Search Count: 47
All
Selected
 |
Organism: Pseudomonas phage phikz
Method: ELECTRON MICROSCOPY Resolution:3.37 Å Release Date: 2026-02-11 Classification: VIRAL PROTEIN |
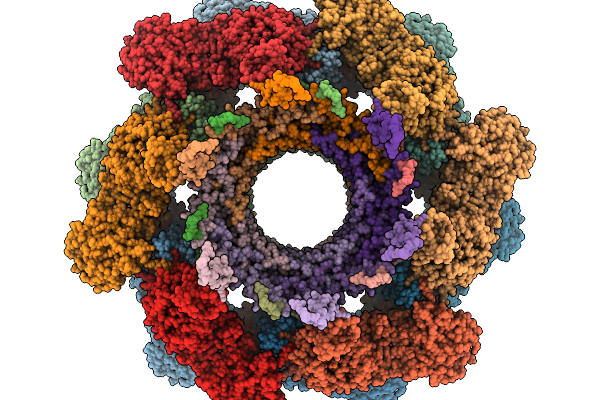 |
Organism: Pseudomonas phage phikz
Method: ELECTRON MICROSCOPY Resolution:3.15 Å Release Date: 2026-02-11 Classification: VIRAL PROTEIN |
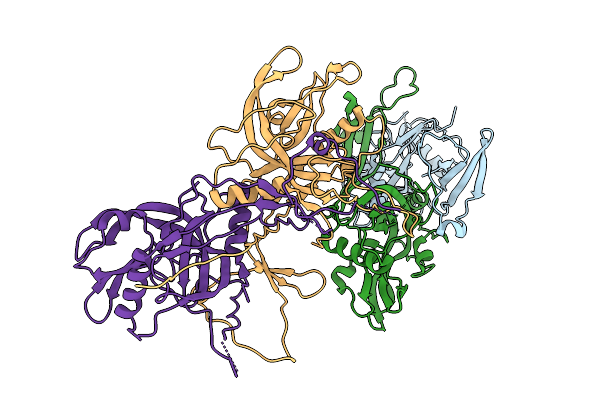 |
Organism: Pseudomonas phage phikz
Method: ELECTRON MICROSCOPY Resolution:4.14 Å Release Date: 2026-02-11 Classification: VIRAL PROTEIN |
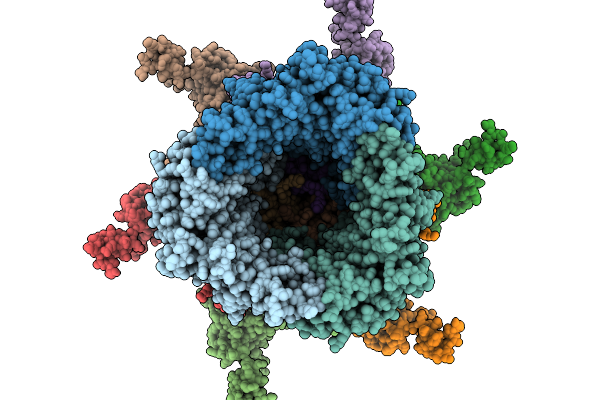 |
Organism: Pseudomonas phage phikz
Method: ELECTRON MICROSCOPY Resolution:4.02 Å Release Date: 2026-02-11 Classification: VIRAL PROTEIN |
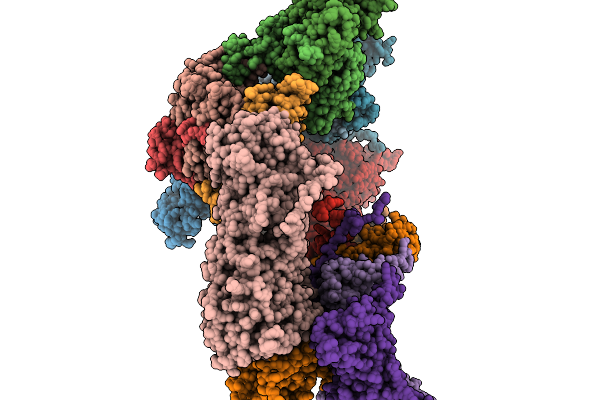 |
Organism: Pseudomonas phage phikz
Method: ELECTRON MICROSCOPY Resolution:3.52 Å Release Date: 2026-02-11 Classification: VIRAL PROTEIN |
 |
The Structure Of Outer Peripheral Region In The Phage Phikz Baseplate Complex
Organism: Pseudomonas phage phikz
Method: ELECTRON MICROSCOPY Resolution:4.53 Å Release Date: 2026-02-11 Classification: VIRAL PROTEIN |
 |
Structure Of The Inner Peripheral Region In The Phage Phikz Baseplate Complex
Organism: Pseudomonas phage phikz
