Search Count: 153
All
Selected
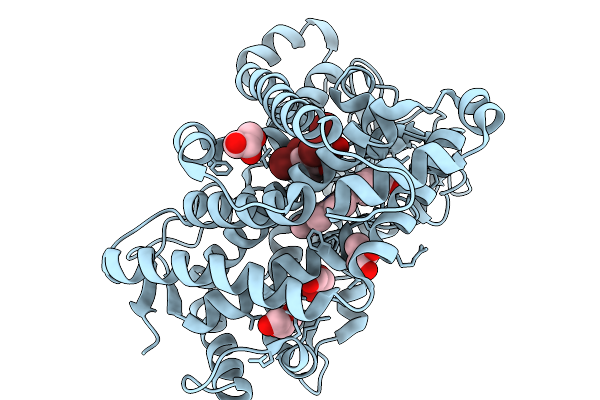 |
Crystal Structure Of Bmp7 In Complex With 2,4-Dibromophenol Generated By Substrate Soaking
Organism: Marinomonas mediterranea
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: GOL, CL, HEM, Y8I |
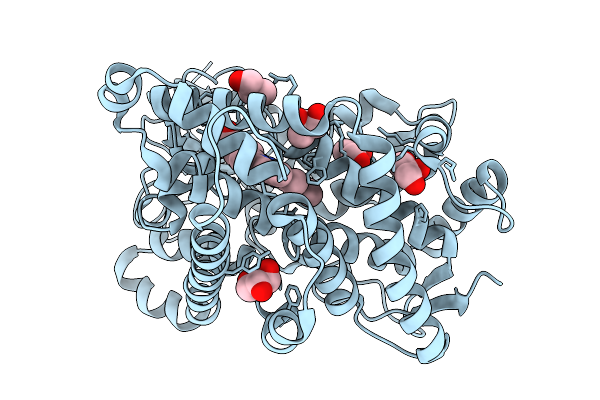 |
Organism: Marinomonas mediterranea
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: HEM, GOL |
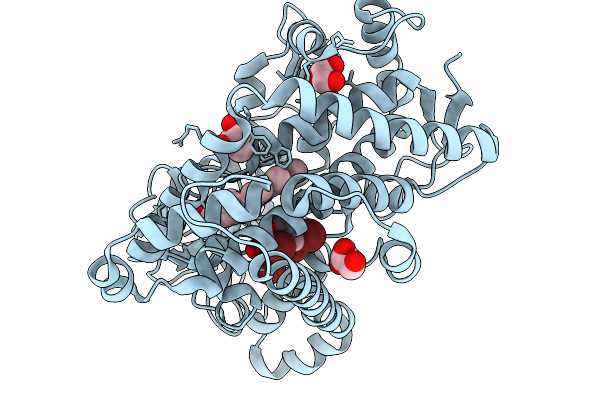 |
Organism: Marinomonas mediterranea
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: GOL, CL, HEM, Y8I |
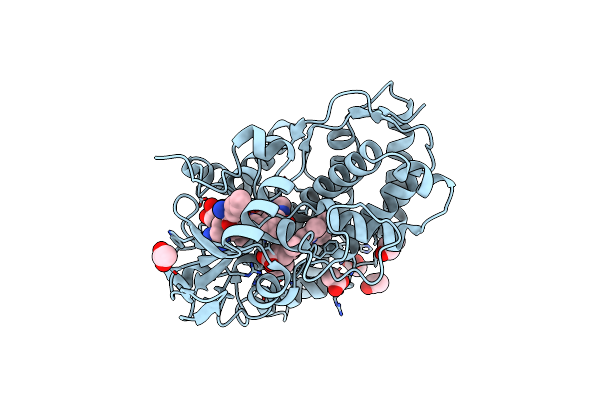 |
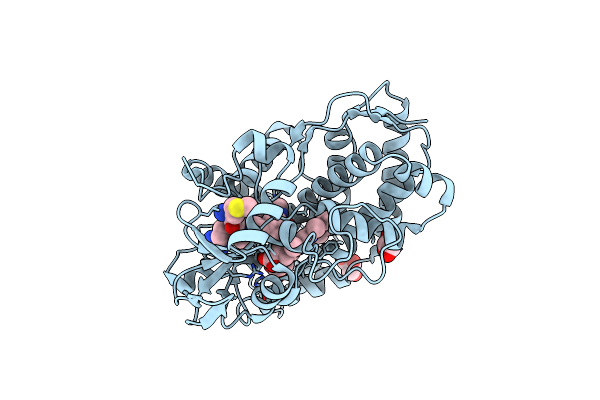
Organism: Streptomyces atratus
