Search Count: 8
All
Selected
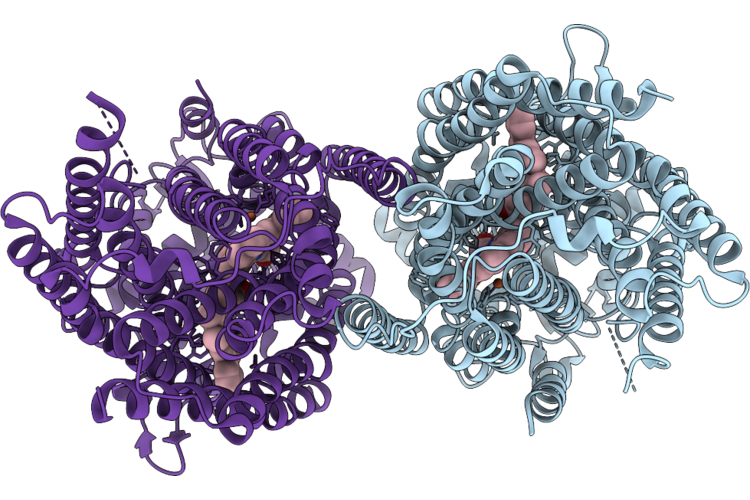 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase At Ph 8.0 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
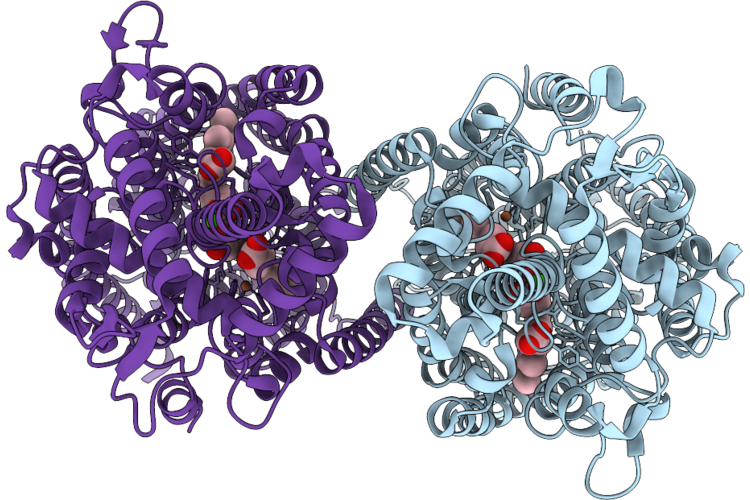 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Arg720Ala Variant At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
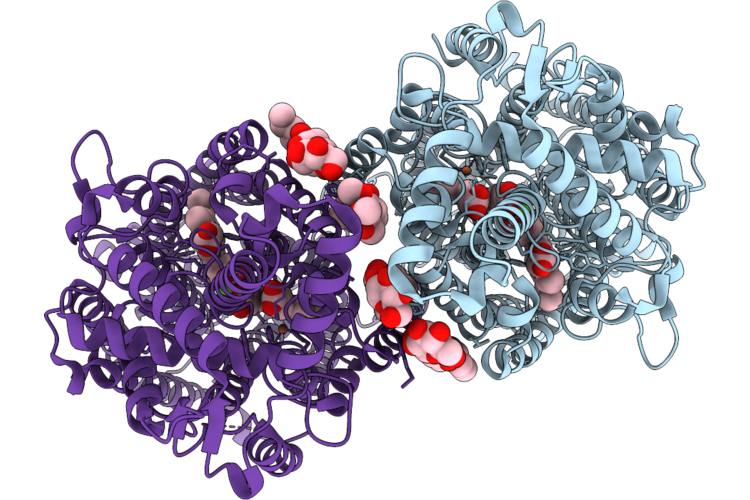 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Trp718Ala Variant At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, LMT, UQ5 |
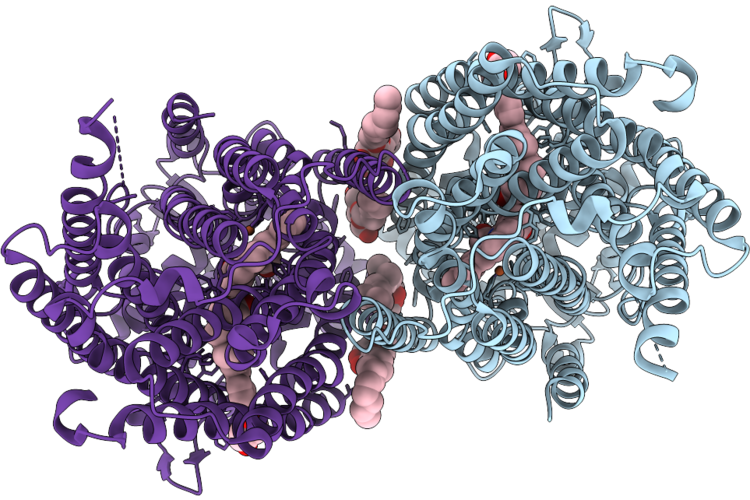 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Trp718Ala Variant With Quino At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, UQ5, HQE, LMT |