Method: ELECTRON MICROSCOPY Resolution:3.65 Å Release Date: 2026-02-11 Classification: VIRAL PROTEIN |
 |
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2025-10-29 Classification: DE NOVO PROTEIN |
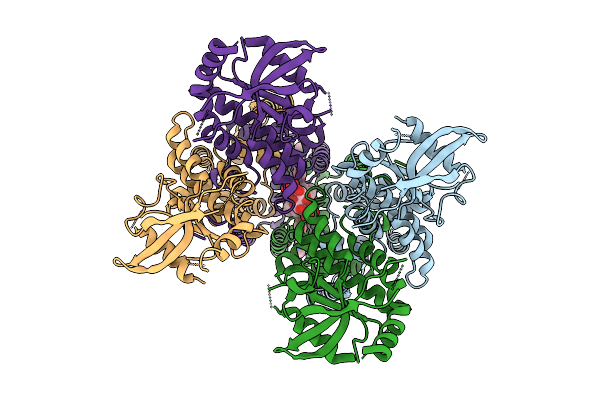 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.71 Å Release Date: 2025-10-15 Classification: MEMBRANE PROTEIN Ligands: A1EDL, A1EER |
 |
Organism: Escherichia phage t1
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2025-03-12 Classification: VIRAL PROTEIN |
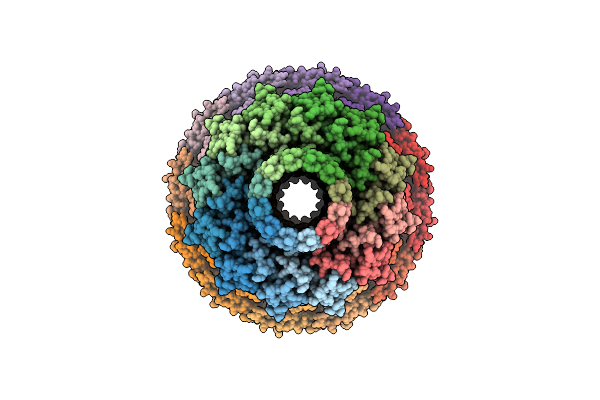 |
Organism: Escherichia phage t1
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2025-03-12 Classification: VIRAL PROTEIN |
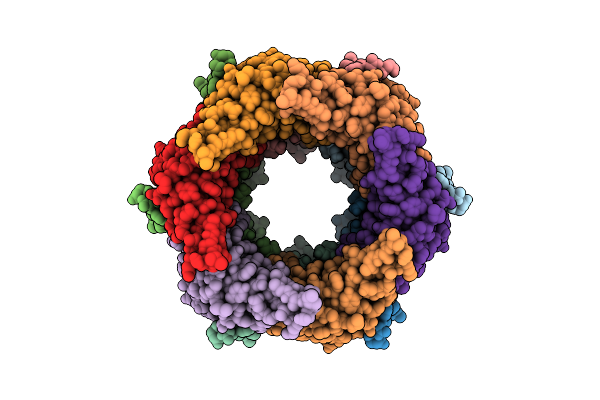 |
Organism: Escherichia phage t1
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2025-03-12 Classification: VIRAL PROTEIN |
 |
Organism: Escherichia phage t1
Method: ELECTRON MICROSCOPY Resolution:4.10 Å Release Date: 2025-03-12 Classification: VIRAL PROTEIN |
 |
Organism: Escherichia phage t1
Method: ELECTRON MICROSCOPY Resolution:4.30 Å Release Date: 2025-03-12 Classification: VIRAL PROTEIN Ligands: SF4 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-11-13 Classification: MEMBRANE PROTEIN |
