Search Count: 161
All
Selected
 |
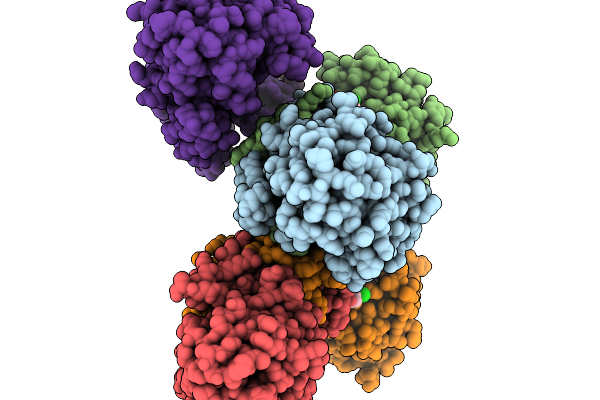
Structure Of Pycr1 Complexed With The Allosteric Inhibitor 1-Benzyl-1H-1,3-Benzodiazole-2-Sulfonic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: A1CFK, SO4 |
 |
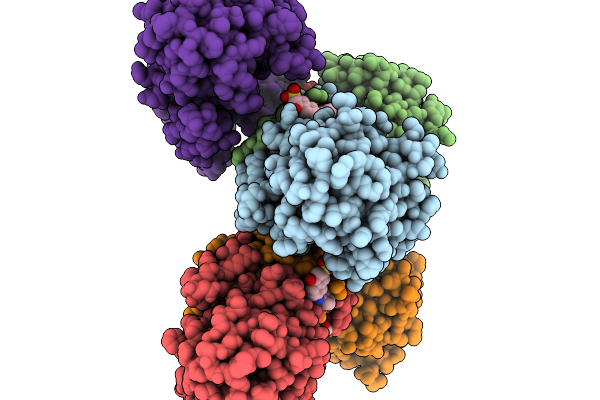
Structure Of Pycr1 Complexed With The Allosteric Inhibitor 2-[(2,6-Dichlorophenyl)Amino]Pyridine-3-Sulfonic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE Ligands: A1CFJ, SO4, PEG, PGE |
 |
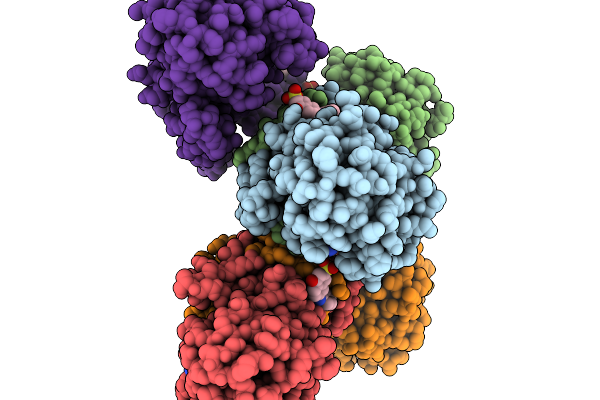
Structure Of Pycr1 Complexed With The Allosteric Inhibitor 4-Hydroxy-7-(Phenylamino)Naphthalene-2-Sulfonic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: A1CFI, SO4, PGE |
 |
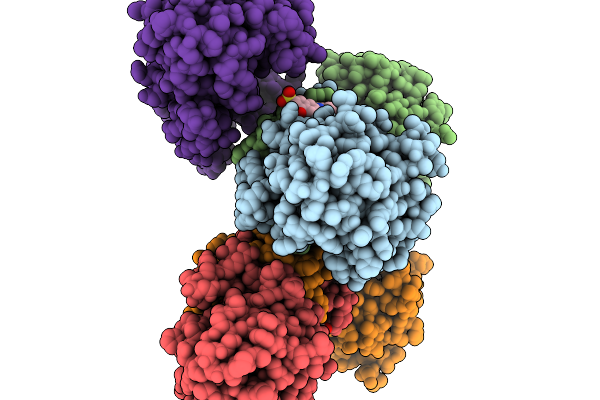
Structure Of Pycr1 Complexed With The Allosteric Inhibitor 4-Hydroxy-7-(Phenylamino)Naphthalene-2-Sulfonic Acid In A Remote Site And Nadh And (S)-(-)-2-Hydroxy-3,3-Dimethylbutyric Acid In The Active Site
