Search Count: 73
All
Selected
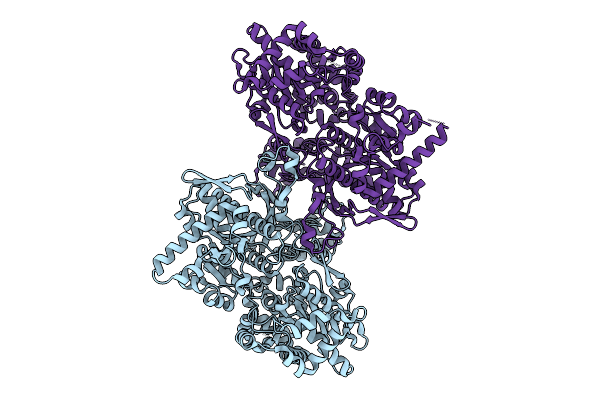 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
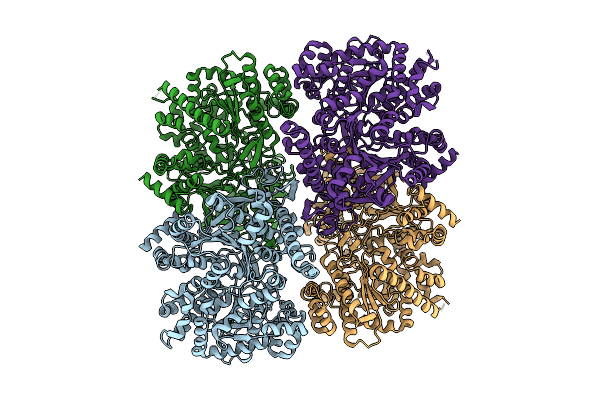 |
Structure Of Glycogen Phosphorylase - Tetrameric Form - From Escherichia Coli
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.97 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
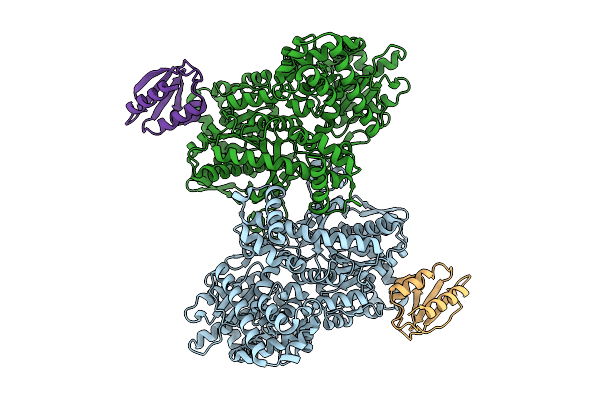 |
Structure Of Glycogen Phosphorylase - Dimeric Form - In Complex With Hpr From Escherichia Coli
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.79 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
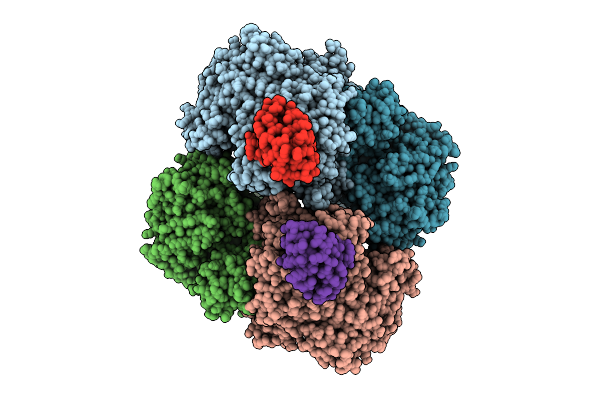 |
Structure Of Glycogen Phosphorylase - Tetrameric Form - In Complex With Hpr From Escherichia Coli
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.06 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
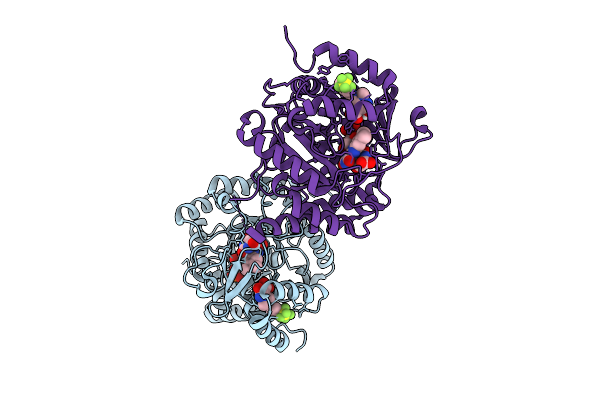 |
Plasmodium Falciparum Dihydroorotate Dehydrogenase In Complex With 3-Hydroxy-1-Methyl Pyrazole Derivatives
Organism: Plasmodium falciparum
Method: X-RAY DIFFRACTION Resolution:2.95 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: FMN, ORO, A1JWR |
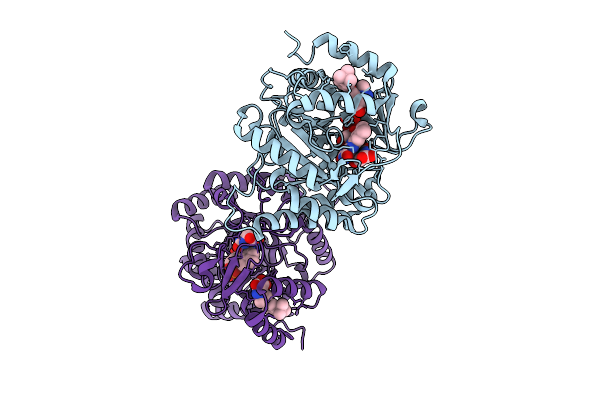 |
Plasmodium Falciparum Dihydroorotate Dehydrogenase In Complex With 3-Hydroxy-1-Methyl Pyrazole Derivatives
Organism: Plasmodium falciparum
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: FMN, ORO, A1JWQ |
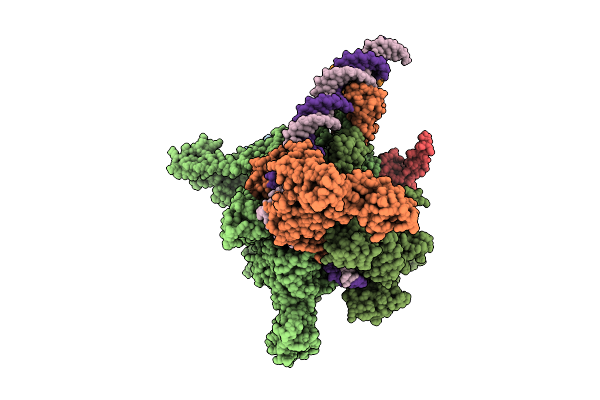 |
Cryo-Em Structure Of Vibrio Cholerae Rna Polymerase Holoenzyme Bound To An Ompu Promoter Dna Fragment
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: MG, ZN |
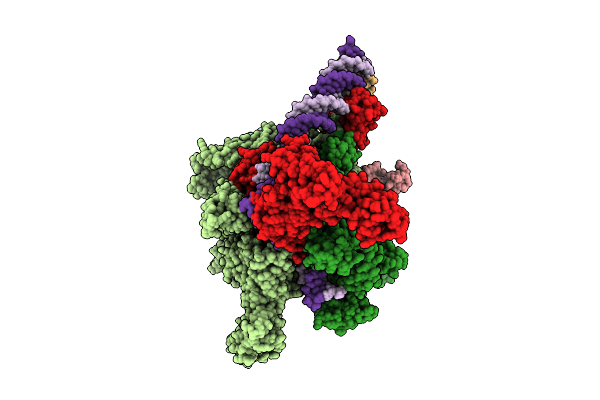 |
