Search Count: 256
All
Selected
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-04-22 Classification: RNA BINDING PROTEIN/RNA |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.47 Å Release Date: 2026-04-22 Classification: RNA BINDING PROTEIN/RNA Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.93 Å Release Date: 2026-04-22 Classification: RNA BINDING PROTEIN/RNA Ligands: ZN |
 |
Organism: Mycobacterium tuberculosis h37rv, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: DNA BINDING PROTEIN |
 |
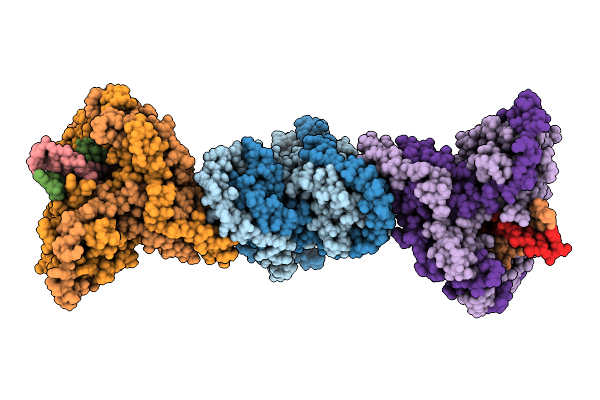
Organism: Mycobacterium tuberculosis h37rv, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: DNA BINDING PROTEIN/DNA |
 |
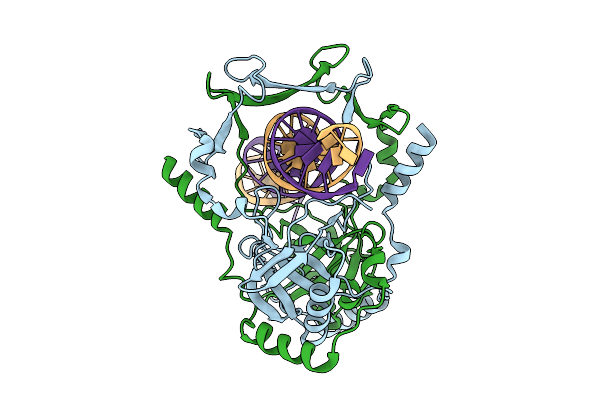
Organism: Mycobacterium tuberculosis h37rv, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: DNA BINDING PROTEIN |
 |
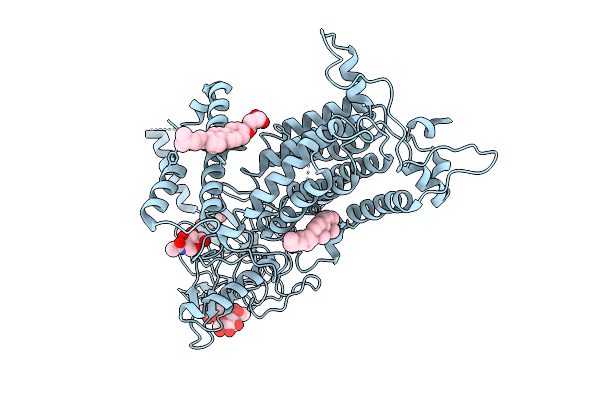
Organism: Mycobacterium tuberculosis h37rv, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: DNA BINDING PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.46 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: NAG, CLR, Y01 |
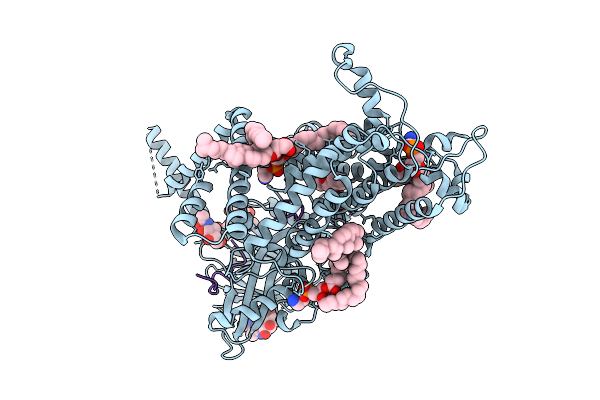 |
Cryo-Em Structure Of Vitamin K-Dependent Gamma-Carboxylase Complexed With Anisindione
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.82 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: NAG, A1AT0, CLR, PEE, Y01, BCT |
 |
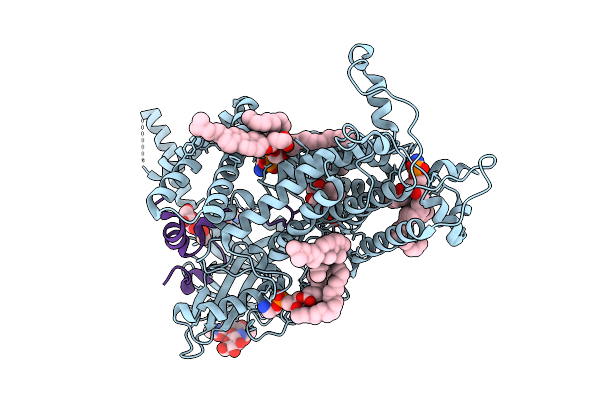
Cryo-Em Structure Of Vitamin K-Dependent Gamma-Carboxylase Complexed With Factor Ix
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.62 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: NAG, A1AVC, BCT, CO2, CLR, PEE, Y01 |
 |
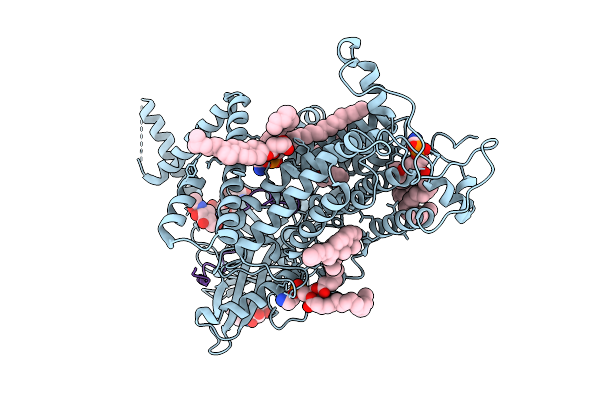
Cryo-Em Structure Of Vitamin K-Dependent Gamma-Carboxylase Complexed With Osteocalcin