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: G8I, A1CFI, NAI, PEG |
 |
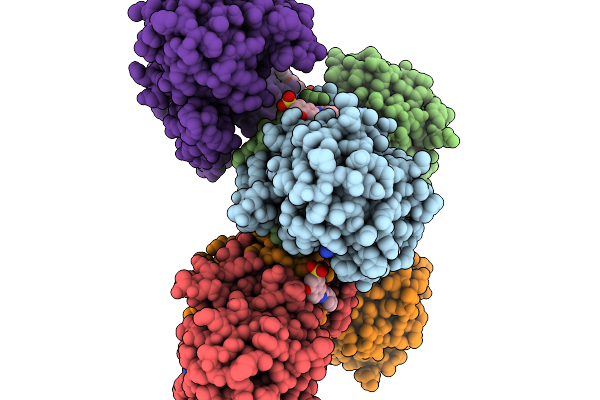
Structure Of Pycr1 Complexed With The Allosteric Inhibitor 7-Benzamido-4-Hydroxynaphthalene-2-Sulfonic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: SO4, PEG, A1CFH, PGE |
 |
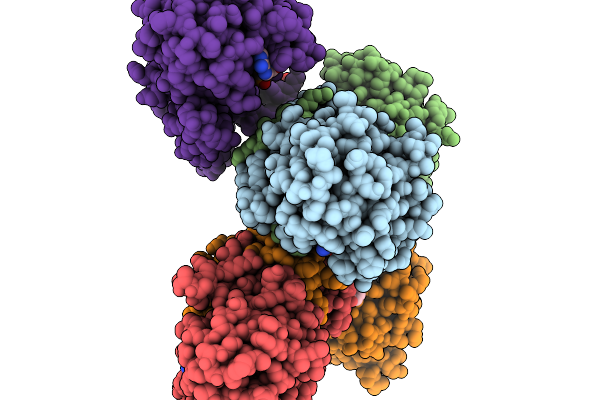
Structure Of Pycr1 Complexed With The Allosteric Inhibitor 7-Benzamido-4-Hydroxynaphthalene-2-Sulfonic Acid In A Remote Site And Nadh And (S)-(-)-2-Hydroxy-3,3-Dimethylbutyric Acid In The Active Site
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: G8I, A1CFH, NAI |
 |
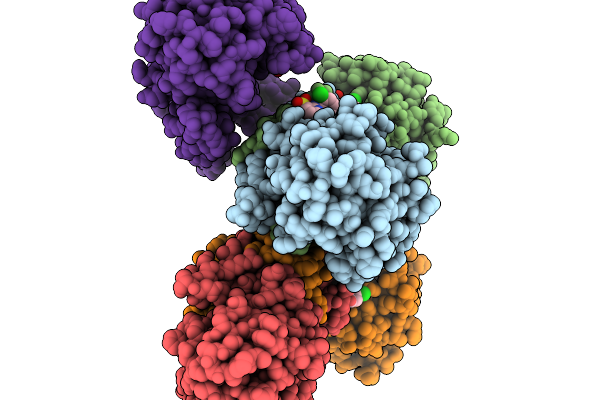
Quaternary Complex Of Pycr1 With The Allosteric Inhibitor 1-Benzyl-1H-1,3-Benzodiazole-2-Sulfonic Acid In A Remote Site And Nadh And (S)-Tetrahydrofuran-2-Carboxylic Acid In The Active Site
