Search Count: 24
All
Selected
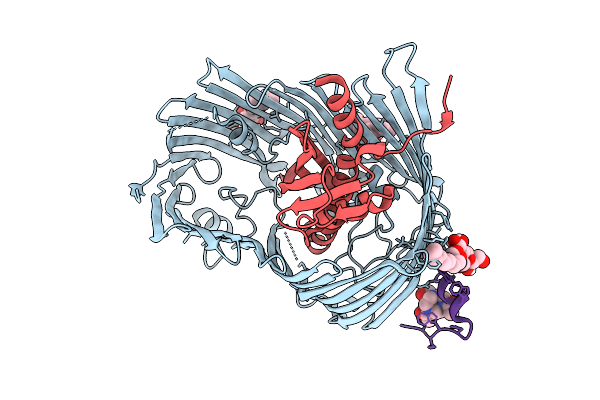 |
Cryo-Em Structure Of Shigella Flexneri Lptde Bound By A Bicyclic Peptide Molecule (Compound 1)
Organism: Shigella flexneri, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.41 Å Release Date: 2025-10-15 Classification: MEMBRANE PROTEIN Ligands: LMT, A1I1D |
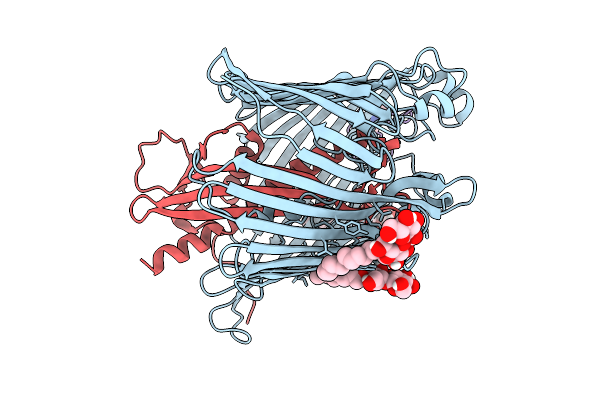 |
Cryo-Em Structure Of Shigella Flexneri Lptde Bound By A Bicyclic Peptide Molecule (Compound 2)
Organism: Shigella flexneri, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.86 Å Release Date: 2025-10-15 Classification: MEMBRANE PROTEIN Ligands: LMT, KZ0 |
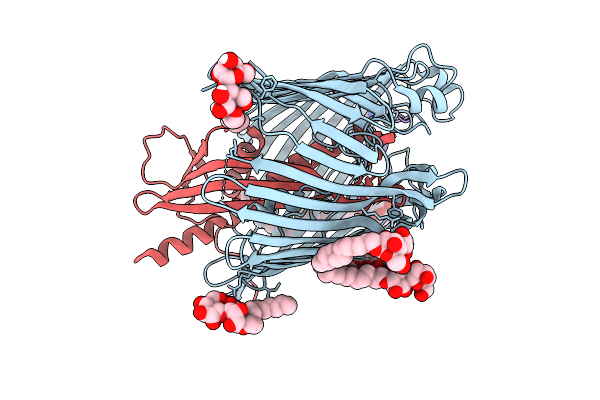 |
Cryo-Em Structure Of Shigella Flexneri Lptde Bound By A Bicyclic Peptide Molecule (Compound 3)
Organism: Shigella flexneri, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2025-10-15 Classification: MEMBRANE PROTEIN Ligands: LMT, KZ0 |
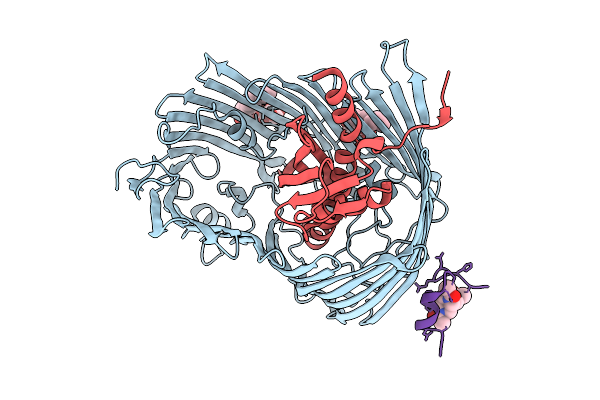 |