 |
Structure Of The Extracellular Subdomain Of A Homomeric Lrrc8C Truncation Disease Mutant
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-11-13 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-11-13 Classification: MEMBRANE PROTEIN |
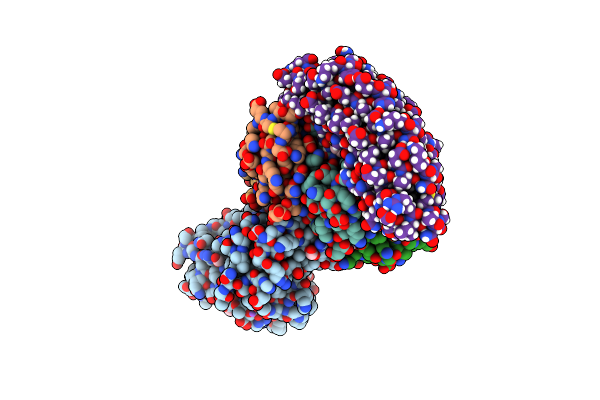 |
Organism: Synthetic construct, Escherichia phage 933w
Method: ELECTRON MICROSCOPY Release Date: 2023-04-12 Classification: ANTITOXIN Ligands: 1PS, FMT, NA, EDO |
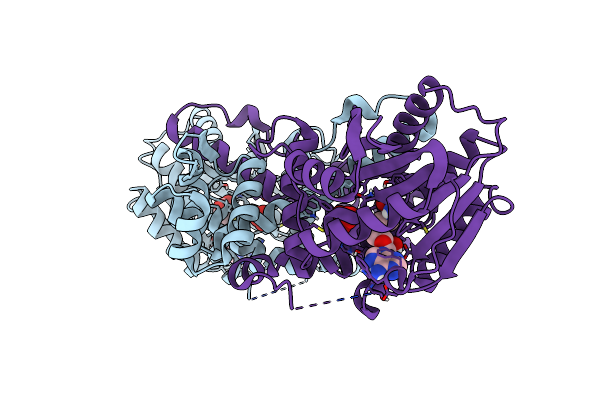 |
Crystal Structure Of Mfng, An L- And D-Tyrosine O-Methyltransferase From The Marformycin Biosynthesis Pathway Of Streptomyces Drozdowiczii, With Sah Bound At 1.35 A Resolution (P212121 - Form I)
Organism: Streptomyces drozdowiczii
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2022-10-12 Classification: TRANSFERASE Ligands: SAH, UNL |
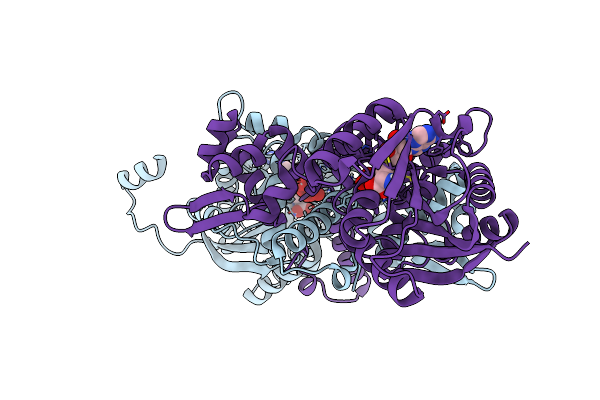 |
Crystal Structure Of Mfng, An L- And D-Tyrosine O-Methyltransferase From The Marformycin Biosynthesis Pathway Of Streptomyces Drozdowiczii, With Sah Bound At 1.2 A Resolution (P212121 - Form Ii)
Organism: Streptomyces drozdowiczii
Method: X-RAY DIFFRACTION Resolution:1.14 Å Release Date: 2022-10-12 Classification: TRANSFERASE Ligands: SAH, UNL |

