Search Count: 59
All
Selected
 |
Structure Of Mycobacterium Tuberculosis Inha In Complex With Pyridomycin (Compound 1)
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: PYW, PGE |
 |
Structure Of Mycobacterium Tuberculosis Inha In Complex With Pyridomycin-Derivative Kv26A (Compound 4)
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: A1JGW |
 |
Structure Of Mycobacterium Tuberculosis Inha In Complex With Pyridomycin-Derivative Kv29A (Compound 5)
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: A1JG1 |
 |
Structure Of Mycobacterium Tuberculosis Inha In Complex With Pyridomycin Derivative Kv37A (Compound 9)
Organism: Mycobacterium tuberculosis '98-r604 inh-rif-em'
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: A1JG0 |
 |
Structure Of Mycobacterium Tuberculosis Inha In Complex With Pyridomycin Derivative Kv41A (Compound 11)
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: A1JG3, PEG |
 |
Structure Of Mycobacterium Tuberculosis Inha In Complex With Pyridomycin Derivative Kv35A (Compound 12)
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: A1JGY |
 |
Structure Of Mycobacterium Tuberculosis Inha In Complex With Pyridomycin Derivative Ep196 (Compound 13)
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: A1JGX |
 |
Structure Of Mycobacterium Tuberculosis Inha In Complex With Pyridomycin Derivative Kv32A (Compound 14)
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: A1JGZ |
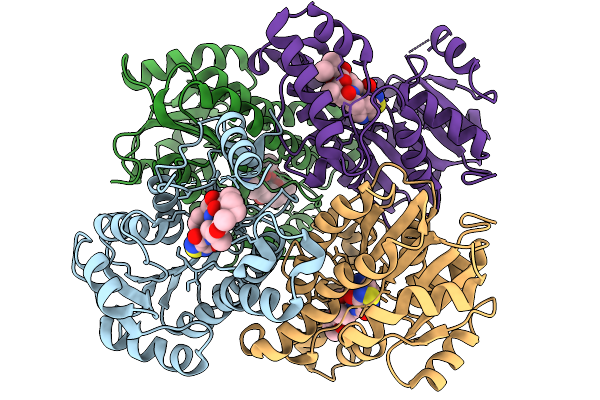 |
Structure Of Mycobacterium Tuberculosis Inha In Complex With Pyridomycin Derivative Kv25A (Compound 15)
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: A1JG2 |
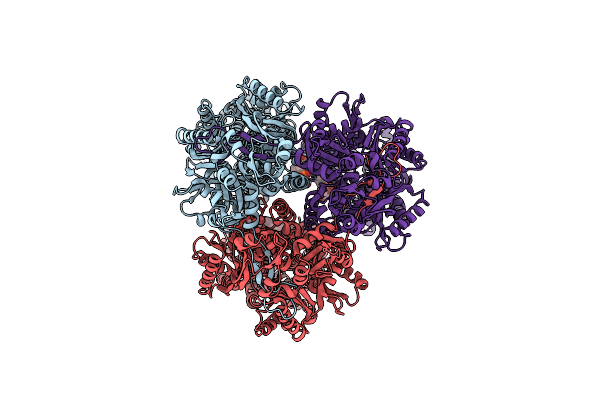 |
Single Particle Cryo-Em Co-Structure Of Klebsiella Pneumoniae Acrb With The Bdm91288 Efflux Pump Inhibitor At 2.97 Angstrom Resolution
Organism: Klebsiella pneumoniae subsp. pneumoniae dsm 30104 = jcm 1662 = nbrc 14940
