Search Count: 70,899
All
Selected
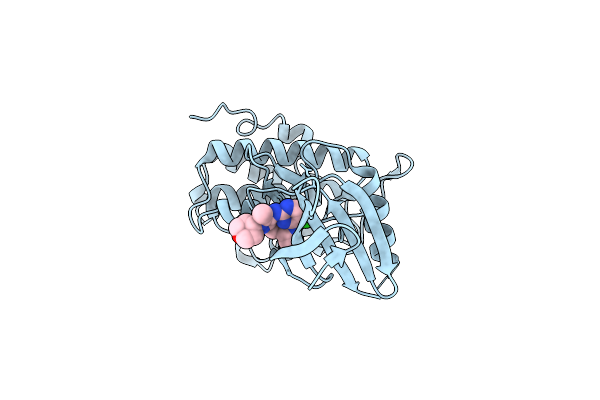 |
Structure Of Chk1 10-Pt. Mutant Complex With Macrocyclic Lrrk2 Inhibitor Compound 1 ((11R)-8-Chloro-3,11-Dimethyl-2-(Oxan-4-Yl)-2,4,10,11,12,13-Hexahydro-9,5-(Azeno)Pyrazolo[3,4-B][1,4,6,10]Oxatriazacyclotridecine)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2026-04-08 Classification: TRANSFERASE/INHIBITOR Ligands: A1C5F |
 |
Structure Of Chk1 10-Pt. Mutant Complex With Macrocyclic Lrrk2 Inhibitor Compound 7 ((10As,13As)-3-Cyclobutyl-1-Methyl-8-(Trifluoromethyl)-3,4,10A,11,13A,14-Hexahydro-10H,13H-9,5-(Azeno)Furo[3,4-K]Pyrazolo[4,3-B][1,4,6,10]Oxatriazacyclotridecine)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2026-04-08 Classification: TRANSFERASE/INHIBITOR Ligands: A1C5G |
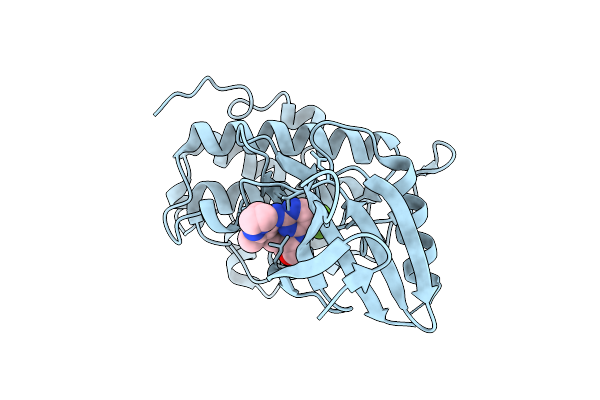 |
Structure Of Chk1 10-Pt. Mutant Complex With Macrocyclic Lrrk2 Inhibitor Compound 12 ((10As,13As)-3-Cyclopropyl-1-Methyl-8-(Trifluoromethyl)-3,4,10A,11,13A,14-Hexahydro-10H,13H-9,5-(Azeno)Furo[3,4-K]Pyrazolo[4,3-B][1,4,6,10]Oxatriazacyclotridecine)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-04-08 Classification: TRANSFERASE/INHIBITOR Ligands: A1C5H |
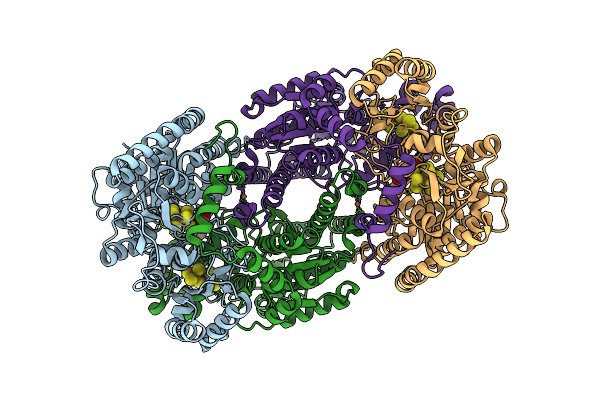 |
Azotobacter Vinelandii Mofep (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0X Surfact
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Resolution:2.69 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: HCA, ICS, CLF, FE |