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: TFB, NAI, A1CFK, PEG |
 |
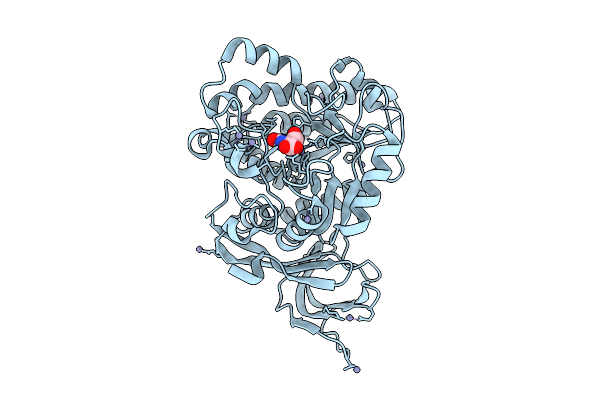
Ternary Complex Of Pycr1 With The Allosteric Inhibitor 2-[(2,6-Dichlorophenyl)Amino]Pyridine-3-Sulfonic Acid In Two Remote Sites And The Active Site, And (S)-(-)-2-Hydroxy-3,3-Dimethylbutyric Acid In The Active Site
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: G8I, A1CFJ, SO4 |
 |
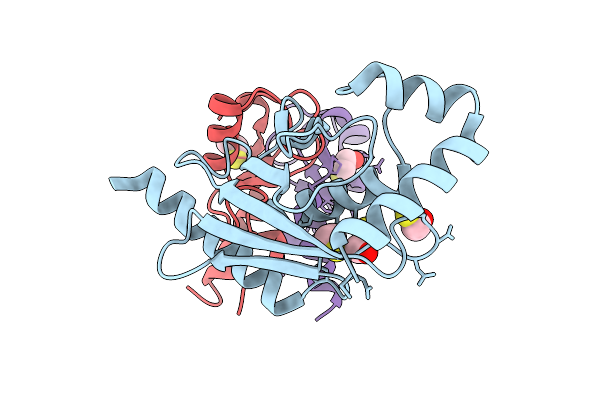
Organism: Methanocaldococcus jannaschii
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2025-12-17 Classification: HYDROLASE Ligands: NCD, ZN |
 |
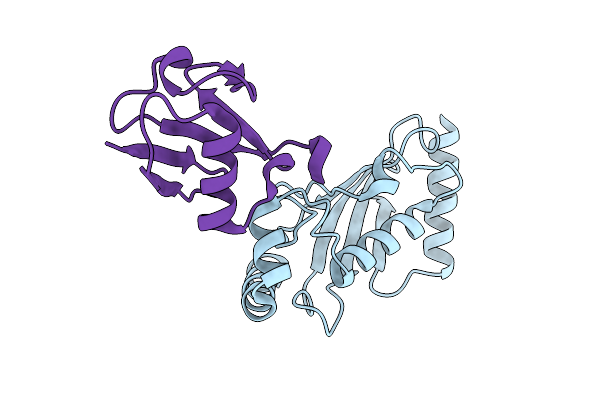
Structure Of Ubch5B In Complex With The U-Box Domain Of The E3 Ubiquitin Ligase Chip
Organism: Homo sapiens, Danio rerio
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2025-11-05 Classification: LIGASE Ligands: BME, CL |
 |
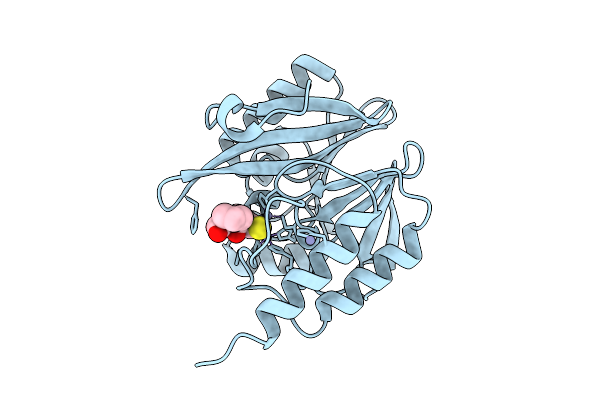