Cryo-Em Structure Of Shigella Flexneri Lptde Bound By A Bicyclic Peptide Molecule (Compound 4)
Organism: Shigella flexneri, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.35 Å Release Date: 2025-10-15 Classification: MEMBRANE PROTEIN Ligands: LMT, A1I1E |
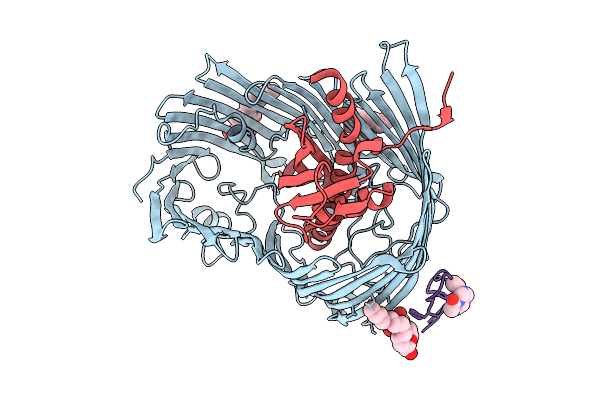 |
Cryo-Em Structure Of Shigella Flexneri Lptde Bound By A Bicyclic Peptide Molecule (Compound 5)
Organism: Shigella flexneri, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.67 Å Release Date: 2025-10-15 Classification: MEMBRANE PROTEIN Ligands: LMT, R06 |
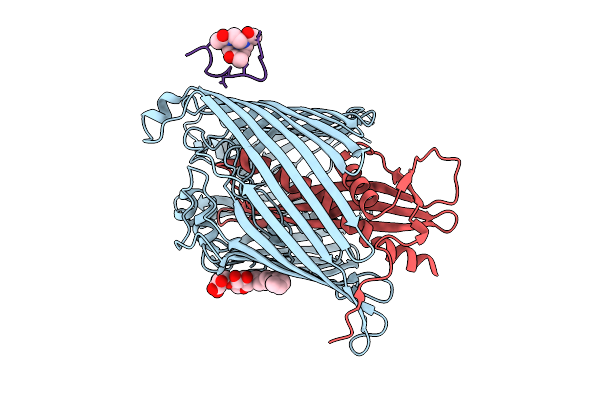 |
Cryo-Em Structure Of Shigella Flexneri Lptde In Complex With A Bicyclic Peptide Binder (Compound 12)
Organism: Shigella flexneri, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.48 Å Release Date: 2025-10-15 Classification: MEMBRANE PROTEIN Ligands: LMT, A1I1D |
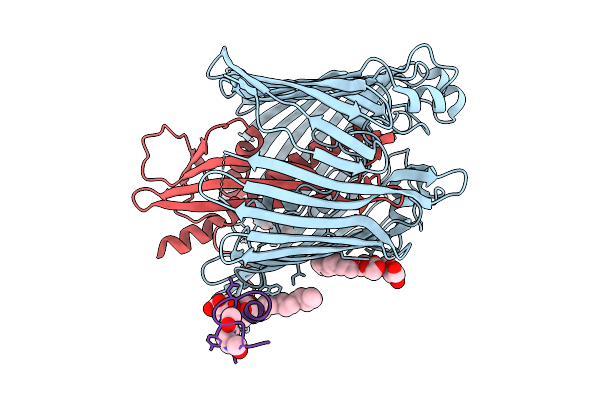 |
Cryo-Em Structure Of Shigella Flexneri Lptde Bound By A Bicyclic Peptide Molecule (Compound 13)
