Search Count: 68
All
Selected
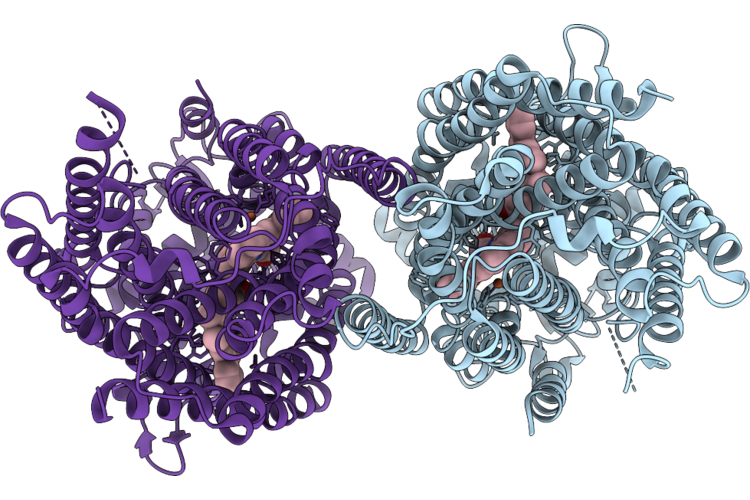 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase At Ph 8.0 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
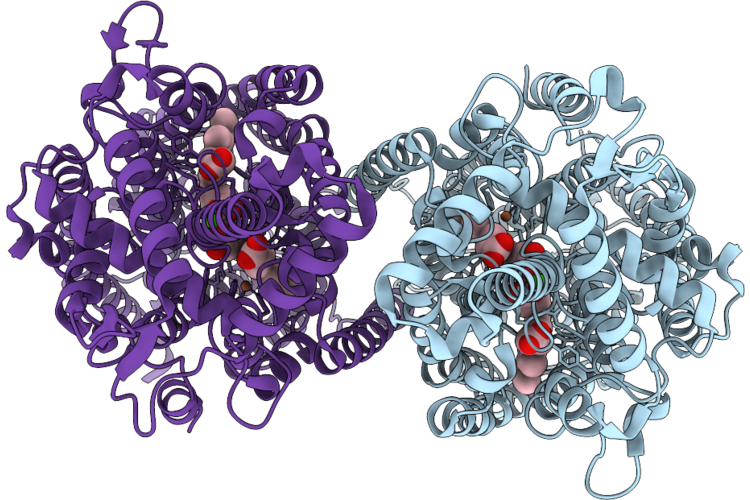 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Arg720Ala Variant At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
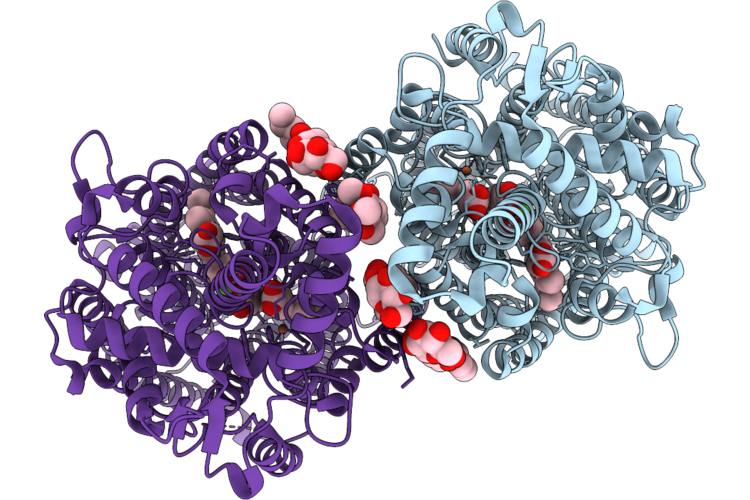 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Trp718Ala Variant At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, LMT, UQ5 |
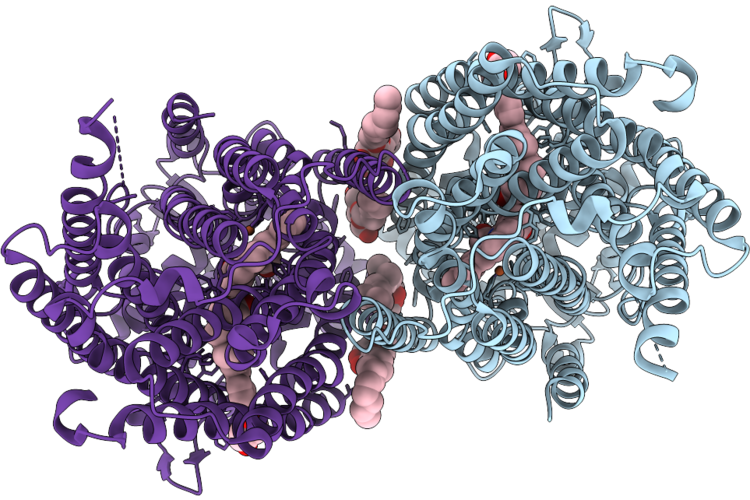 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Trp718Ala Variant With Quino At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, UQ5, HQE, LMT |
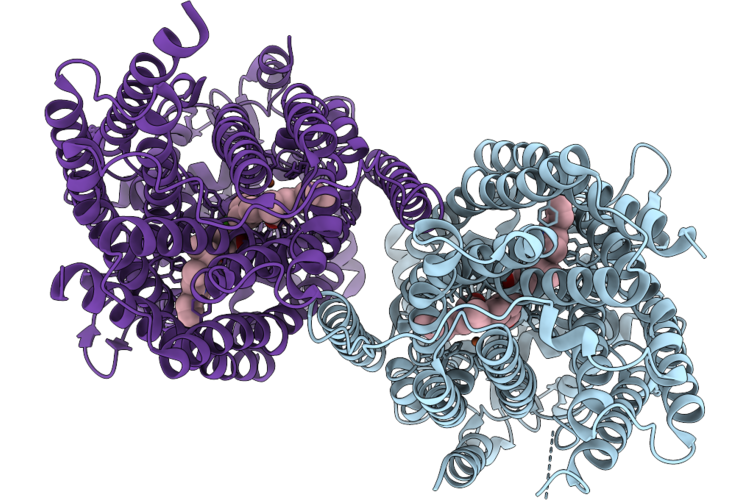 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase At Ph 6.5
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
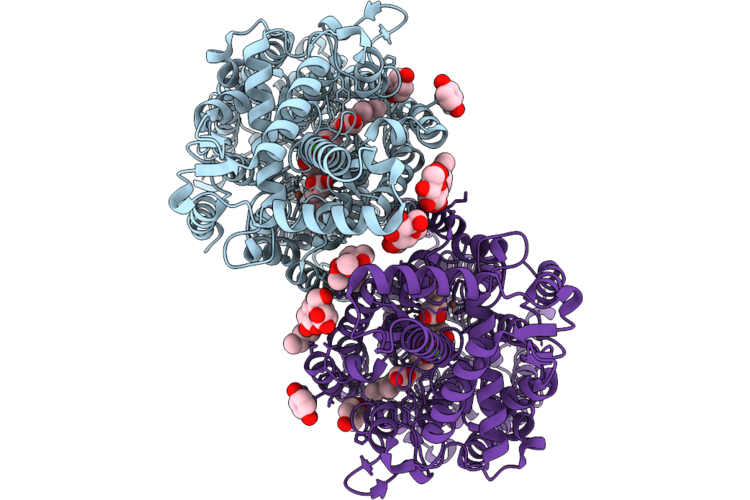 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase With Hqe At Ph 6.5
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, LMT, HQE, UQ5 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
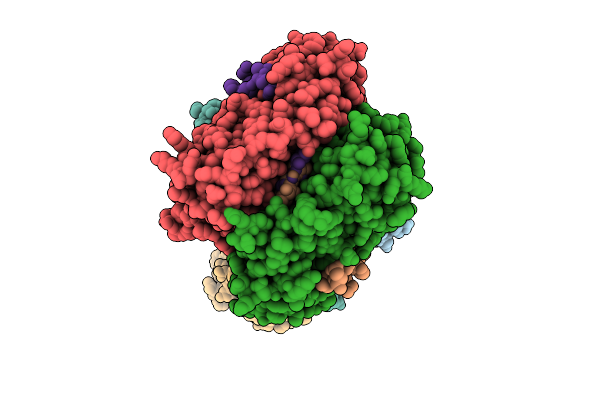 |
Cryoem Structure Of Human Mata2 In Complex With Mat2B Isoform V1 At 2.6 A Resolution
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
