Search Count: 2,211
All
Selected
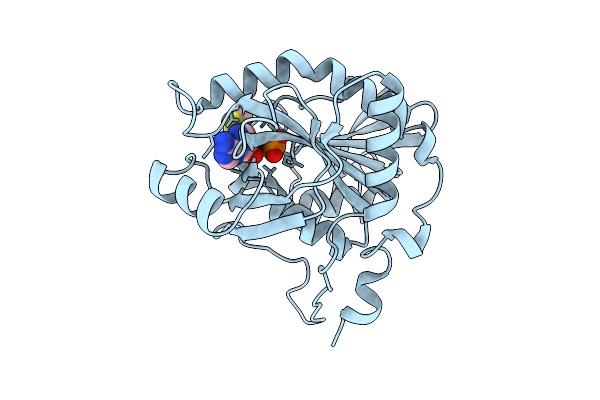 |
Crystal Structure Of Mtap From Aeropyrum Pernix Complex With Mta At 343K, Soaked In Potassium Phosphate Ph7.0 With Mta
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: PO4, MTA |
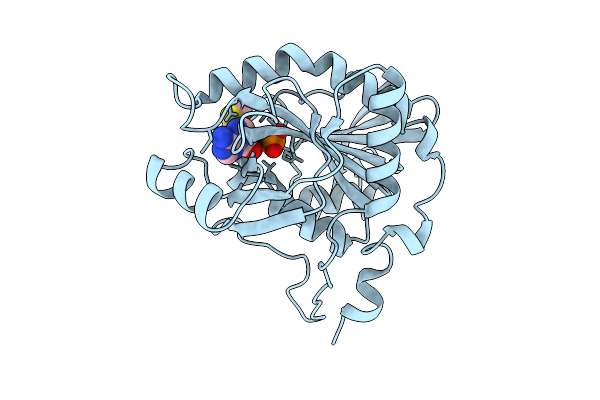 |
Crystal Structure Of Mtap From Aeropyrum Pernix Complex With Mta At 323K, Soaked In Potassium Phosphate Ph7.0 With Mta
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: PO4, MTA |
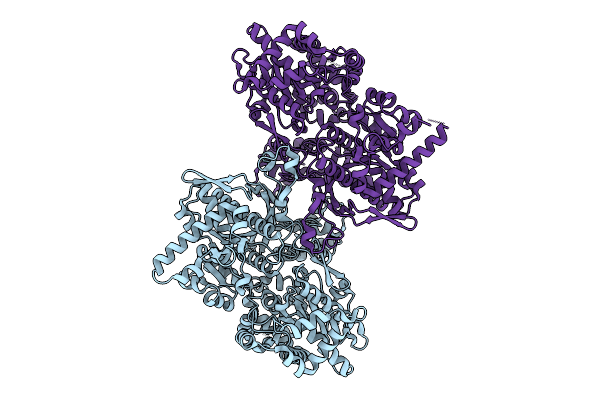 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
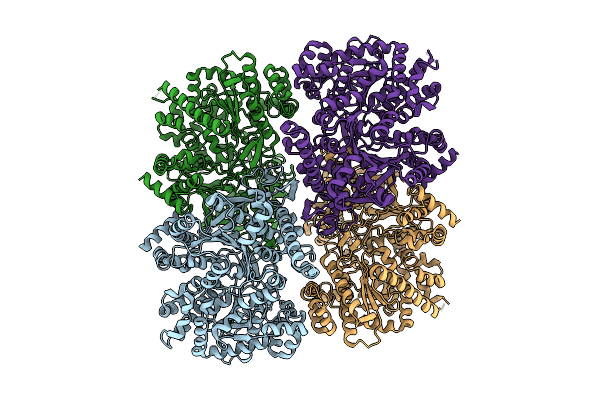 |
Structure Of Glycogen Phosphorylase - Tetrameric Form - From Escherichia Coli
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.97 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
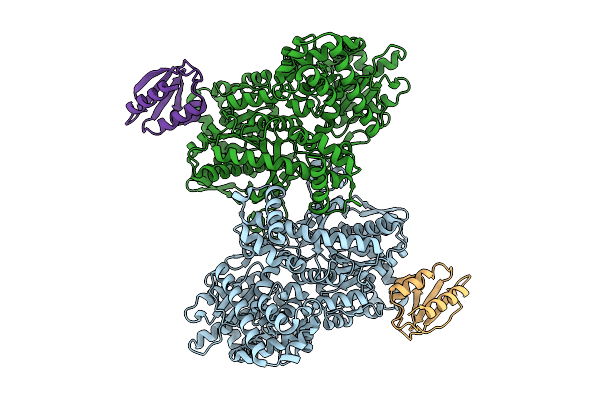 |
Structure Of Glycogen Phosphorylase - Dimeric Form - In Complex With Hpr From Escherichia Coli