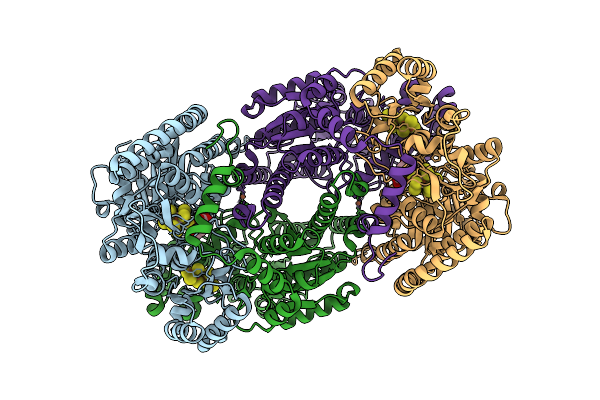 |
Azotobacter Vinelandii Mofep (C2 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0X Surfact
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Resolution:2.46 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: HCA, ICS, FE, CLF |
 |
Azotobacter Vinelandii Mofep (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Resolution:2.16 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: HCA, ICS, FE, CLF |
 |
Azotobacter Vinelandii Mofep (C2 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Resolution:2.06 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: HCA, ICS, FE, CLF |
 |
Azotobacter Vinelandii Mofep (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 1X Surfact
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Resolution:2.49 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: HCA, ICS, FE, CLF |
 |
Azotobacter Vinelandii Mofep (C2 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 1X Surfact
Organism: Azotobacter vinelandii
Method: ELECTRON MICROSCOPY Resolution:2.34 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: HCA, ICS, FE, CLF |
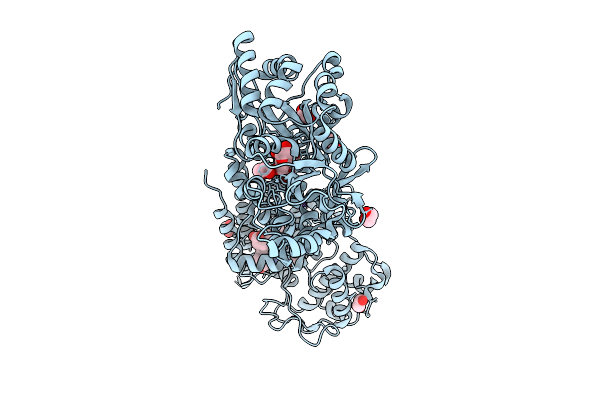 |
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) H143Y Mutant Complexed With 5-Methoxy-L-Trp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: A1A2Y, EDO, HEM, CYN, NA |
 |
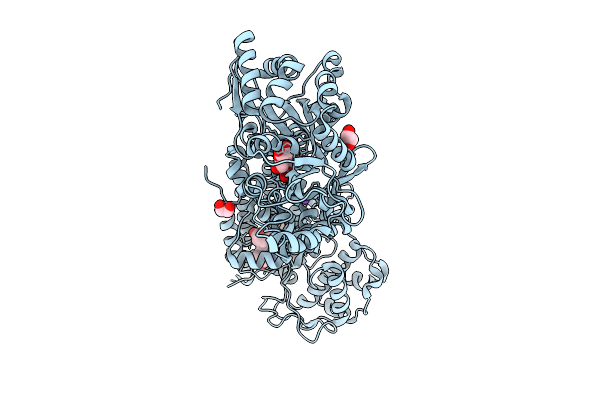