Method: ELECTRON MICROSCOPY Release Date: 2024-01-17 Classification: TRANSPORT PROTEIN Ligands: WQW, WR6, PG8 |
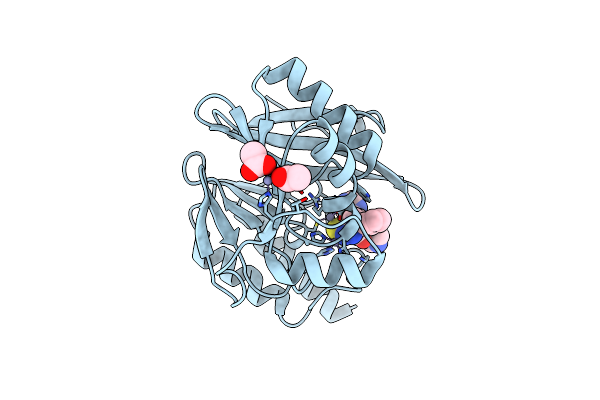 |
Crystal Structure Of The Vim-2 Acquired Metallo-Beta-Lactamase In Complex With Compound 8 (Jmv-7061)
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2023-04-26 Classification: HYDROLASE Ligands: ZN, ACT, L2R |
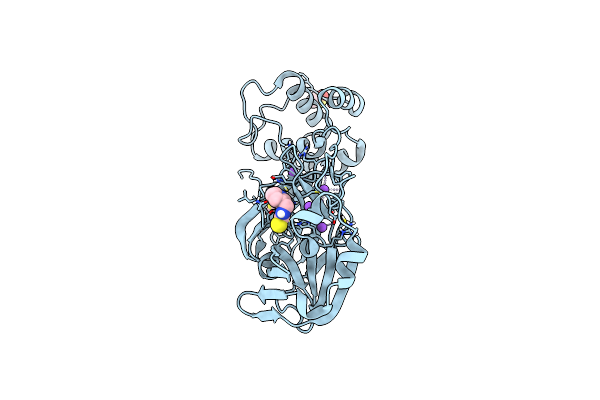 |
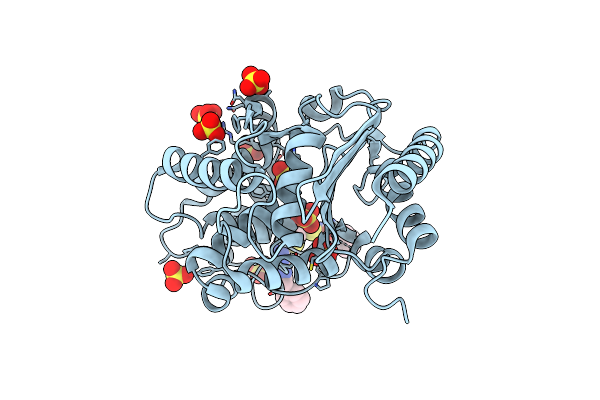
Sars-Cov-2 Main Protease Complexed With N-(Pyridin-3-Ylmethyl)Thioformamide
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2023-03-15 Classification: VIRAL PROTEIN Ligands: DMS, 35J, NA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2023-01-18 Classification: HYDROLASE Ligands: SO4, 04A |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.38 Å Release Date: 2023-01-18 Classification: HYDROLASE Ligands: R90 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.78 Å Release Date: 2023-01-18 Classification: HYDROLASE Ligands: SO4, 5XX |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2023-01-18 Classification: HYDROLASE Ligands: SO4, R0C |
 |
Metallo-Beta-Lactamase Ndm-1 In Complex With 1,2,4-Triazole-3-Thione Compound 26
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2022-12-14 Classification: HYDROLASE Ligands: ZN, CA, L82, EDO, PEG, EPE |
 |
Crystal Structure Of Endoplasmic Reticulum Aminopeptidase 2 (Erap2) Complex With A Highly Selective And Potent Small Molecule
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2022-07-20 Classification: IMMUNE SYSTEM,HYDROLASE/INHIBITOR Ligands: ZN, GIY, EPE, MES, NA |
 |
Crystal Structure Of The Sars-Cov-2 Main Protease Complexed With N-(Pyridin-3-Ylmethyl)Thioformamide
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2022-03-02 Classification: VIRAL PROTEIN Ligands: 35J, NA, FMT, DMS |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.48 Å Release Date: 2021-10-06 Classification: VIRAL PROTEIN Ligands: DMS, GOL, FMT |