Cryo-Em Structure Of Vibrio Cholerae Rna Polymerase Holoenzyme Bound To An Ompu Promoter Dna Fragment And 5-Mer Rna
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: ZN, MG |
 |
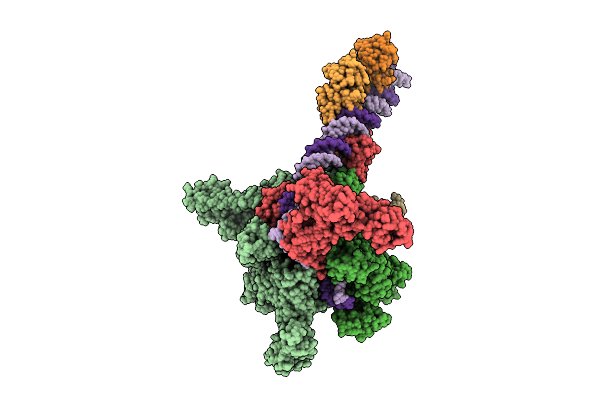
Cryo-Em Structure Of Vibrio Cholerae Rna Polymerase Transcription Activation Complex With Toxr Transcription Factor And Ompu Promoter Dna
Organism: Vibrio cholerae o395, Vibrio cholerae o1 biovar el tor str. n16961
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: ZN, MG |
 |
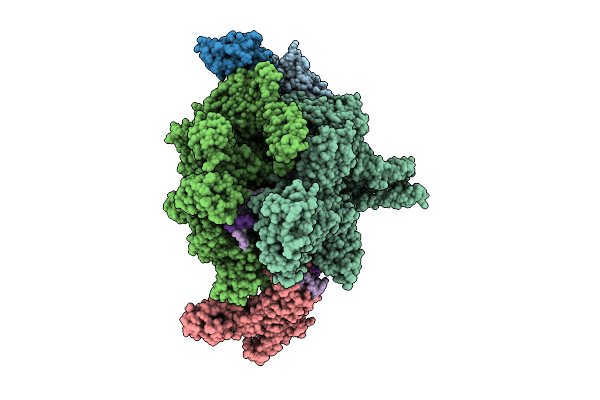
Cryo-Em Structure Of Vibrio Cholerae Rna Polymerase Transcription Activation Complex With Tcpp Transcription Factor And A Toxt Promoter Dna Fragment
Organism: Vibrio cholerae o395
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: ZN, MG |
 |
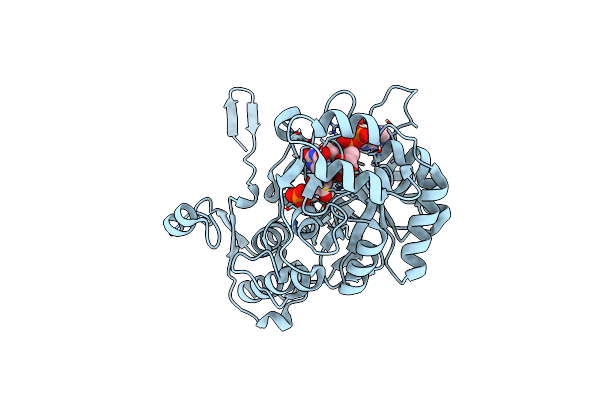
Cryo-Em Structure Of Vibrio Cholerae Rna Polymerase Transcription Activation Complex With Toxr And Tcpp Transcription Factors And A Toxt Promoter Dna Fragment
Organism: Vibrio cholerae o395, Vibrio cholerae o1 biovar el tor str. n16961
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: TRANSCRIPTION Ligands: MG, ZN |
 |
Structure A Catalytically Inactive Mutant Of The Imp Dehydrogenase Related Protein Guab3 From Synechocystis Pcc 6803
Organism: Synechocystis sp. pcc 6803
Method: X-RAY DIFFRACTION Resolution:1.22 Å Release Date: 2024-03-27 Classification: BIOSYNTHETIC PROTEIN Ligands: XMP, NAD |
 |
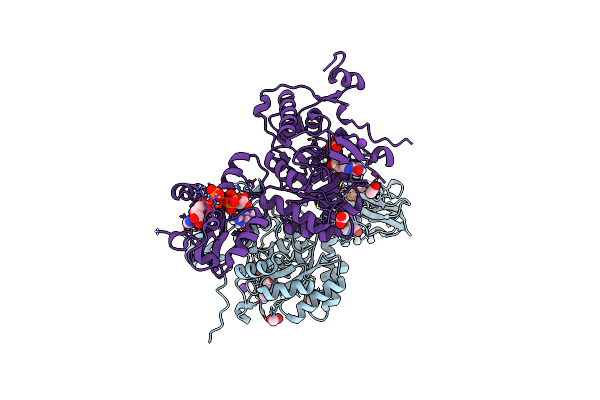
Structure Of The Imp Dehydrogenase Related Protein Guab3 From Synechocystis Pcc 6803
Organism: Synechocystis sp. pcc 6803
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2024-03-27 Classification: BIOSYNTHETIC PROTEIN Ligands: IMP, XMP |