Organism: Shigella flexneri, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.86 Å Release Date: 2025-10-15 Classification: MEMBRANE PROTEIN Ligands: LMT, R06 |
 |
Cryo-Em Structure Of Shigella Flexneri Lptde Bound By A Bicyclic Peptide Molecule (Compound 16)
Organism: Shigella flexneri, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.54 Å Release Date: 2025-10-15 Classification: MEMBRANE PROTEIN Ligands: LMT, PLM, Z41, A1I4O |
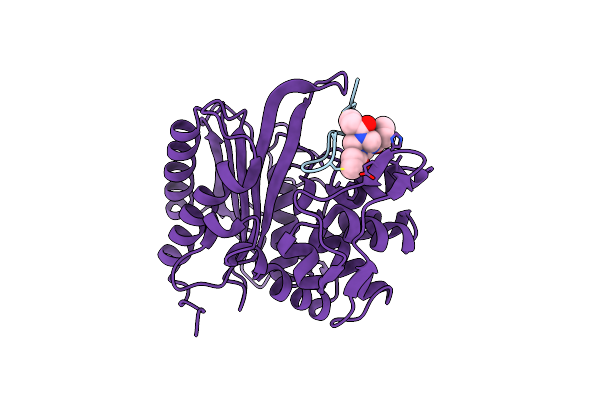 |
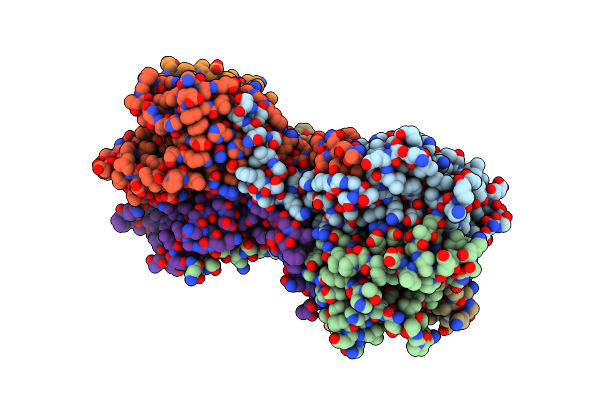
The Structure Of E. Coli Penicillin Binding Protein 3 (Pbp3) In Complex With A Bicyclic Peptide Inhibitor
Organism: Escherichia coli, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2024-04-03 Classification: HYDROLASE Ligands: 29N |
 |
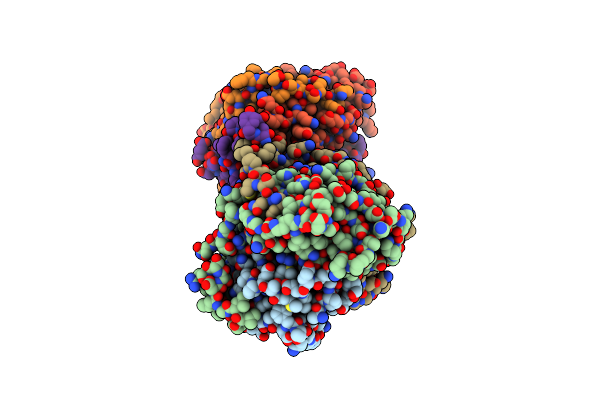
Organism: Pisum sativum
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2024-01-31 Classification: PLANT PROTEIN |
 |
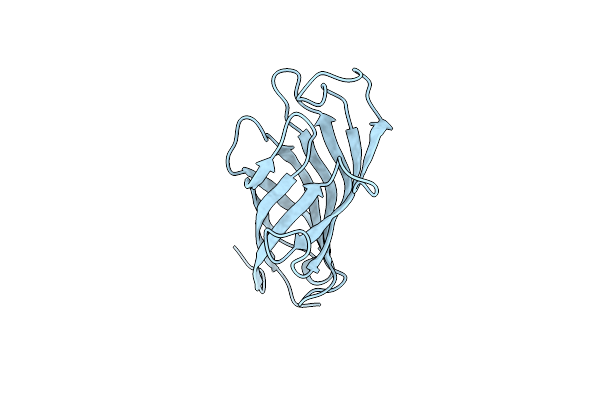
Organism: Glycyrrhiza echinata
