Search Count: 6
All
Selected
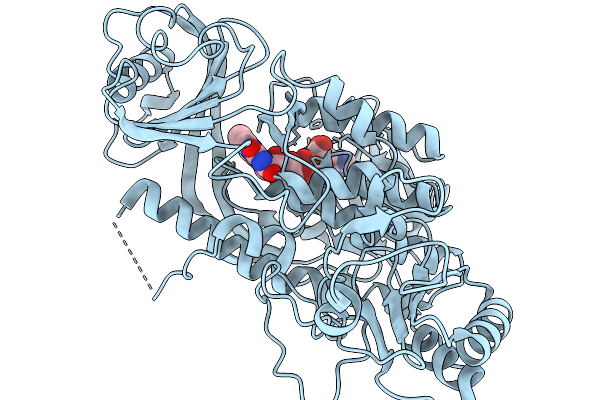 |
Crystal Structure Of The Flavoprotein Monooxygenase Rslo9 From Streptomyces Bottropensis
Organism: Streptomyces bottropensis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: FAD, CL |
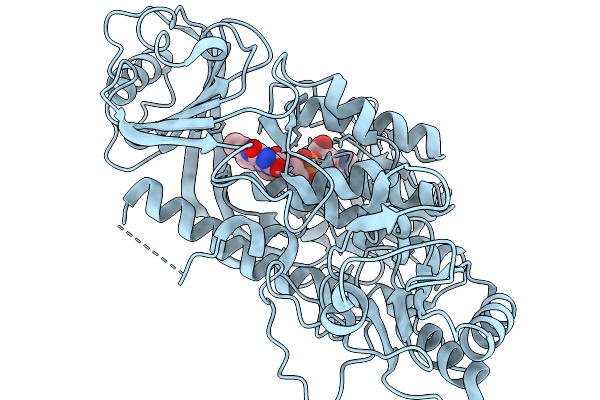 |
Crystal Structure Of The Flavoprotein Monooxygenase Rslo9 From Streptomyces Bottropensis (Alternative Loop Conformation)
Organism: Streptomyces bottropensis
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: FAD |
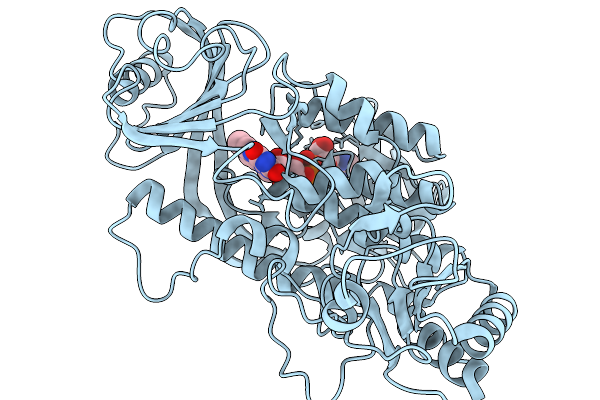 |
Crystal Structure Of The Flavoprotein Monooxygenase Rslo9 From Streptomyces Bottropensis (Alternative N-Terminus)
Organism: Streptomyces bottropensis
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Crystal Structure Of The Flavoprotein Monooxygenase Trle From Streptomyces Cyaneofuscatus Soc7
Organism: Streptomyces cyaneofuscatus
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2024-05-15 Classification: FLAVOPROTEIN Ligands: FAD, SO4, CL |
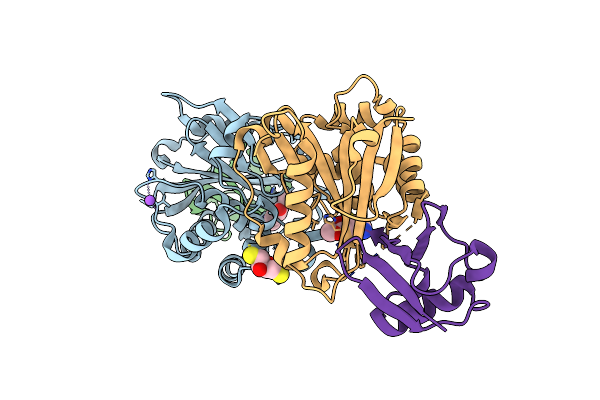 |
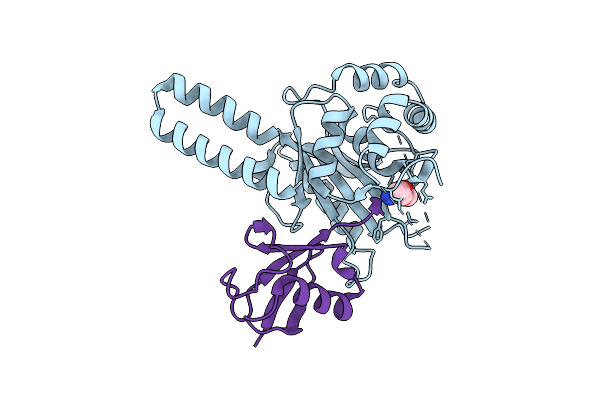
Crystal Structure Of Trichinella Spiralis Uch37 Catalytic Domain Bound To Ubiquitin Vinyl Methyl Ester
Organism: Trichinella spiralis, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2013-05-29 Classification: Hydrolase/Signaling Protein Ligands: DTT, NA, GVE |
 |
Crystal Structure Of Trichinella Spiralis Uch37 Bound To Ubiquitin Vinyl Methyl Ester
Organism: Trichinella spiralis, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2013-05-29 Classification: Hydrolase/Signaling Protein Ligands: GVE |

