Search Count: 23
All
Selected
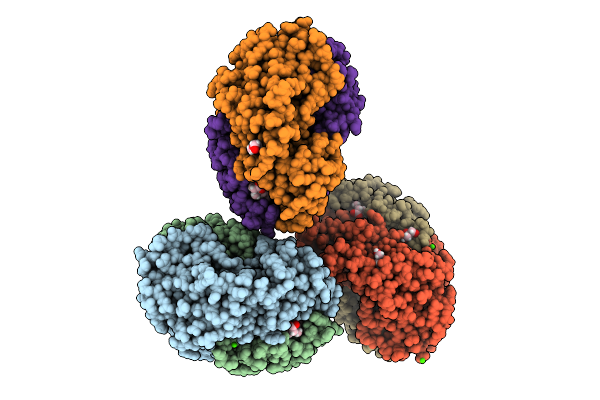 |
Organism: Streptomyces sp. b93
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2026-04-08 Classification: LYASE Ligands: HEM, GOL, CA, PEG |
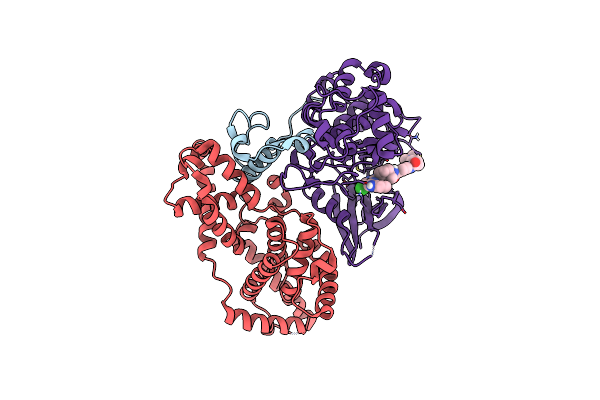 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-12-25 Classification: TRANSCRIPTION Ligands: A1H46 |
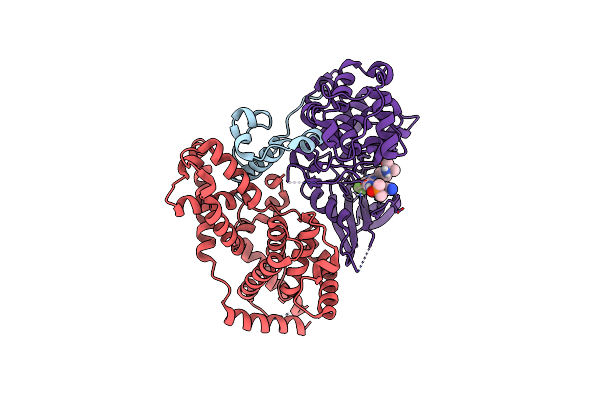 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-12-25 Classification: TRANSCRIPTION Ligands: YNK |
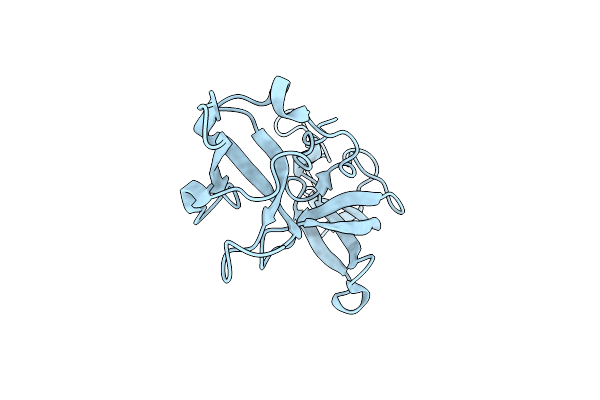 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2021-02-24 Classification: LIGASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2021-01-27 Classification: LIGASE Ligands: RVV, BGC |
 |
Organism: Pleurotus ostreatus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2020-08-12 Classification: SUGAR BINDING PROTEIN Ligands: CA, MLI |
 |
Crystal Structure Of Lectin From Pleurotus Ostreatus In Complex With Glycerol
Organism: Pleurotus ostreatus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2020-08-05 Classification: SUGAR BINDING PROTEIN Ligands: CA, GOL |
 |
Crystal Structure Of Lectin From Pleurotus Ostreatus In Complex With Rhamnose
Organism: Pleurotus ostreatus