 |
Crystal Structure Of Pseudomonas Aeruginosa Guab (Imp Dehydrogenase) Bound To Atp And Gdp At 1.65A Resolution
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2022-07-06 Classification: OXIDOREDUCTASE Ligands: ATP, GDP, IMP, MG, K, ACT |
 |
Crystal Structure Of Streptomyces Coelicolor Guab (Imp Dehydrogenase) Bound To Atp And Ppgpp At 2.0 A Resolution
Organism: Streptomyces coelicolor a3(2)
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2022-05-11 Classification: OXIDOREDUCTASE Ligands: ATP, G4P, MG |
 |
Crystal Structure Of The Pdz Tandem Of Syntenin In Complex With Fragment C58
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2021-02-03 Classification: SIGNALING PROTEIN Ligands: PO4, JVK, ACT |
 |
Crystal Structure Of The Pdz Tandem Of Syntenin In Complex With Fragment F13
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2021-02-03 Classification: SIGNALING PROTEIN Ligands: K7Z, ACT |
 |
Cryo-Em Structure Of The Glucagon-Like Peptide-1 Receptor In Complex With G Protein, Glp-1 Peptide And A Positive Allosteric Modulator
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2020-07-22 Classification: SIGNALING PROTEIN/MEMBRANE PROTEIN Ligands: QW7 |
 |
Crystal Structure Of The Syntenin Pdz1 Domain In Complex With The Peptide Inhibitor Ksl-128018
Organism: Rattus norvegicus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2019-09-04 Classification: SIGNALING PROTEIN/INHIBITOR |
 |
Crystal Structure Of The Syntenin Pdz1 And Pdz2 Tandem In Complex With The Frizzled 7 C-Terminal Fragment And Pip2
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2016-06-29 Classification: SIGNALING PROTEIN Ligands: ACT, GOL, IP2 |