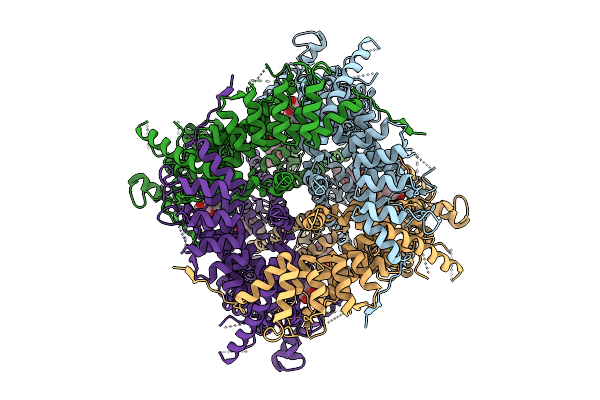 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: PROTEIN TRANSPORT Ligands: Y01, CA, A1IMP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: PROTEIN TRANSPORT Ligands: A1IMO, CA, Y01 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN Ligands: A1IIE |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN Ligands: ZN, A1IIF |
 |
Organism: Danio rerio
Method: ELECTRON MICROSCOPY Release Date: 2025-07-09 Classification: MEMBRANE PROTEIN Ligands: A1IGW |
 |
Organism: Danio rerio
Method: ELECTRON MICROSCOPY Release Date: 2025-07-09 Classification: MEMBRANE PROTEIN Ligands: A1IGY |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2025-06-18 Classification: ENDOCYTOSIS Ligands: CA, NAG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2025-06-18 Classification: ENDOCYTOSIS Ligands: NAG, CA, HEM, OXY |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:4.00 Å Release Date: 2025-06-18 Classification: ENDOCYTOSIS Ligands: CA, HEM, OXY |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-02-05 Classification: MEMBRANE PROTEIN |
 |
Organism: Mus musculus, Oryctolagus cuniculus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: MOTOR PROTEIN Ligands: PO4, MG, ADP |
 |
Organism: Mus musculus, Oryctolagus cuniculus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: MOTOR PROTEIN Ligands: MG, ADP |