Structure Of The Isopeptide Bond-Linked Ubch5B~Ubiquitin Conjugate Complex For An M1K/C85K Ubch5B Mutant
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2025-11-05 Classification: LIGASE |
 |
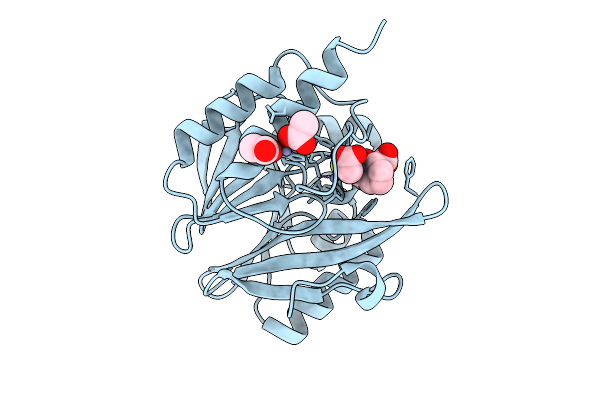
Organism: Enterobacter cloacae
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2025-07-09 Classification: HYDROLASE Ligands: ZN, MCO |
 |
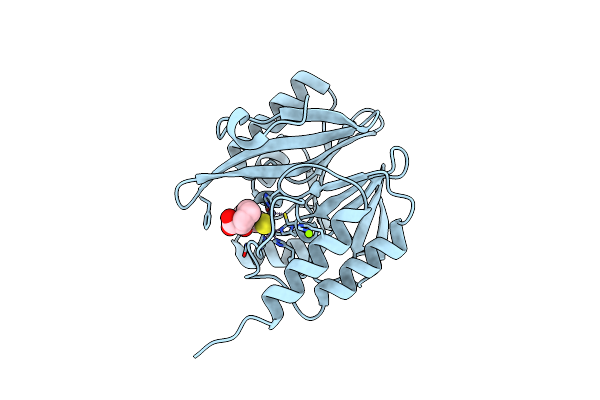
Organism: Enterobacter cloacae
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2025-07-09 Classification: HYDROLASE Ligands: ZN, X8Z, ACT |
 |
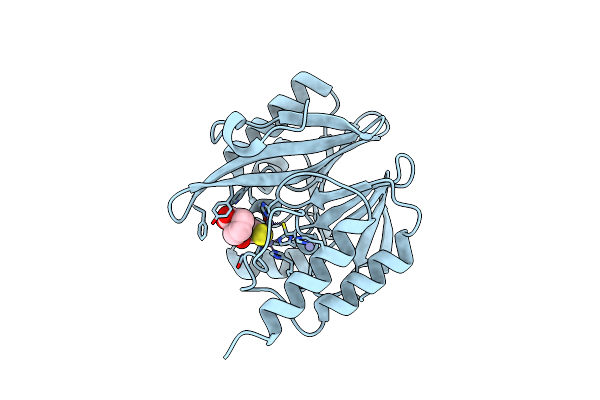
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.29 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: ZN, X8Z, MG |
 |
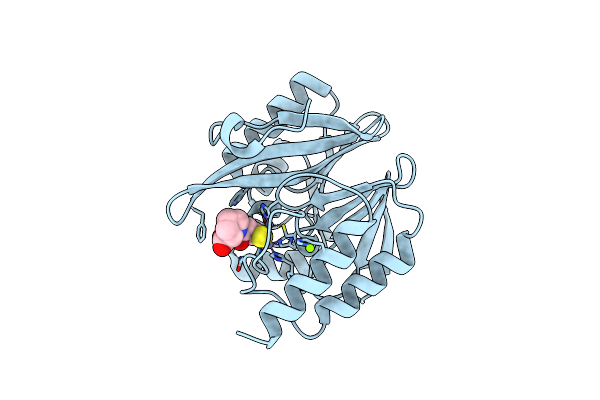
Organism: Enterobacter cloacae
Method: X-RAY DIFFRACTION Resolution:1.36 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: ZN, MCO |
