Search Count: 34
All
Selected
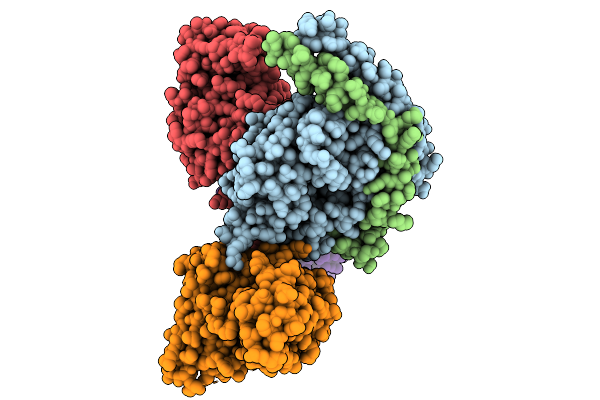 |
Cryo-Em Structure Of Gi Coupled Sphingosine 1-Phosphate Receptor Bound With Cym5442
Organism: Homo sapiens, Mus musculus, Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.69 Å Release Date: 2026-01-28 Classification: SIGNALING PROTEIN Ligands: A1L3G |
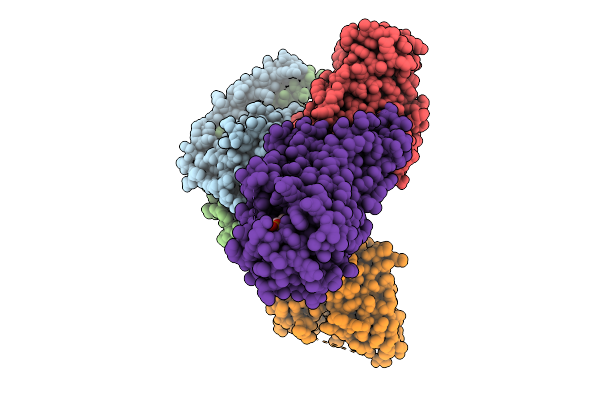 |
Cryo-Em Structure Of Gi Coupled Sphingosine 1-Phosphate Receptor Bound With Hy-X-1011
Organism: Homo sapiens, Mus musculus, Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.79 Å Release Date: 2026-01-28 Classification: SIGNALING PROTEIN Ligands: A1ESY |
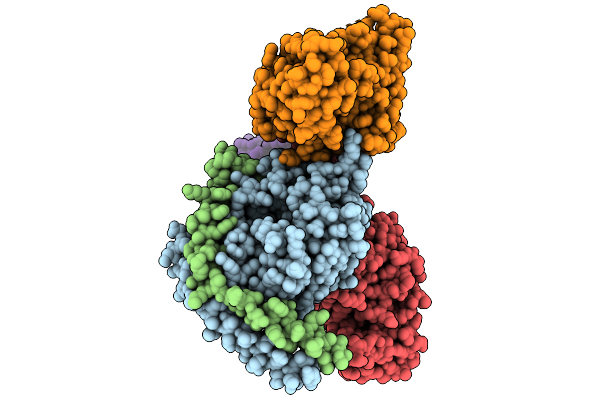 |
Cryo-Em Structure Of Gi Coupled Sphingosine 1-Phosphate Receptor Bound With Ponesimod
Organism: Homo sapiens, Mus musculus, Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.79 Å Release Date: 2026-01-28 Classification: SIGNALING PROTEIN Ligands: A1ESZ |
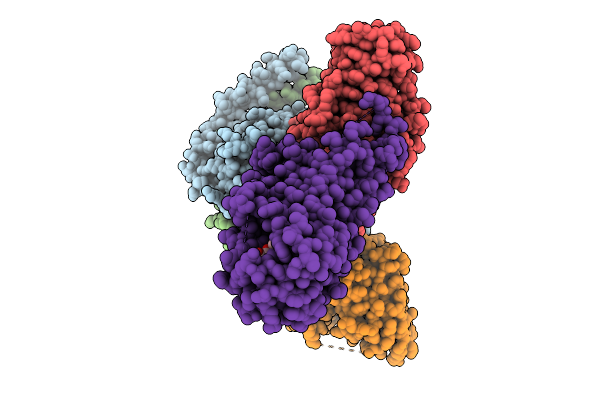 |
Cryo-Em Structure Of Gi Coupled Sphingosine 1-Phosphate Receptor Bound With Sar247799
Organism: Homo sapiens, Mus musculus, Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.97 Å Release Date: 2026-01-28 Classification: SIGNALING PROTEIN Ligands: A1LYQ |
 |
Organism: Echinacea
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-03-05 Classification: PLANT PROTEIN |
 |
Organism: Echinacea
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2025-03-05 Classification: PLANT PROTEIN |
