Search Count: 19
All
Selected
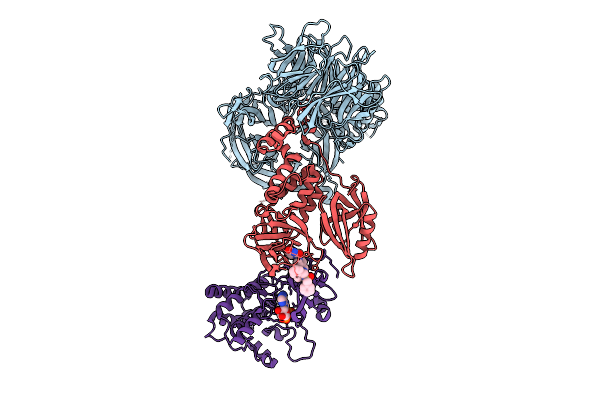 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.25 Å Release Date: 2025-07-09 Classification: DNA Binding Protein/Transferase Ligands: A1BX6, ZN, ADP, MG |
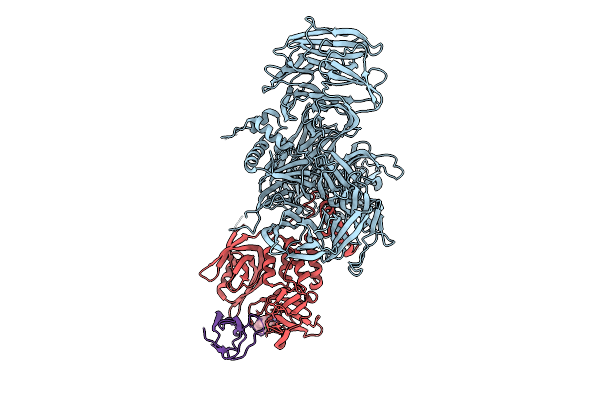 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.40 Å Release Date: 2025-07-09 Classification: LIGASE Ligands: ZN, A1BYX |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.95 Å Release Date: 2025-07-09 Classification: DNA Binding Protein/Transferase Ligands: ZN, A1BC8 |
 |
A Degradation Product Of Pd 404182 (P2742) Bound To Histone Deacetylase-Like Amidohydrolase
Organism: Alcaligenaceae bacterium fb188
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2019-01-16 Classification: HYDROLASE Ligands: ZN, K, 1PE, PEG, F0Z, ACT, MLT |
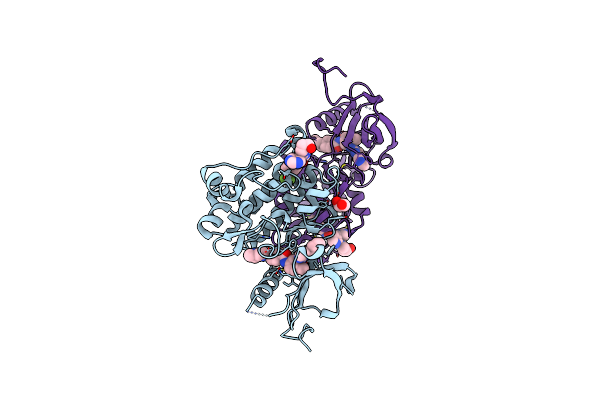 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2018-10-24 Classification: TRANSFERASE Ligands: GOL, CL, FYW, STI |