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) Complexed With 5-Hydroxy-L-Trp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: EDO, HEM, CYN, 4PQ, NA |
 |
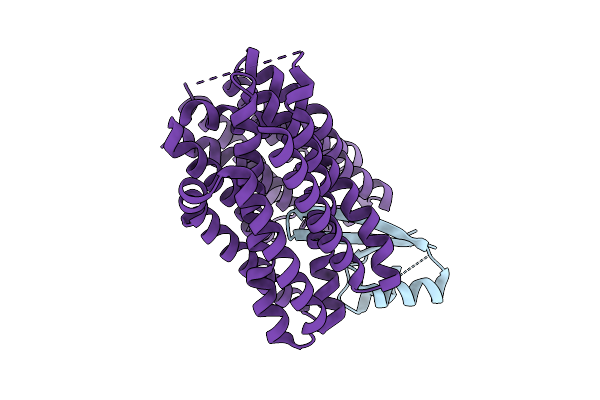
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) H143Y Mutant Complexed With 5-Methyl-L-Trp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: EDO, HEM, CYN, D0Q, NA |
 |
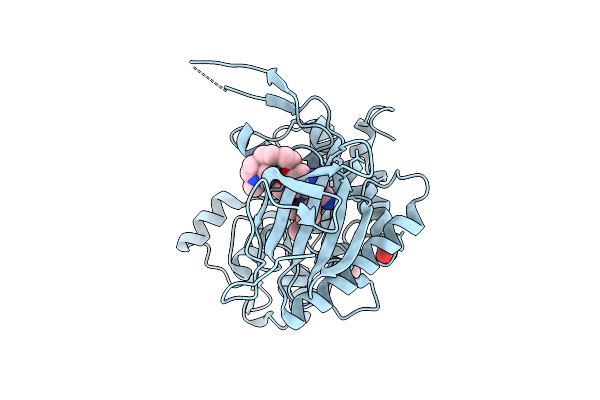
Organism: Synthetic construct, Staphylococcus aureus, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSPORT PROTEIN |
 |
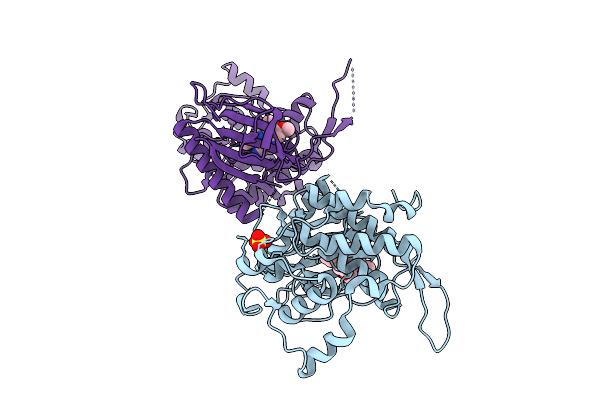
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: EDO, A1J3A, IOD |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: SO4, A1J3B |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.03 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN,HYDROLASE/INHIBITOR Ligands: A1A5Z, CL |
 |
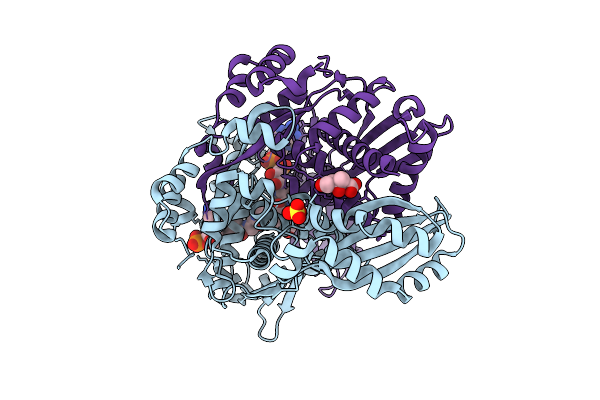
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN,HYDROLASE/INHIBITOR Ligands: A1A54, CL |
 |
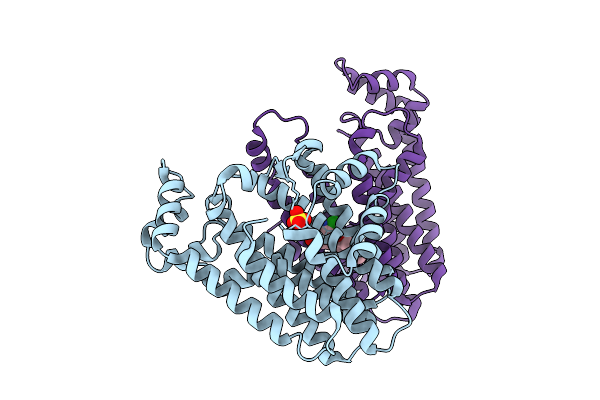
Structure Of Pmhmgr Bound To Mevalonate, Coa And Nad 2 Minutes 20 Seconds After Reaction Initiation At Ph 9
Organism: Pseudomonas sp. 'mevalonii'
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE/SUBSTRATE/INTERMEDIATE Ligands: COA, SO4, A1AFY, ZKE, NAI, MEV |
 |
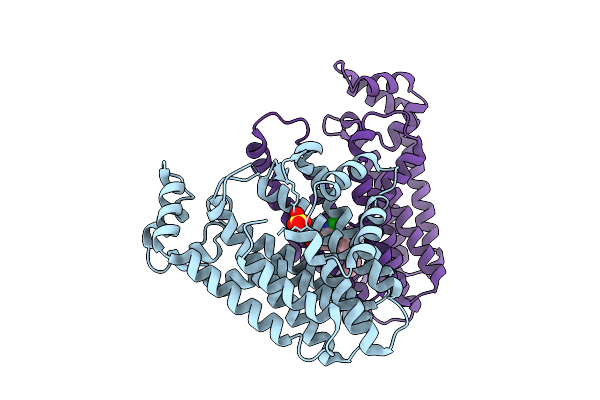
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: SO4, A1EC1 |
 |
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: SO4, A1EC2 |