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.62 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: NAG, A1AVC, CLR, PEE, Y01, CO2, BCT |
 |
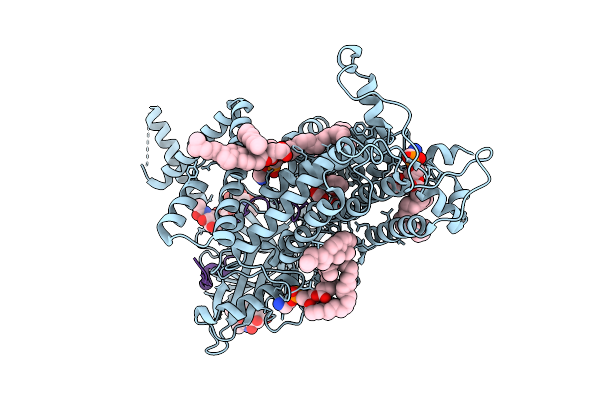
Cryo-Em Structure Of Vitamin K-Dependent Gamma-Carboxylase Complexed With Matrix Gla Protein
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: NAG, A1AVC, CLR, PEE, Y01 |
 |
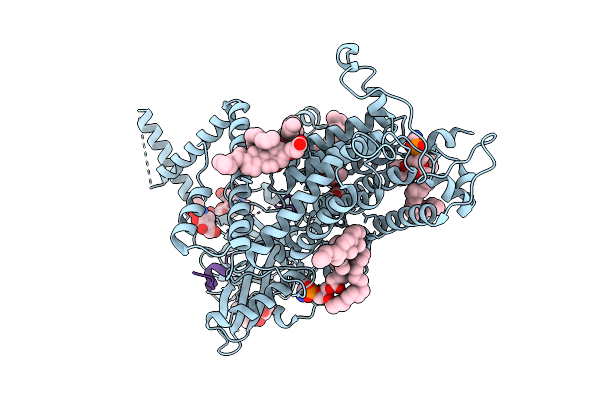
Cryo-Em Structure Of Vitamin K-Dependent Gamma-Carboxylase Complexed With Factor Ix(Gla)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.41 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: NAG, A1AVC, CLR, PEE, Y01, CO2, BCT |
 |
Cryo-Em Structure Of Vitamin K-Dependent Gamma-Carboxylase Complexed With Vitamin K1 2,3-Epoxide
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.04 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: NAG, A1EIL, CLR, PEE, Y01, BCT |
 |
Organism: Clostridium acetobutylicum
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2025-07-02 Classification: OXIDOREDUCTASE Ligands: NAP, EDO |
 |
Clostridium Acetobutylicum Alcohol Dehydrogenase Bound To Nadp+, Disordered Nicotinamide
Organism: Clostridium acetobutylicum
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-07-02 Classification: OXIDOREDUCTASE Ligands: NAP, EDO |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2024-10-16 Classification: RNA BINDING PROTEIN/RNA |
 |
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2024-10-16 Classification: HYDROLASE Ligands: PEG, WFH |
 |
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-10-16 Classification: HYDROLASE Ligands: WF9, PEG |
 |
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2024-10-16 Classification: HYDROLASE Ligands: WFU, PEG |