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2020-08-05 Classification: SUGAR BINDING PROTEIN Ligands: CA, RAM |
 |
Crystal Structure Of Lectin From Pleurotus Ostreatus In Complex With Galnac
Organism: Pleurotus ostreatus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2020-08-05 Classification: SUGAR BINDING PROTEIN Ligands: CA, GOL, NGA |
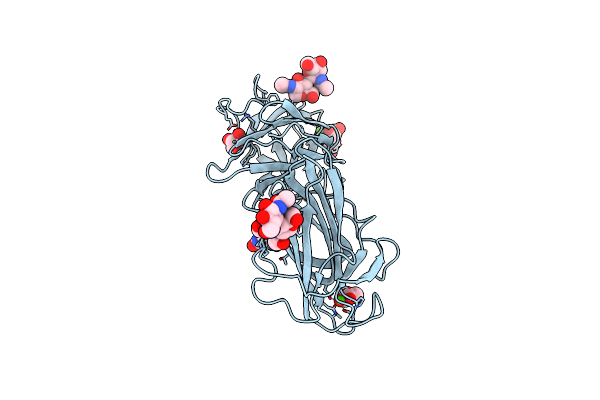 |
Crystal Structure Of Lectin From Pleurotus Ostreatus In Complex With Galactose
Organism: Pleurotus ostreatus
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2020-08-05 Classification: SUGAR BINDING PROTEIN Ligands: CA, GOL, GLA |
 |
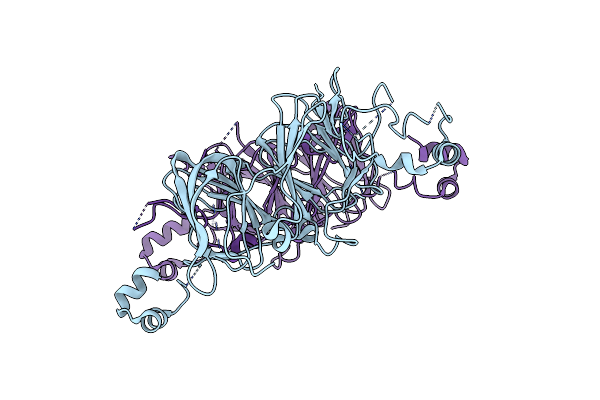
Crystal Structure Of Alocasin, Protease Inhibitor From Giant Taro (Arum Macrorrhizon)
Organism: Alocasia macrorrhizos
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2018-06-13 Classification: PLANT PROTEIN |
 |
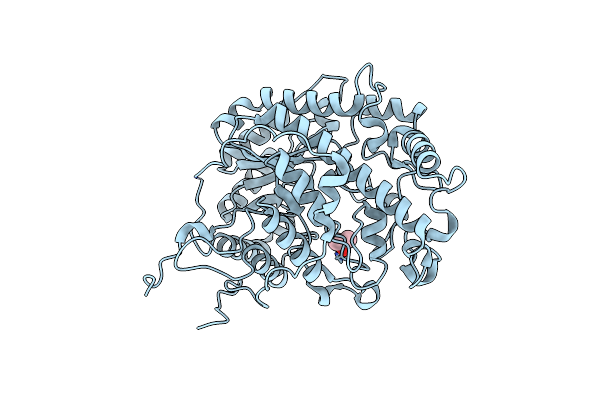
Organism: Cocos nucifera
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2018-06-06 Classification: ALLERGEN |
 |
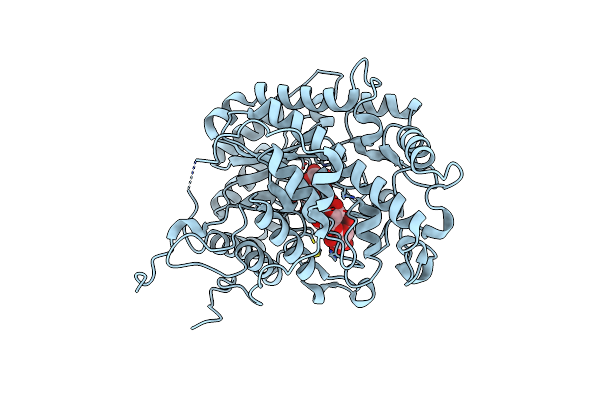
Organism: Ipomoea batatas
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2017-12-06 Classification: HYDROLASE Ligands: IPA |
 |
Crystal Structure Of Sweet Potato Beta-Amylase Complexed With Maltotetraose
Organism: Ipomoea batatas