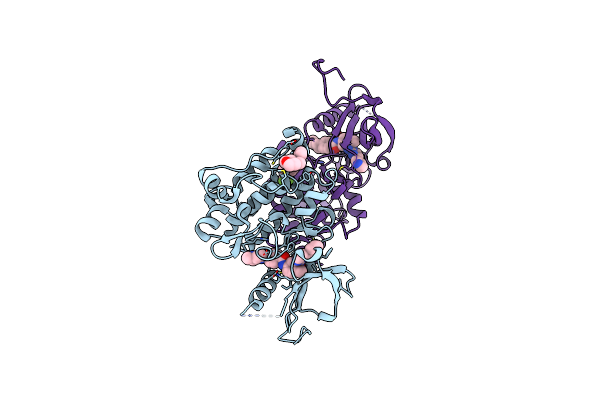 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2018-09-12 Classification: TRANSFERASE Ligands: CL, FYH, STI |
 |
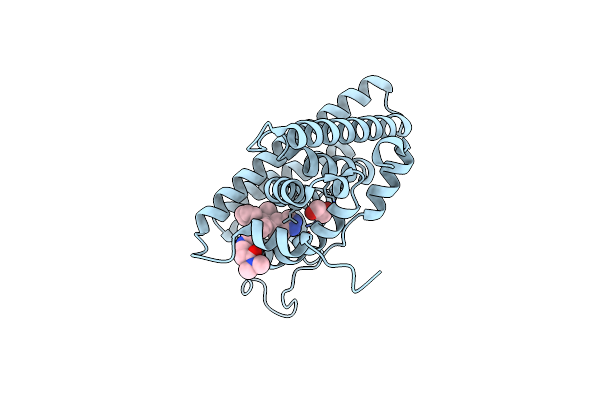
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2018-03-21 Classification: nuclear protein/antagonist Ligands: F3D, EDO, DMS |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2018-03-21 Classification: nuclear protein/antagonist Ligands: F3D, EDO |
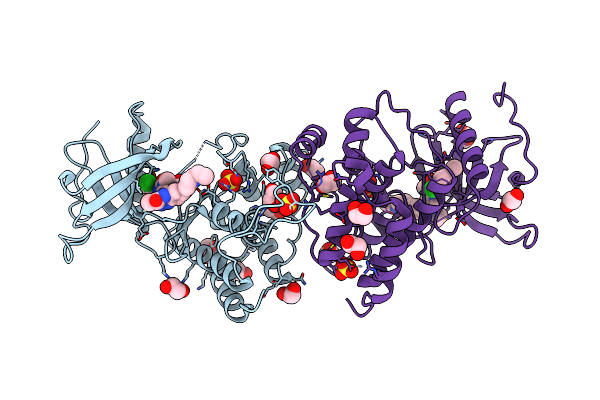 |
Crystal Structure Of Fgfr1-Y563C (Fgfr4 Surrogate) Covalently Bound To H3B-6527
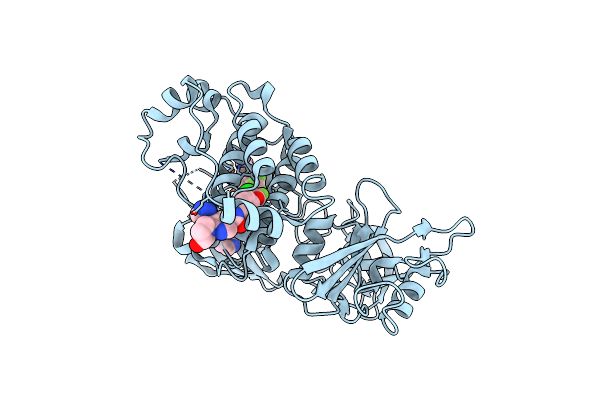
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2017-05-24 Classification: transferase/transferase inhibitor Ligands: 9ES, SO4, EDO |
 |
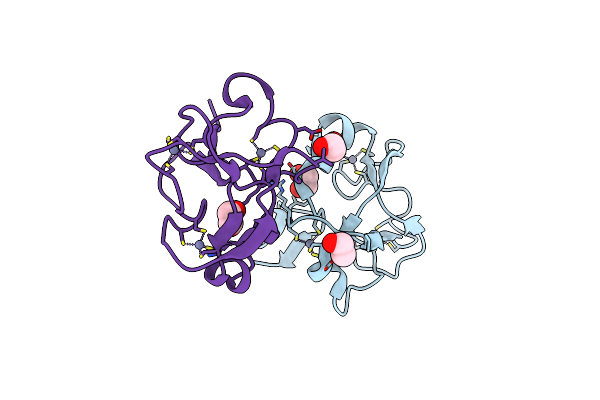
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.17 Å Release Date: 2017-04-05 Classification: TRANSFERASE Ligands: NIL, AY7 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2016-09-07 Classification: TRANSCRIPTION Ligands: ZN, EDO |
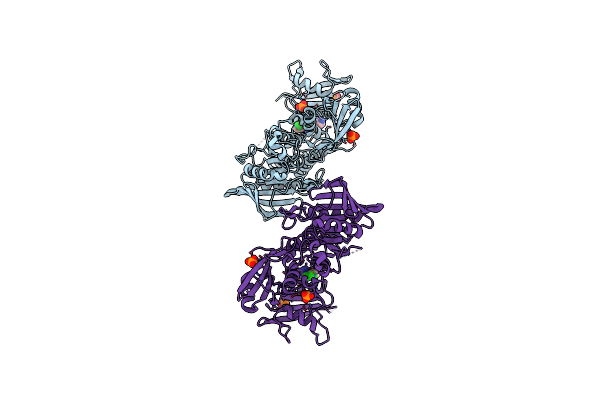 |
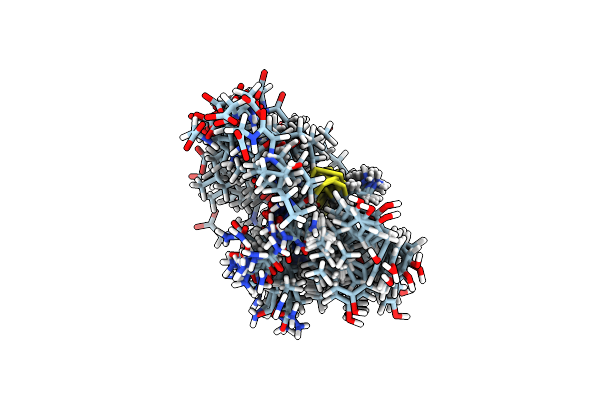
