Search Count: 34
All
Selected
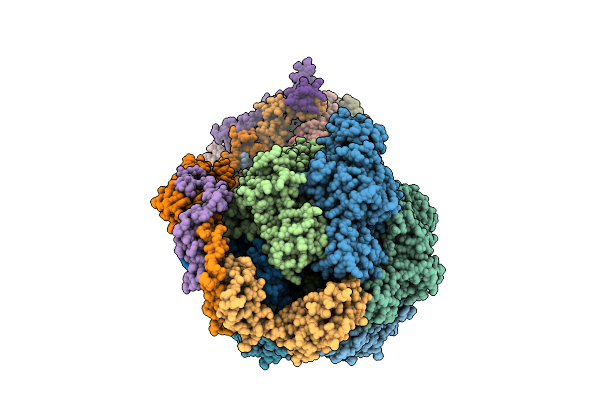 |
Cryo-Em Structure Of Fof1-Atpase Monomer State 1 On The Bovine Heart Submitochondrial Particles (Fof1-1)
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2026-04-01 Classification: MOTOR PROTEIN Ligands: ATP, MG, ADP |
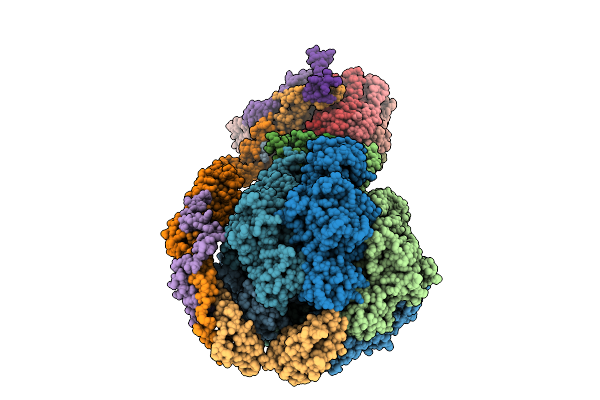 |
Cryo-Em Structure Of Fof1-Atpase Monomer State 3 On The Bovine Heart Submitochondrial Particles (Fof1-2)
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Resolution:4.00 Å Release Date: 2026-04-01 Classification: MOTOR PROTEIN |
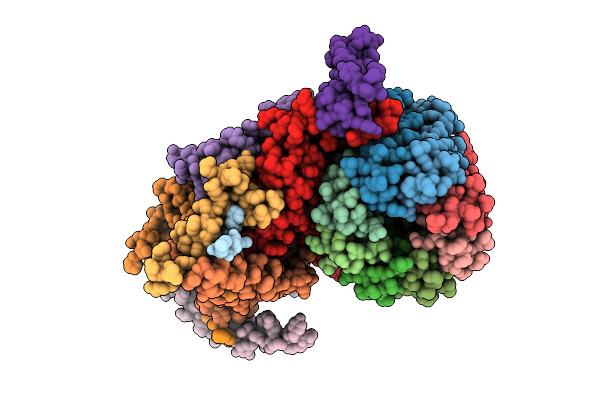 |
Cryo-Em Structure Of Fo Domain Of Fof1-Atpase Monomer State On The Bovine Heart Submitochondrial Particles
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Resolution:4.10 Å Release Date: 2026-04-01 Classification: MOTOR PROTEIN |
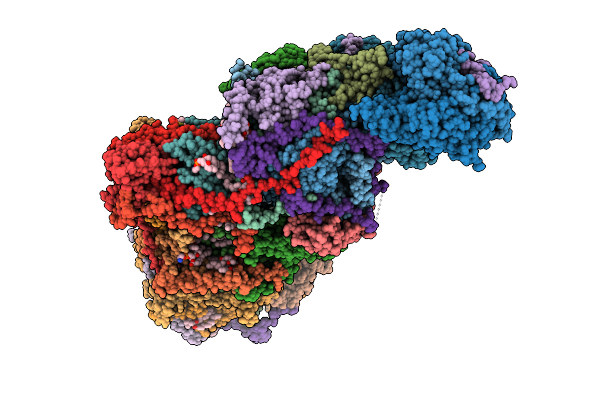 |
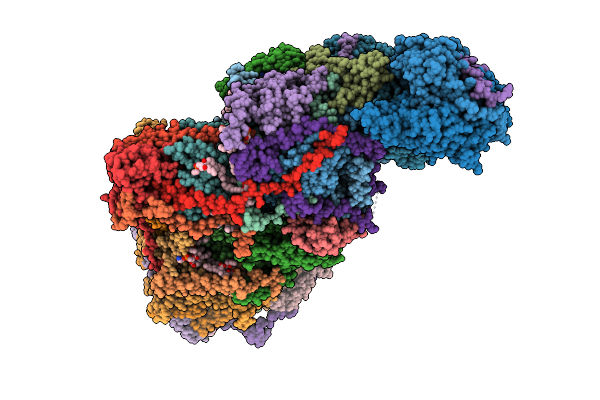
Cryo-Em Structure Of Complex I On The Bovine Heart Submitochondrial Particles, Open
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: FME, PC1, SF4, FES, FMN, K, 3PE, CDL, CHD, GTP, MG, NDP, ZN, MYR |
 |
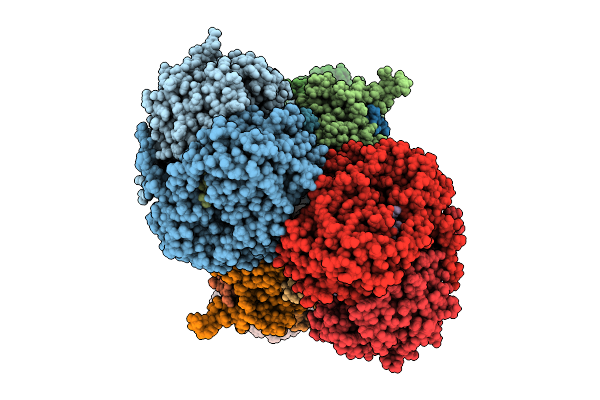
Cryo-Em Structure Of Complex I On The Bovine Heart Submitochondrial Particles, Closed
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: FME, 3PE, PC1, SF4, FES, FMN, K, CDL, MG, GTP, NDP, ZN |
 |
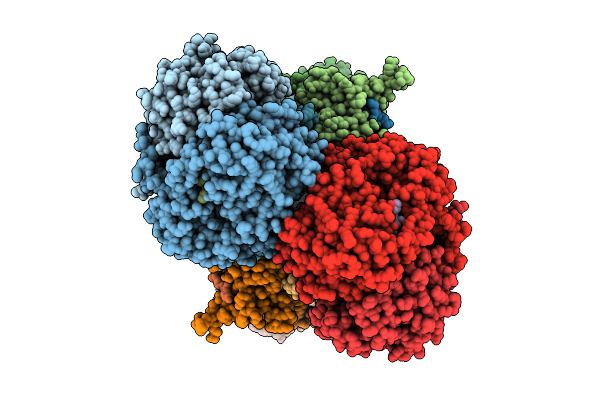
Cryo-Em Structure Of Complex Iii On The Bovine Heart Submitochondrial Particles, Iii-1
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MOTOR PROTEIN Ligands: HEM, HEC, FES |
 |
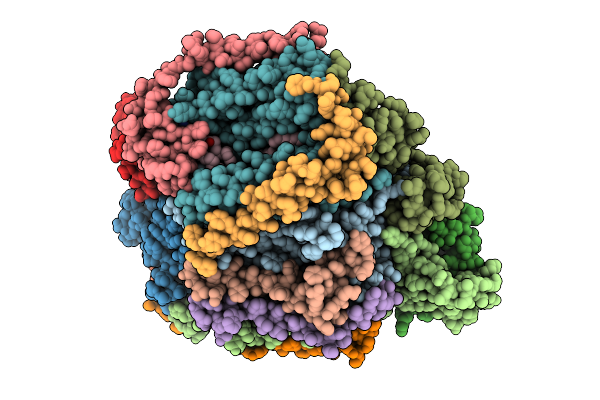
Cryo-Em Structure Of Complex Iii On The Bovine Heart Submitochondrial Particles, Iii-2
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: HEM, HEC, FES |
 |
Cryo-Em Structure Of Complex Iv On The Bovine Heart Submitochondrial Particles, Iv-A
