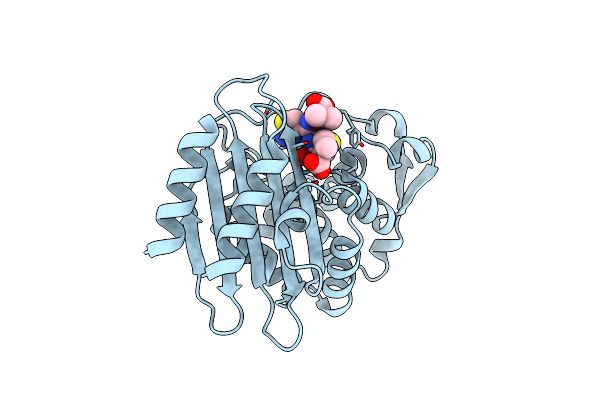 |
Crystal Structure Of Ctx-M-14 E166A/P167S/D240G Beta-Lactamase In Complex With Ceftazidime-2
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2020-04-22 Classification: HYDROLASE Ligands: CAZ |
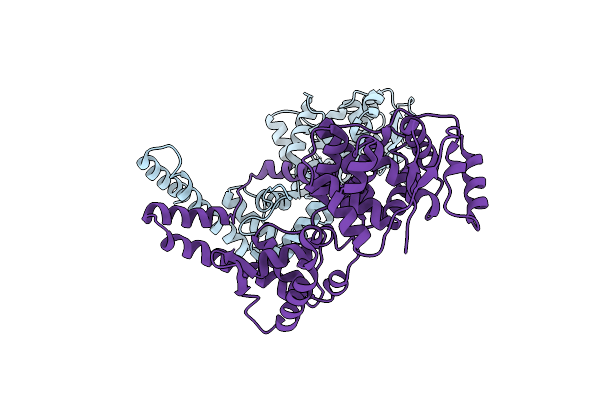 |
Organism: Zaire ebolavirus
Method: ELECTRON MICROSCOPY Release Date: 2018-03-07 Classification: RNA BINDING PROTEIN |
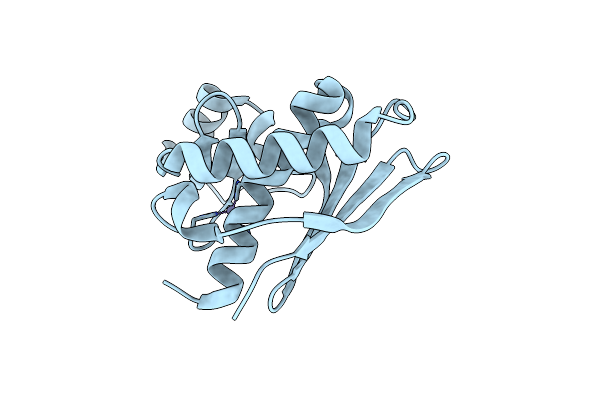 |
Epitope Backbone Grafting By Computational Design For Improved Presentation Of Linear Epitopes On Scaffold Proteins
Organism: Pyrococcus horikoshii
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2011-11-09 Classification: DE NOVO PROTEIN Ligands: ZN |
 |
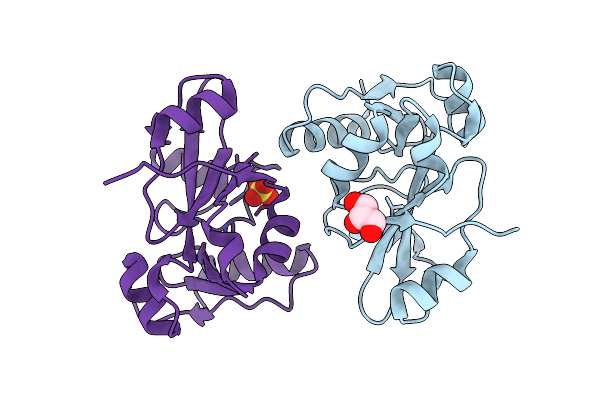
Epitope Backbone Grafting By Computational Design For Improved Presentation Of Linear Epitopes On Scaffold Proteins
Organism: Pyrococcus horikoshii
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2011-11-09 Classification: DE NOVO PROTEIN |
 |
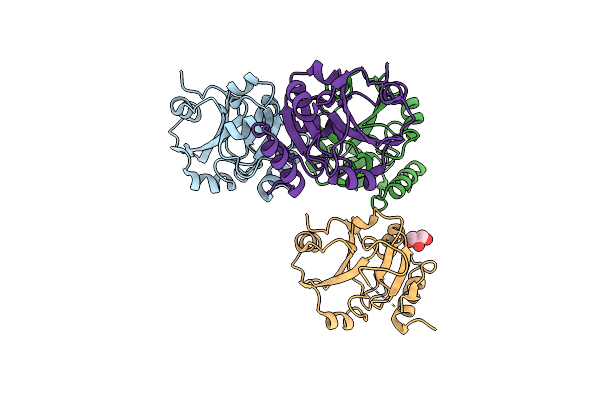
Epitope Backbone Grafting By Computational Design For Improved Presentation Of Linear Epitopes On Scaffold Proteins
Organism: Thermus thermophilus
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2011-11-09 Classification: DE NOVO PROTEIN Ligands: GOL, SO4 |
 |
Epitope Backbone Grafting By Computational Design For Improved Presentation Of Linear Epitopes On Scaffold Proteins
Organism: Thermus thermophilus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2011-11-09 Classification: DE NOVO PROTEIN Ligands: GOL |