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2020-04-29 Classification: PLANT PROTEIN |
 |
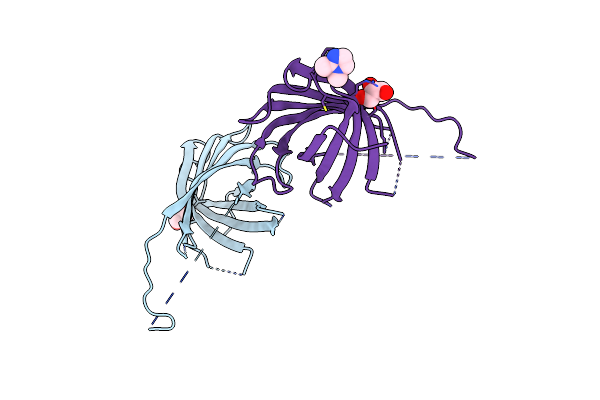
Organism: Pisum sativum
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2020-04-29 Classification: PLANT PROTEIN |
 |
Organism: Pisum sativum
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2014-11-26 Classification: PLANT PROTEIN Ligands: GOL, DMI, CL |
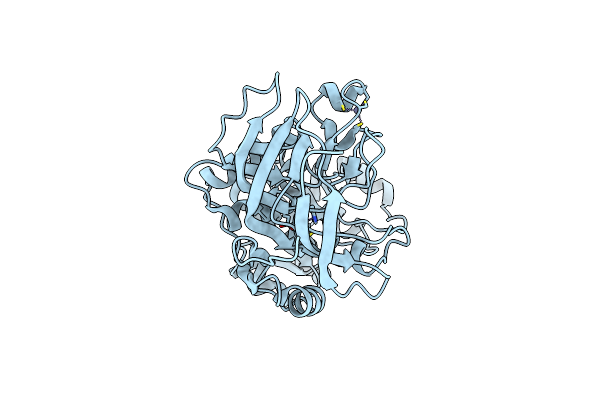 |
Crystal Structures Of The Arabidopsis Cinnamyl Alcohol Dehydrogenases, Atcad5
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2007-02-20 Classification: OXIDOREDUCTASE Ligands: ZN |
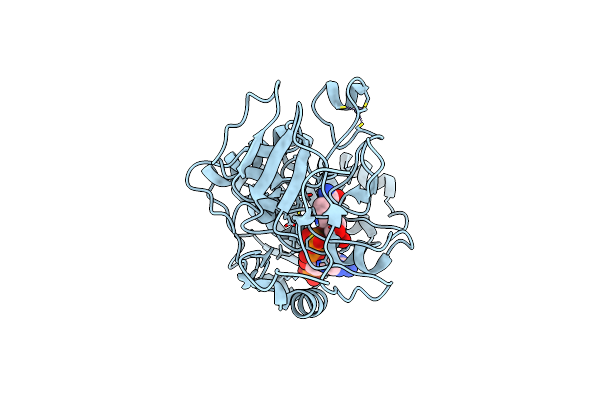 |
Crystal Structures Of The Arabidopsis Cinnamyl Alcohol Dehydrogenases Atcad5
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2007-02-20 Classification: OXIDOREDUCTASE Ligands: ZN, NAP |
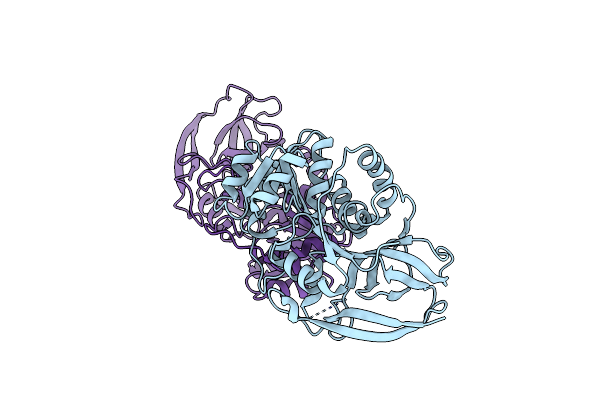 |
Crystal Structure Of Arabidopsis Thaliana Double Bond Reductase (At5G16970)-Apo Form