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: HEA, CU, MG, PER, PGV, CUA, NA, ZN, PEK |
 |
Cryo-Em Structure Of Na+-Translocating Nadh-Ubiquinone Oxidoreductase From Vibrio Cholerae Reduced By Nadh, In The Absence Of Na+, Middle State
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Resolution:3.75 Å Release Date: 2025-08-06 Classification: MEMBRANE PROTEIN Ligands: FMN, RBF, CA, LMT, FES, FAD |
 |
Cryo-Em Structure Of Na+-Translocating Nadh-Ubiquinone Oxidoreductase Nqrc-T225Y Mutant From Vibrio Cholerae
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Resolution:2.66 Å Release Date: 2025-06-25 Classification: MEMBRANE PROTEIN Ligands: FMN, RBF, LMT, CA, FES, FAD |
 |
Cryo-Em Structure Of Na+-Translocating Nadh-Ubiquinone Oxidoreductase Nqrc-T225Y Mutant From Vibrio Cholerae Reduced By Nadh
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Resolution:2.59 Å Release Date: 2025-06-25 Classification: MEMBRANE PROTEIN Ligands: FMN, RBF, LMT, CA, FES, FAD |
 |
Cryo-Em Structure Of Na+-Translocating Nadh-Ubiquinone Oxidoreductase Nqrb-T236Y Mutant From Vibrio Cholerae
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Resolution:3.80 Å Release Date: 2025-06-25 Classification: MEMBRANE PROTEIN Ligands: RBF, LMT, FMN, CA, FES, FAD |
 |
Cryo-Em Structure Of Na+-Translocating Nadh-Ubiquinone Oxidoreductase Nqrb-T236Y Mutant From Vibrio Cholerae Reduced By Nadh
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Resolution:3.31 Å Release Date: 2025-06-25 Classification: MEMBRANE PROTEIN Ligands: RBF, LMT, CA, FMN, FES, FAD |
 |
Cryo-Em Structure Of Na+-Translocating Nadh-Ubiquinone Oxidoreductase From Vibrio Cholerae Reduced By Nadh, With Bound Korormicin A
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2025-06-25 Classification: MEMBRANE PROTEIN Ligands: FMN, RBF, IQT, LMT, CA, FES, FAD |
 |