Non-Receptor Protein Tyrosine Phosphatase Shp2 In Complex With Allosteric Inhibitor Shp099
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2016-06-29 Classification: Hydrolase/Hydrolase Inhibitor Ligands: 5OD, PO4 |
 |
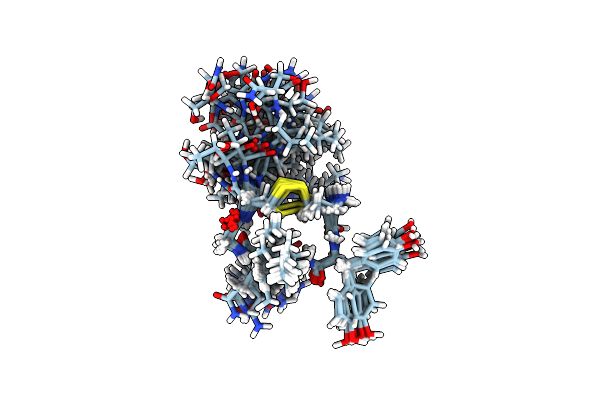 |
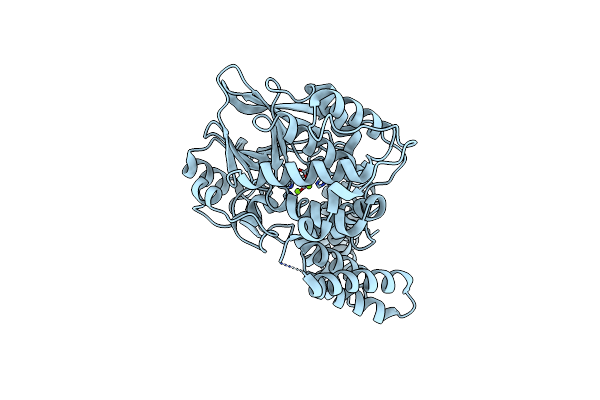 |
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2014-02-19 Classification: HYDROLASE Ligands: MG |
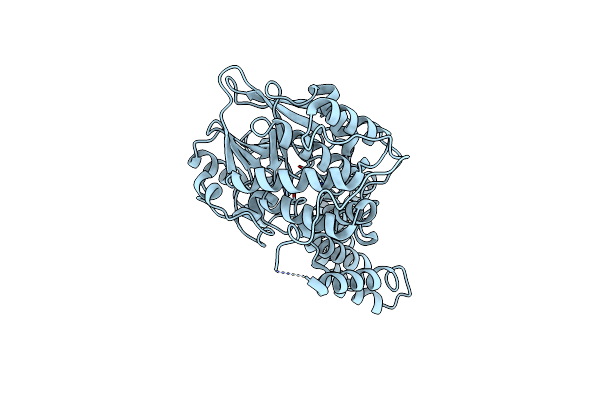 |
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2014-02-19 Classification: HYDROLASE Ligands: MG |
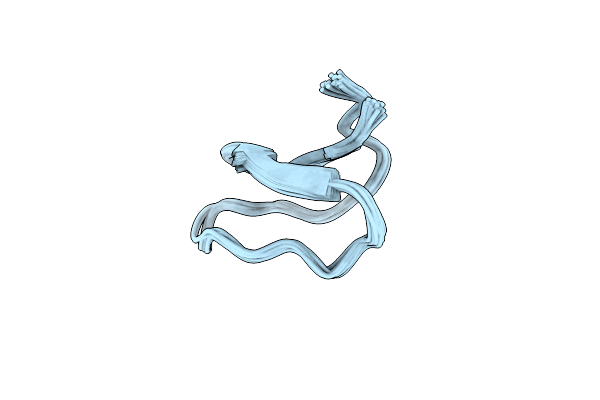 |
Organism: Oldenlandia affinis
Method: SOLUTION NMR Release Date: 2013-11-20 Classification: PLANT PROTEIN |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2010-01-19 Classification: TRANSFERASE Ligands: STJ, STI, CL |
 |
Crystal Structure Of Inactive Conformation Abl Kinase Catalytic Domain Complexed With Type Ii Inhibitor
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2006-08-22 Classification: TRANSFERASE Ligands: 7MP |