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.79 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
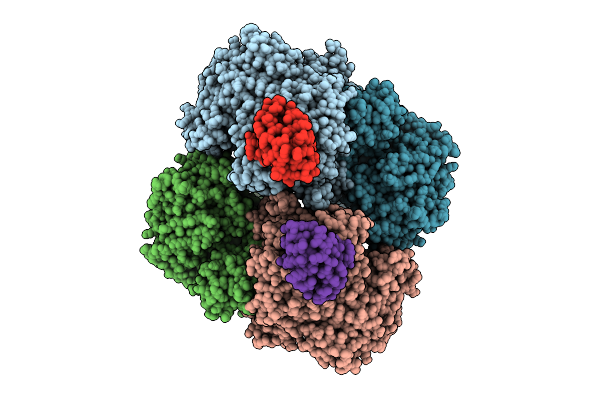 |
Structure Of Glycogen Phosphorylase - Tetrameric Form - In Complex With Hpr From Escherichia Coli
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.06 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
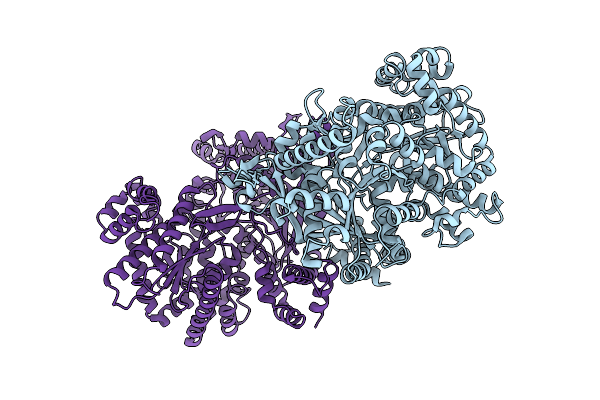 |
Organism: Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: STRUCTURAL PROTEIN |
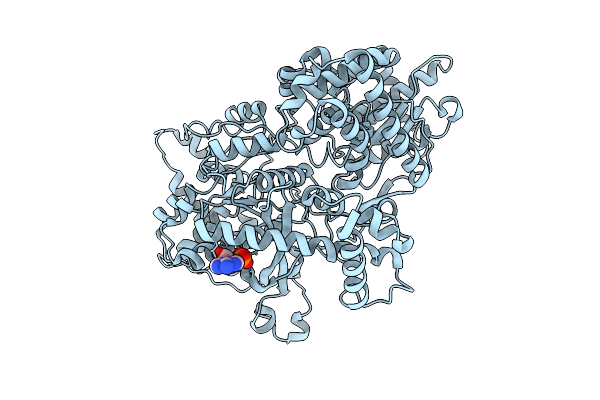 |
Organism: Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: STRUCTURAL PROTEIN Ligands: AMP |
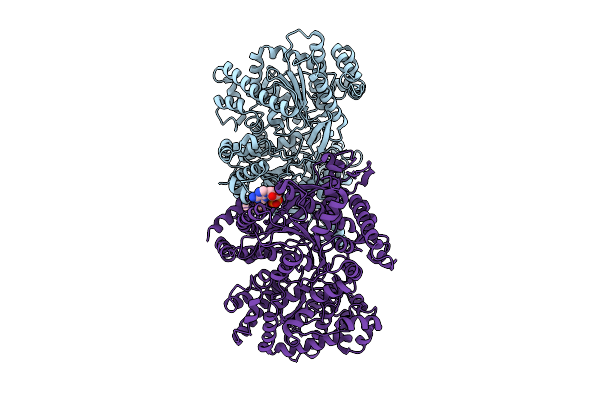 |
Organism: Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: STRUCTURAL PROTEIN Ligands: AMP |
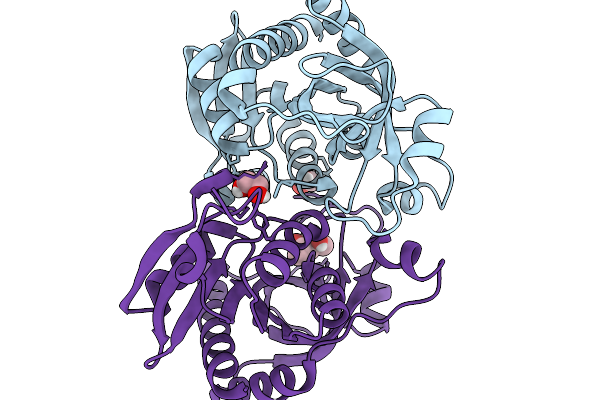 |
Wild-Type Apo Purine Nucleoside Phosphorylase From Parageobacillus Thermoglucosidasius
Organism: Parageobacillus thermoglucosidasius