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2017-12-06 Classification: HYDROLASE |
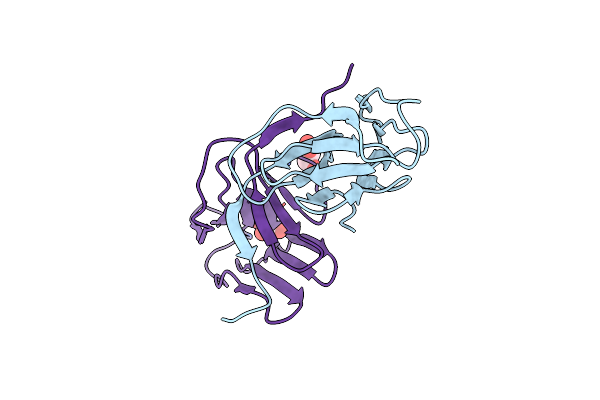 |
Organism: Colocasia esculenta
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2016-06-22 Classification: SUGAR BINDING PROTEIN Ligands: GOL |
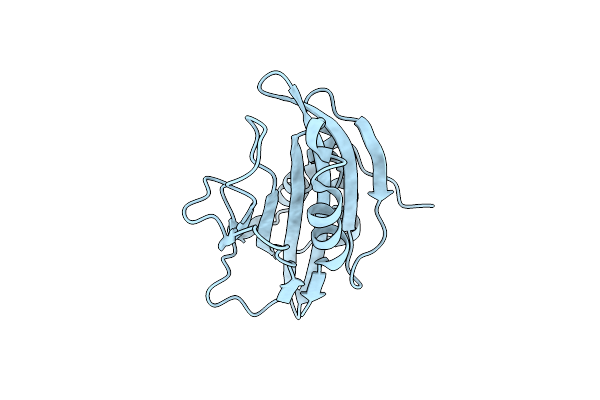 |
Crystal Structure Of Secreted Proline Rich Antigen Mtc28 (Rv0040C) At 2.15 A With Bound Chloride From Mycobacterium Tuberculosis
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2015-03-25 Classification: IMMUNE SYSTEM Ligands: CL |
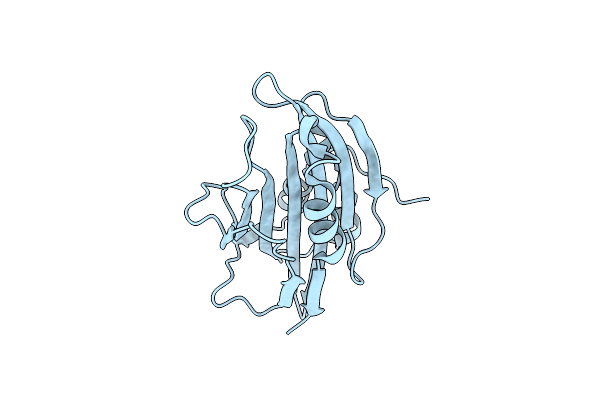 |
Crystal Structure Of Secreted Proline Rich Antigen Mtc28 (Rv0040C) From Mycobacterium Tuberculosis
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2015-01-28 Classification: LIPID BINDING PROTEIN |
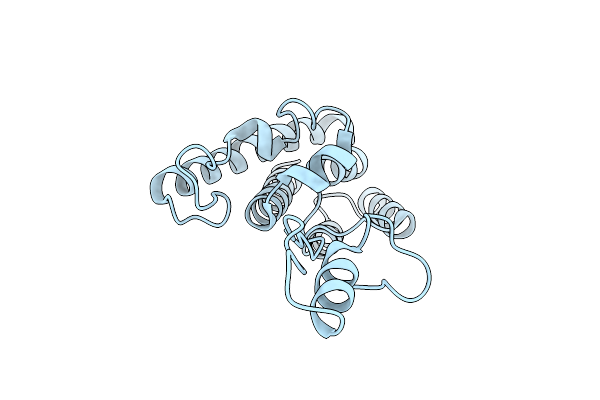 |
Dimeric Rous Sarcoma Virus Capsid Protein Structure With An Upstream 25-Amino Acid Residue Extension Of C-Terminal Of Gag P10 Protein
Organism: Rous sarcoma virus
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2003-12-23 Classification: VIRAL PROTEIN |
 |
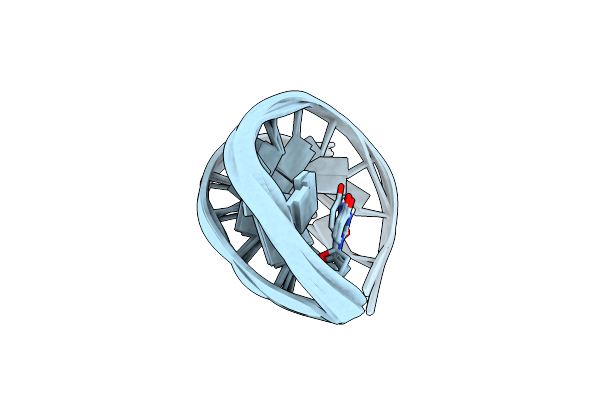
Organism: Paramecium bursaria chlorella virus 1
Method: ELECTRON MICROSCOPY Release Date: 2002-12-04 Classification: VIRUS |
 |