 |
Organism: Homo sapiens, Pyrococcus abyssi (strain ge5 / orsay)
Method: ELECTRON MICROSCOPY Release Date: 2024-01-10 Classification: LIPID TRANSPORT Ligands: S1P |
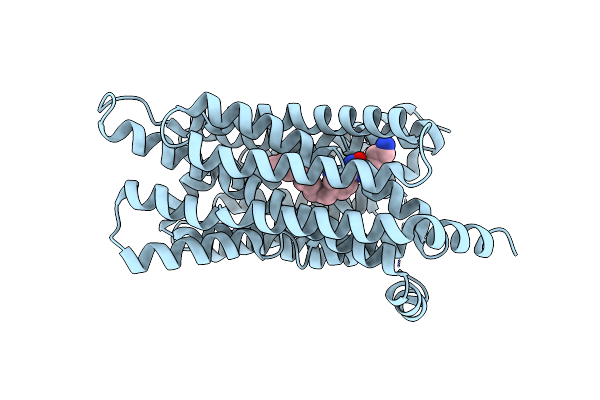 |
Cryo-Em Structure Of Human S1P Transporter Spns2 Bound With An Inhibitor 16D
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-01-10 Classification: LIPID TRANSPORT Ligands: YUX |
 |
Organism: Homo sapiens, Escherichia coli, Oplophorus gracilirostris
Method: ELECTRON MICROSCOPY Release Date: 2023-11-29 Classification: SIGNALING PROTEIN Ligands: CLR |
 |
Organism: Homo sapiens, Escherichia coli, Oplophorus gracilirostris
Method: ELECTRON MICROSCOPY Release Date: 2023-11-29 Classification: SIGNALING PROTEIN Ligands: CLR, 4AI |
 |
Organism: Homo sapiens, Escherichia coli, Oplophorus gracilirostris
Method: ELECTRON MICROSCOPY Release Date: 2023-11-29 Classification: SIGNALING PROTEIN Ligands: CLR, 4IE |
 |
Organism: Lama glama, Escherichia coli, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-11-29 Classification: SIGNALING PROTEIN Ligands: 43I, CLR |
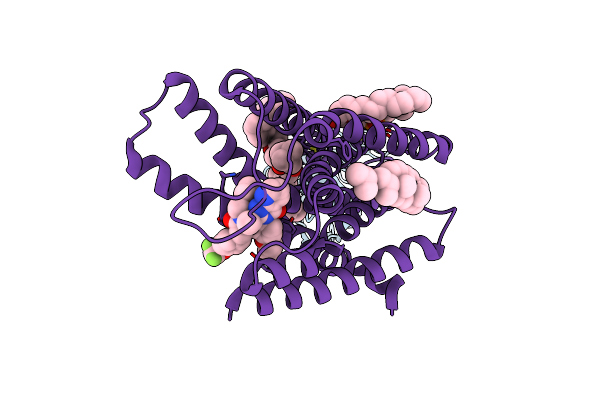 |
Organism: Lama glama, Escherichia coli, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-11-29 Classification: SIGNALING PROTEIN Ligands: CLR, PCW, LPC, FI6 |
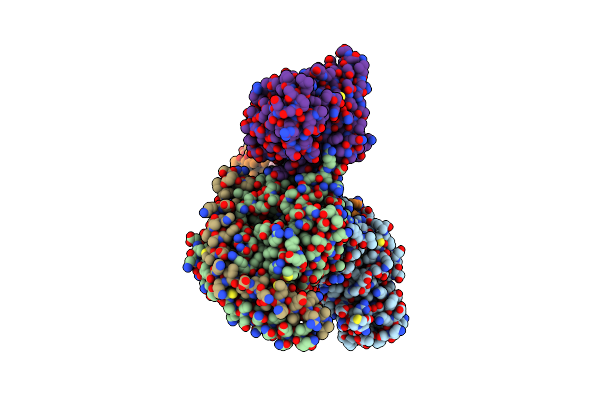 |
Organism: Homo sapiens, Escherichia coli, Oplophorus gracilirostris