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-03-25 Classification: TRANSFERASE Ligands: EDO, CL, MPD |
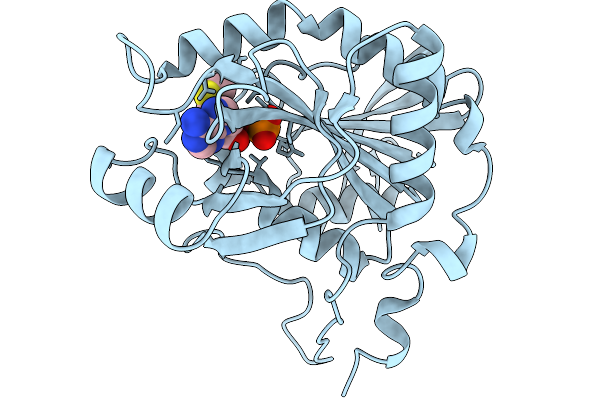 |
Crystal Structure Of Mtap From Aeropyrum Pernix Complex With Mta At 333K, Soaked In Potassium Phosphate Ph7.0 With Mta
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2026-03-25 Classification: TRANSFERASE Ligands: PO4, MTA |
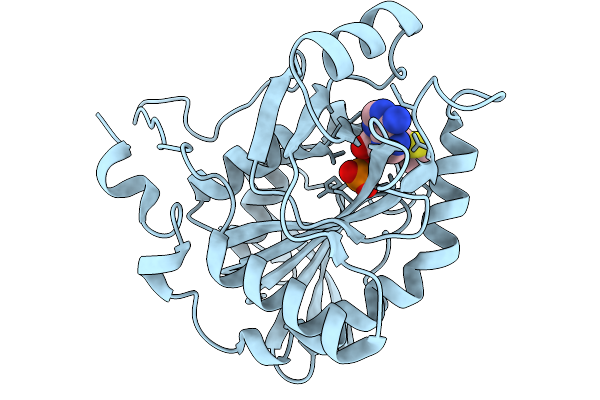 |
Crystal Structure Of Mtap From Aeropyrum Pernix Complex With Mta At 293K, Soaked In Potassium Phosphate Ph7.0 With Mta
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2026-03-25 Classification: TRANSFERASE Ligands: PO4, MTA |
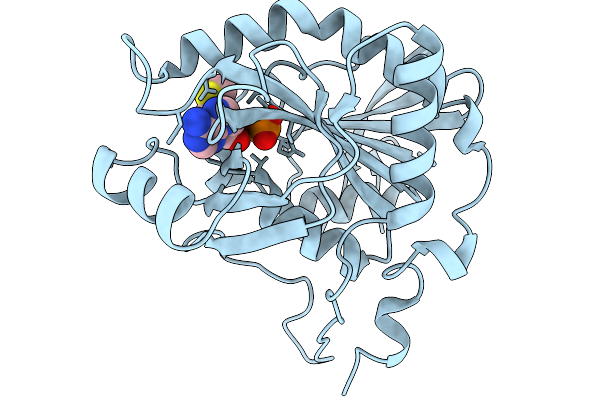 |
Crystal Structure Of 5'-Deoxy-5'-Methylthioadenosine Phosphorylase From Aeropyrum Pernix Complex With 5'-Deoxy-5'-Methylthioadenosine 353K 1.77 Angstrom
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2026-03-25 Classification: TRANSFERASE Ligands: PO4, MTA |