Cryo-Em Structure Of Na+-Translocating Nadh-Ubiquinone Oxidoreductase From Vibrio Cholerae Reduced By Nadh, In The Absence Of Na+, Upper State
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Resolution:2.65 Å Release Date: 2025-06-25 Classification: MEMBRANE PROTEIN Ligands: FMN, RBF, LMT, CA, FES, FAD |
 |
Cryo-Em Structure Of Na+-Translocating Nadh-Ubiquinone Oxidoreductase From Vibrio Cholerae Reduced By Nadh, In The Absence Of Na+, Down State
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Resolution:3.11 Å Release Date: 2025-06-25 Classification: MEMBRANE PROTEIN Ligands: FMN, RBF, CA, FES, FAD |
 |
Cryo-Em Structure Of Na+-Translocating Nadh-Ubiquinone Oxidoreductase Nqrb-G141A Mutant From Vibrio Cholerae Reduced By Nadh, With Bound Korormicin A, Stable State
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Resolution:2.61 Å Release Date: 2025-06-25 Classification: MEMBRANE PROTEIN Ligands: FMN, RBF, IQT, CA, FES, FAD |
 |
Cryo-Em Structure Of Na+-Translocating Nadh-Ubiquinone Oxidoreductase Nqrb-G141A Mutant From Vibrio Cholerae Reduced By Nadh, With Bound Korormicin A, Shifted State
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Resolution:2.93 Å Release Date: 2025-06-25 Classification: MEMBRANE PROTEIN Ligands: FMN, RBF, PEE, LMT, IQT, CA, FES, FAD |
 |
Cryo-Em Structure Of Na+-Translocating Nadh-Ubiquinone Oxidoreductase From Vibrio Cholerae Reduced By Nadh, With Bound Aurachin D-42
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Resolution:3.18 Å Release Date: 2025-06-25 Classification: MEMBRANE PROTEIN Ligands: FMN, RBF, 0NI, PEE, LMT, CA, FES, FAD |
 |
Cryo-Em Structure Of Na+-Translocating Nadh-Ubiquinone Oxidoreductase From Vibrio Cholerae Reduced By Nadh
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Release Date: 2025-06-25 Classification: MEMBRANE PROTEIN Ligands: FMN, RBF, CA, FES, FAD, NAI |