Method: ELECTRON MICROSCOPY Release Date: 2023-11-29 Classification: SIGNALING PROTEIN Ligands: CLR |
 |
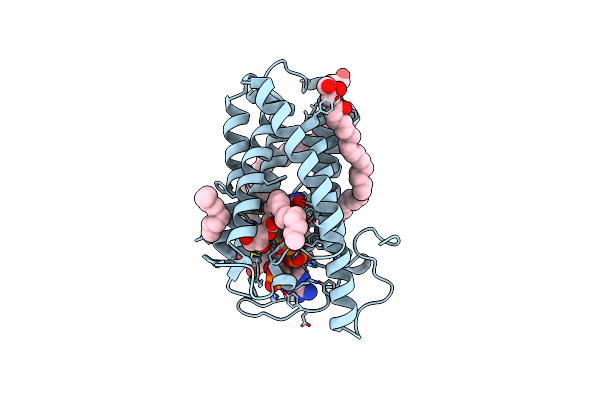
Organism: Homo sapiens, Mus musculus, Escherichia coli, Vicugna pacos
Method: ELECTRON MICROSCOPY Release Date: 2023-05-31 Classification: SIGNALING PROTEIN Ligands: U2U |
 |
Crystal Structure Of An Integral Membrane Steroid 5-Alpha-Reductase Pbsrd5A
Organism: Proteobacteria bacterium
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2021-01-27 Classification: OXIDOREDUCTASE Ligands: NDP, OLC |
 |
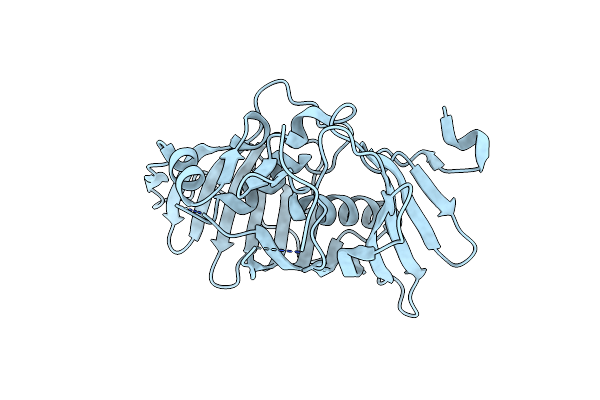
Organism: Influenza a virus (strain a/wilson-smith/1933 h1n1)
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2020-01-01 Classification: VIRAL PROTEIN Ligands: B7O |
 |
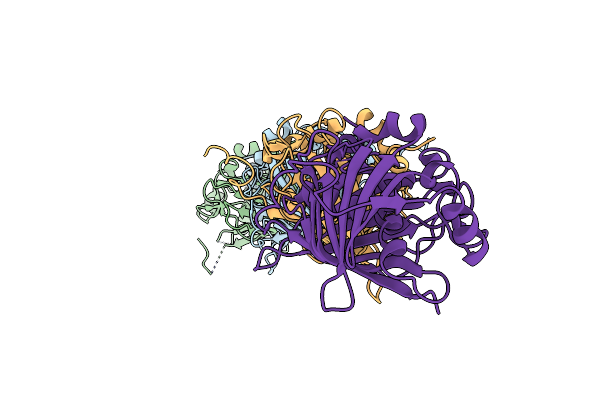
Structural Insights Into Dehydratase Substrate Selection For The Borrelidin And Fluvirucin Polyketide Synthases
Organism: Streptomyces parvulus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2019-06-05 Classification: LYASE |
 |
Structural Insights Into Dehydratase Substrate Selection For The Borrelidin And Fluvirucin Polyketide Synthases
Organism: Actinomadura vulgaris
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2019-06-05 Classification: LYASE |
 |
Crystal Structure Of Ndm-1 At Ph5.5 (Bis-Tris) In Complex With Hydrolyzed Ampicillin
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:1.00 Å Release Date: 2018-08-22 Classification: HYDROLASE/ANTIBIOTIC Ligands: ZZ7, ZN, OH |