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2006-10-05 Classification: OXIDOREDUCTASE |
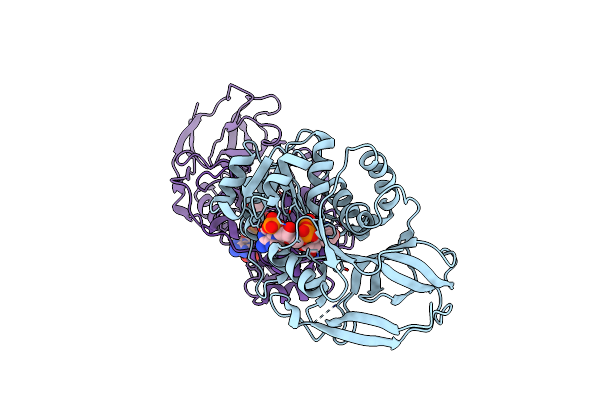 |
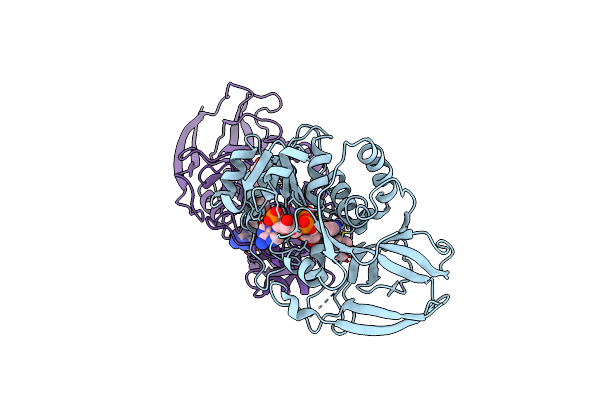
Crystal Structure Of Arabidopsis Thaliana Double Bond Reductase (At5G16970)-Binary Complex
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2006-10-05 Classification: OXIDOREDUCTASE Ligands: NAP |
 |
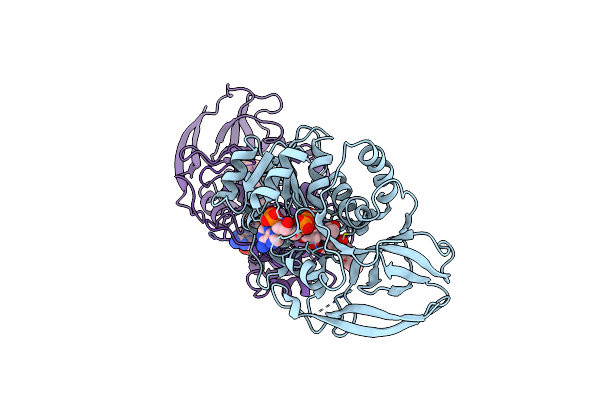
Crystal Structure Of Arabidopsis Thaliana Double Bond Reductase (At5G16970)-Ternary Complex I
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2006-10-05 Classification: OXIDOREDUCTASE Ligands: NAP, HC4 |
 |
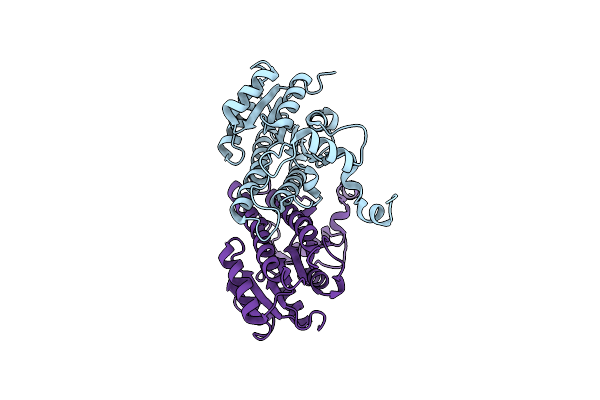
Crystal Structure Of Arabidopsis Thaliana Double Bond Reductase (At5G16970)-Ternary Complex Ii
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2006-10-05 Classification: OXIDOREDUCTASE Ligands: NAP, HNE |
 |
Organism: Podophyllum peltatum
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2005-01-13 Classification: OXIDOREDUCTASE |

