Search Count: 377
All
Selected
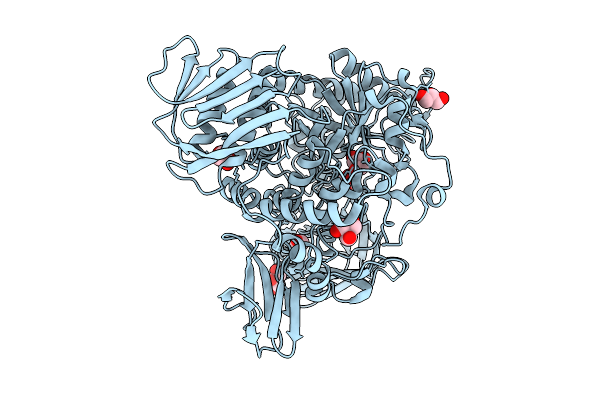 |
Sdacs: A Synergistic Strategy Using Both Crystallographic And Alphafold Structures To Improve Thermostability And Activity Of N-Acetylhexosaminidase
Organism: Cellvibrio mixtus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: TRS, GOL |
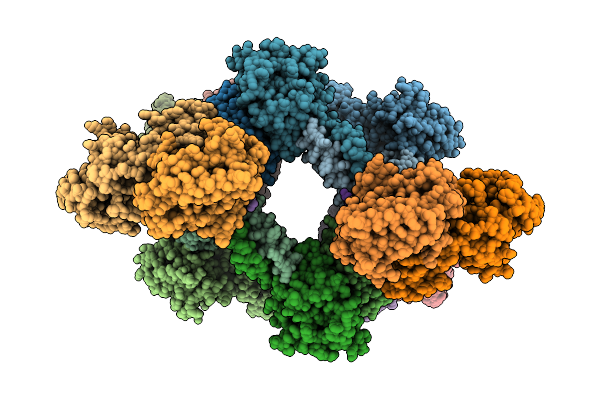 |
Organism: Enterococcus faecalis
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN Ligands: CA |
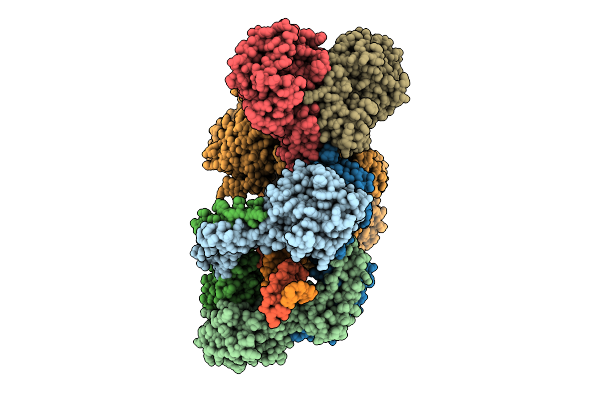 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA/RNA |
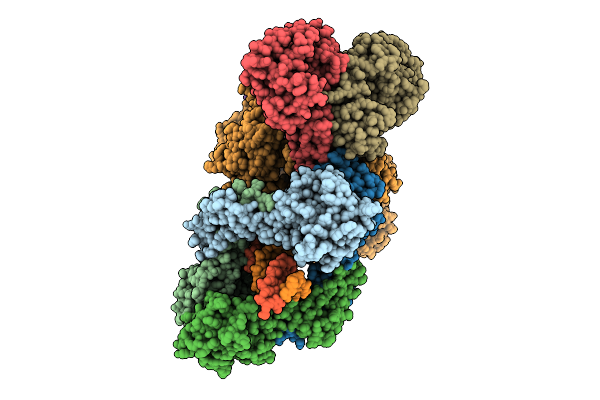 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.01 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.76 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA Ligands: CA |
 |
Organism: Psychrobacter lutiphocae dsm 21542, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN/RNA Ligands: CA, GOL, PEG |
 |
Organism: Psychrobacter lutiphocae dsm 21542, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN/RNA Ligands: MG |
 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.17 Å Release Date: 2025-12-31 Classification: HYDROLASE Ligands: PEG, A1EDS |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: HYDROLASE Ligands: 4BW |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2025-11-19 Classification: HYDROLASE |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2025-11-19 Classification: HYDROLASE |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.04 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: 4BW |
 |
Organism: Human immunodeficiency virus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Human immunodeficiency virus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Human immunodeficiency virus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Human immunodeficiency virus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Human immunodeficiency virus 1, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Loop-Deleted Dna Polymerase Theta In Complex With A Dsdna Overhang And An Allostertic Inhibitor
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2025-08-20 Classification: DNA BINDING PROTEIN Ligands: MG, DG3, A1A2C |
 |
Loop-Deleted Dna Polymerase Theta Polymerase Domain In Complex With A Dsdna Overhang And An Allosteric Inhibitor
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.31 Å Release Date: 2025-08-20 Classification: DNA BINDING PROTEIN Ligands: MG, DG3, A1A2D |