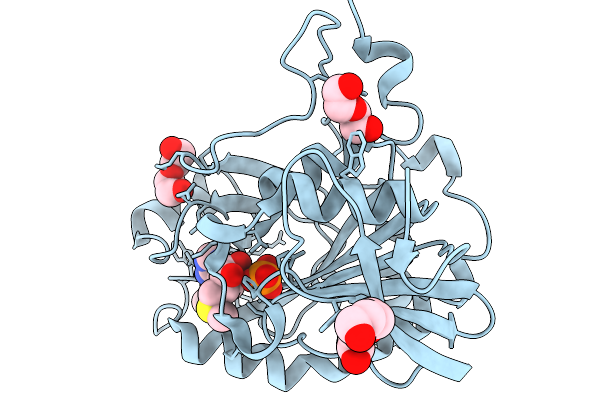 |
Crystal Structure Of Mtap From Aeropyrum Pernix Complex With Mta At 100K, Soaked In Potassium Phosphate Ph7.0 With Mta
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.19 Å Release Date: 2026-03-25 Classification: TRANSFERASE Ligands: PO4, MTA, PEG, PGE |
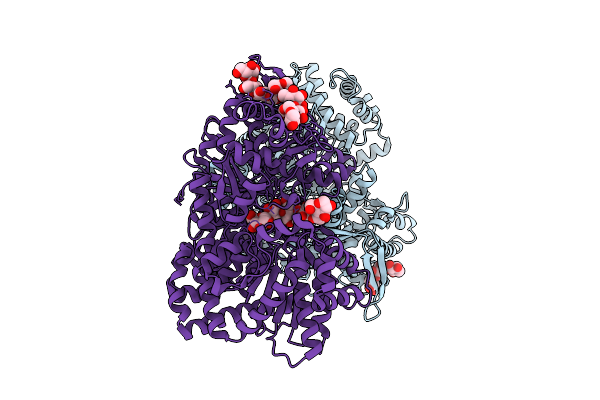 |
Carbohydrate-Bound Structure Of Alpha-Glucan Phosphorylase From Crocosphaera Subtropica Atcc 51142
Organism: Crocosphaera subtropica atcc 51142
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-03-18 Classification: TRANSFERASE |
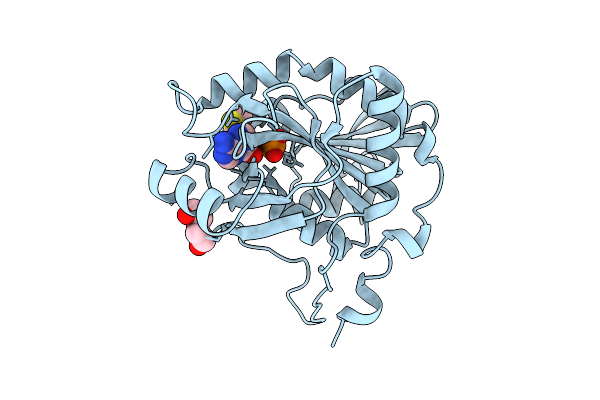 |
Crystal Structure Of 5'-Deoxy-5'-Methylthioadenosine Phosphorylase From Aeropyrum Pernix Complex With 5'-Deoxy-5'-Methylthioadenosine 343K 1.58 Angstrom
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-03-11 Classification: TRANSFERASE Ligands: PO4, MTA, PEG |
 |