 |
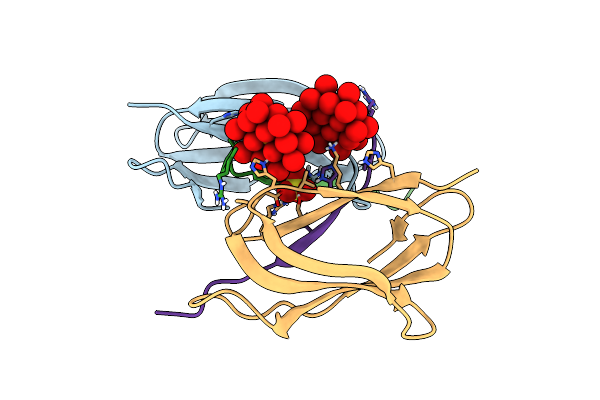
Organism: Enterobacter cloacae
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: ZN, X8Z, MG |
 |
Organism: Streptococcus pyogenes
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2025-06-25 Classification: PEPTIDE BINDING PROTEIN Ligands: TEW, SO4 |
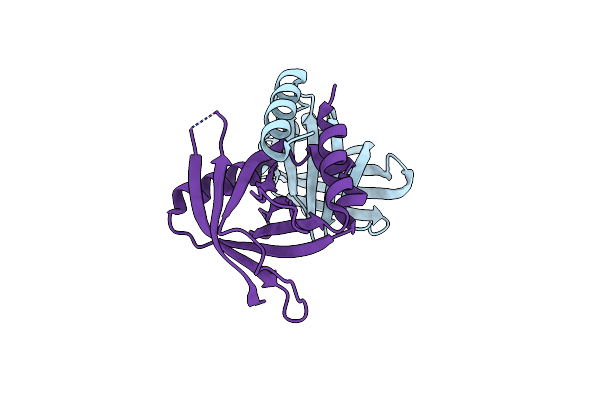 |
Crystal Structure Of Human Respiratory Syncytial Virus Ns1 Bound To Human Med25 Acid
Organism: Human respiratory syncytial virus a, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.53 Å Release Date: 2025-04-09 Classification: VIRAL PROTEIN |
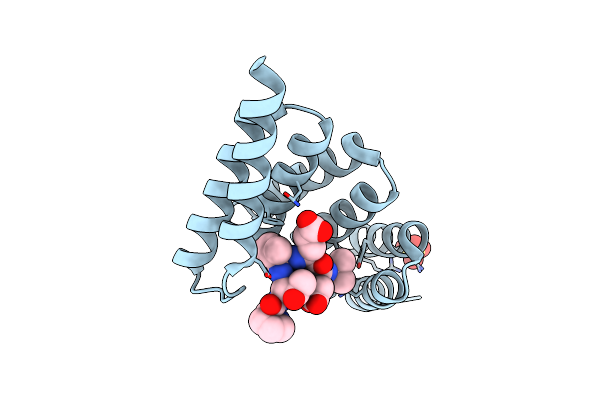 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2025-03-26 Classification: LIGASE Ligands: SO4 |
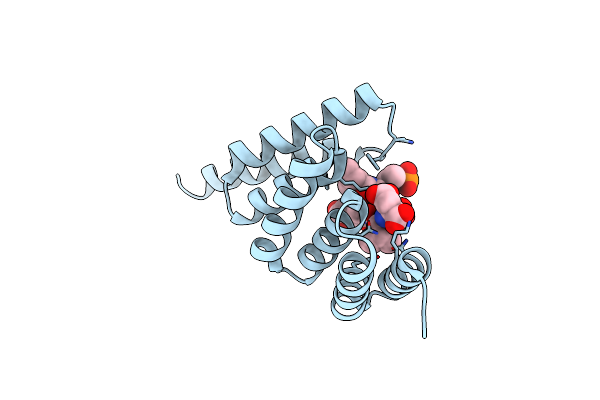 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-03-26 Classification: LIGASE Ligands: CL |