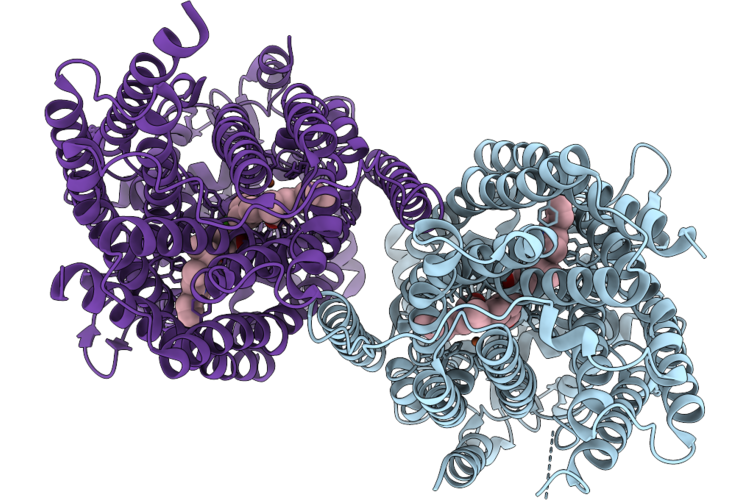 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase At Ph 6.5
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
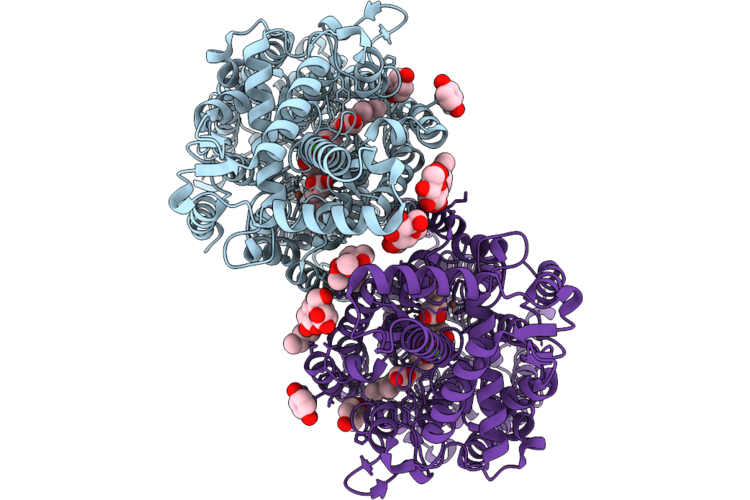 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase With Hqe At Ph 6.5
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, LMT, HQE, UQ5 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2023-10-25 Classification: RNA BINDING PROTEIN |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2023-10-25 Classification: RNA BINDING PROTEIN |