Crystal Structure Of 5'-Deoxy-5'-Methylthioadenosine Phosphorylase From Aeropyrum Pernix Complex With 5'-Deoxy-5'-Methylthioadenosine 100K
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2026-03-04 Classification: TRANSFERASE Ligands: PO4, MTA, PEG, PGE |
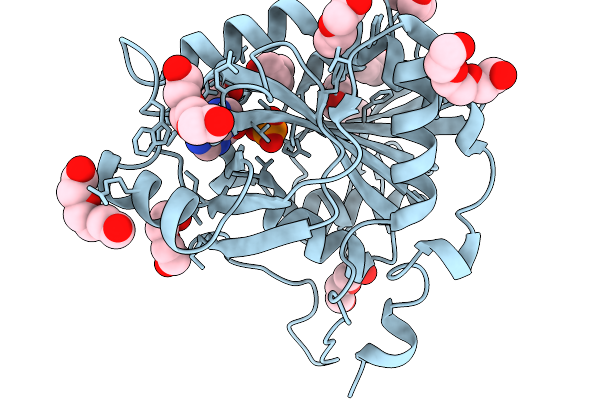 |
Crystal Structure Of 5'-Deoxy-5'-Methylthioadenosine Phosphorylase From Aeropyrum Pernix 100K
Organism: Aeropyrum pernix k1
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2026-03-04 Classification: TRANSFERASE Ligands: PO4, ADE, PGE, PEG |
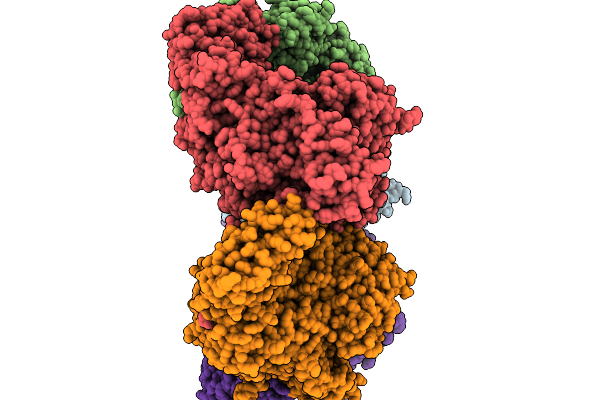 |
Organism: Segatella copri
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-02-18 Classification: TRANSFERASE |
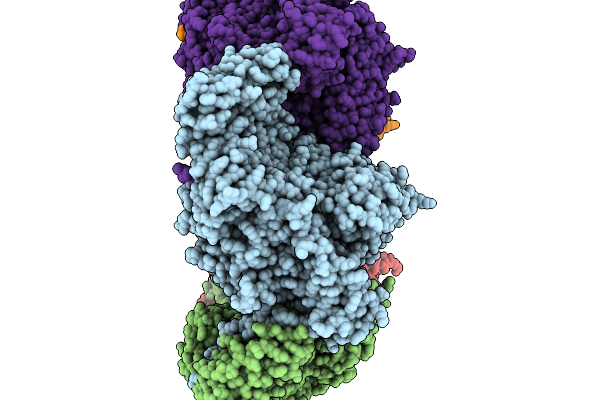 |
R583A Mutant Of Glycogen Phosphorylase From Segatella Copri In The Presence Of Amp
Organism: Segatella copri
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2026-02-18 Classification: TRANSFERASE |

