Search Count: 347
All
Selected
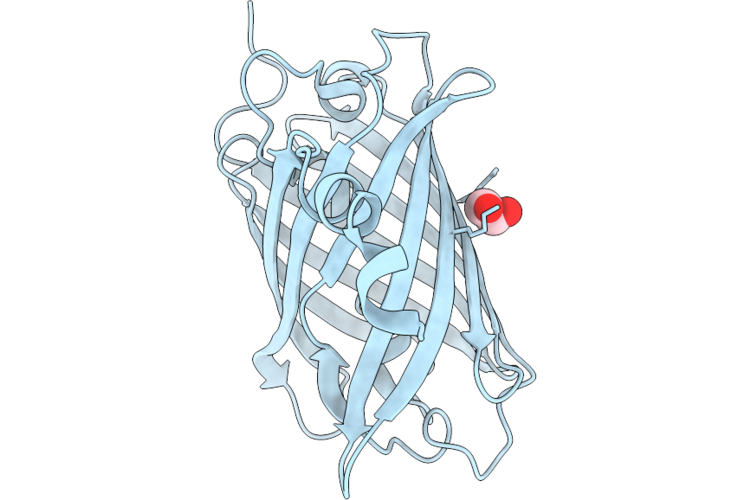 |
Organism: Discosoma sp.
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-22 Classification: FLUORESCENT PROTEIN Ligands: EDO |
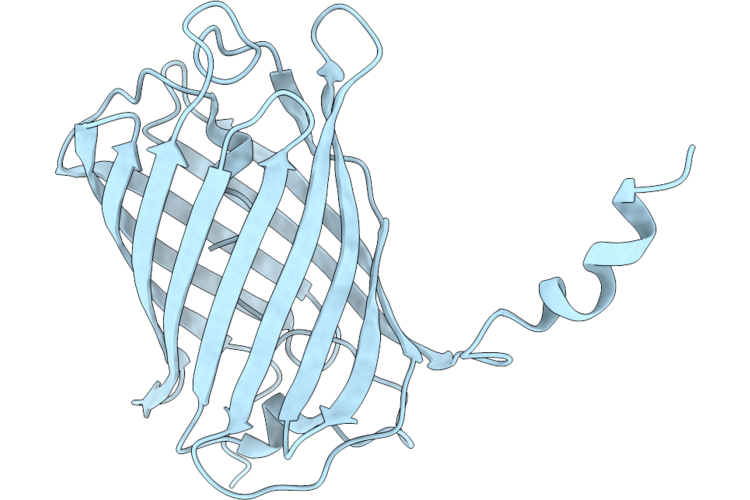 |
Organism: Discosoma sp.
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-22 Classification: FLUORESCENT PROTEIN |
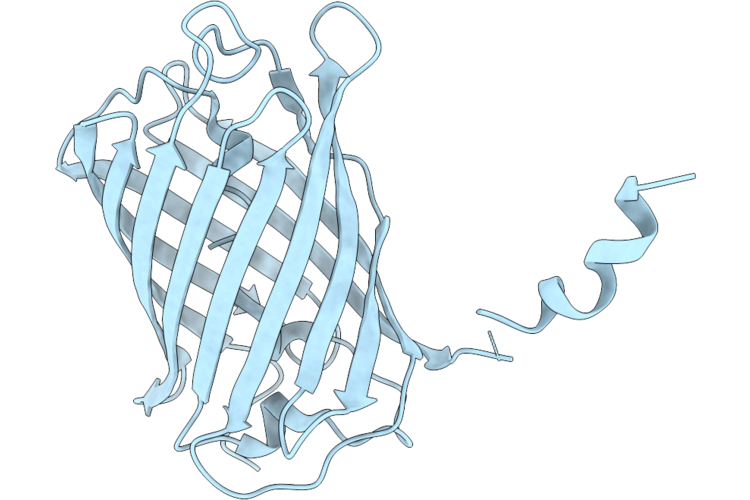 |
Crystal Structure Of Lssmorange Under Room Temperature At Ph 8.0, 250Ps After Pump Laser, Refined Against 8.33 Times Extrapolated Structure Factor
Organism: Discosoma sp.
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: FLUORESCENT PROTEIN |
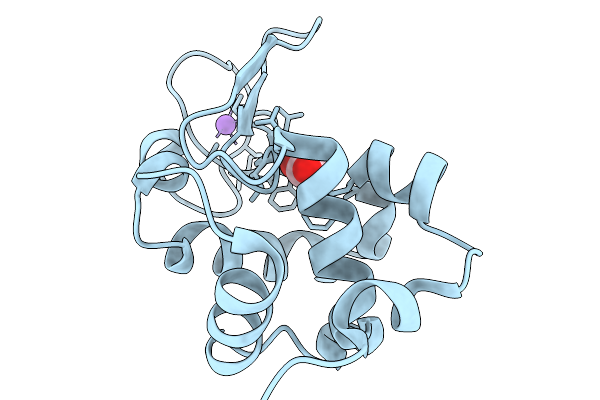 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-03-25 Classification: HYDROLASE |
 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-03-25 Classification: HYDROLASE |
 |
Time-Resolved Mixing Crystallography Structure Of 3-5 Micrometers Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 2 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 3-5 Micrometers Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 5 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 3-5 Micrometers Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 9.7 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 1.3 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 5.0 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 7.5 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 226 Mm N-Acetyl-D-Glucosamine At 9.7 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 452 Mm N-Acetyl-D-Glucosamine At 1.3 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 452 Mm N-Acetyl-D-Glucosamine At 2.5 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 452 Mm N-Acetyl-D-Glucosamine At 4 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Time-Resolved Mixing Crystallography Structure Of 1 Micrometer Lysozyme Microcrystals With 452 Mm N-Acetyl-D-Glucosamine At 5.0 Seconds Delay
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: CL, NA, NDG |
 |
Crystal Structure Of Hen Egg-White Lysozyme Determined In The Commissioning Of Nanoterasu Mx-Es
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2026-03-11 Classification: HYDROLASE Ligands: CL, NA |
 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured By Mix-And-Inject Serial Crystallography At 100-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: PEA, NA, CU |
 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured By Mix-And-Inject Serial Crystallography At 200-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: PEA, NA, CU |
 |
Reductive-Half Reaction Intermediate Of Copper Amine Oxidase From Arthrobacter Globiformis Captured By Mix-And-Inject Serial Crystallography At 300-Ms Time Delay
Organism: Arthrobacter globiformis
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE |