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2025-06-18 Classification: OXIDOREDUCTASE Ligands: HEM, CL, ACT, GOL |
 |
Organism: Streptomyces atratus
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2025-06-18 Classification: OXIDOREDUCTASE Ligands: HEM, GOL |
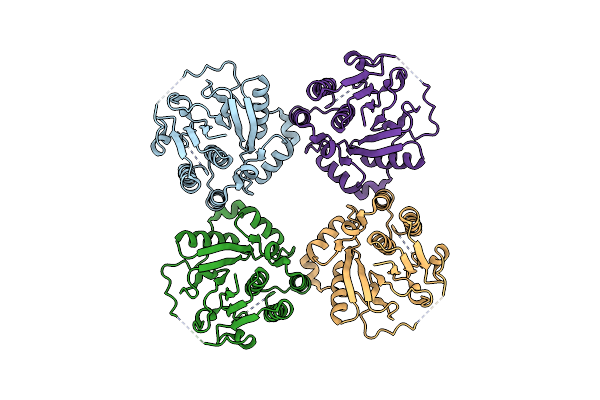 |
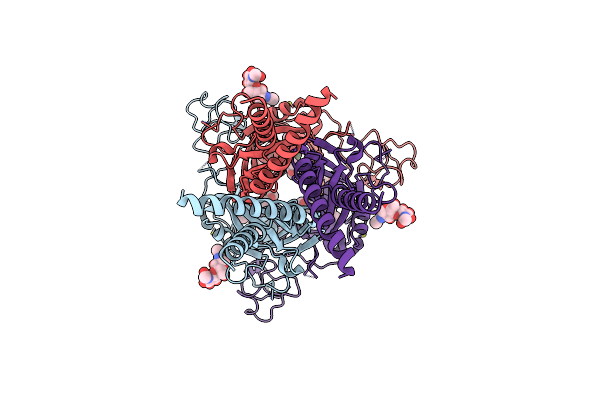
Organism: Pasteurella
Method: ELECTRON MICROSCOPY Release Date: 2025-06-11 Classification: TRANSFERASE |
 |
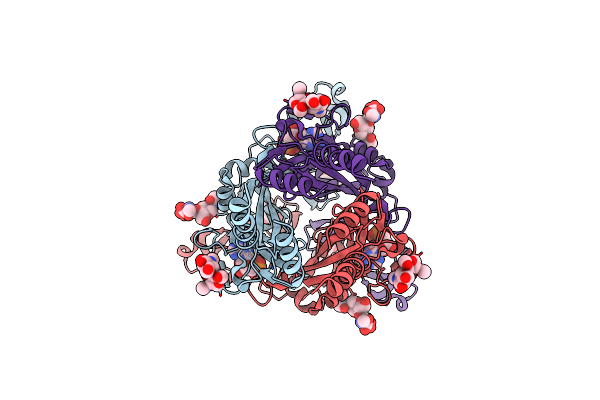
Organism: Canis lupus
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: MEMBRANE PROTEIN Ligands: A1AQX, NAG |
 |
Organism: Canis lupus
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: MEMBRANE PROTEIN Ligands: NAG, MG, ATP |
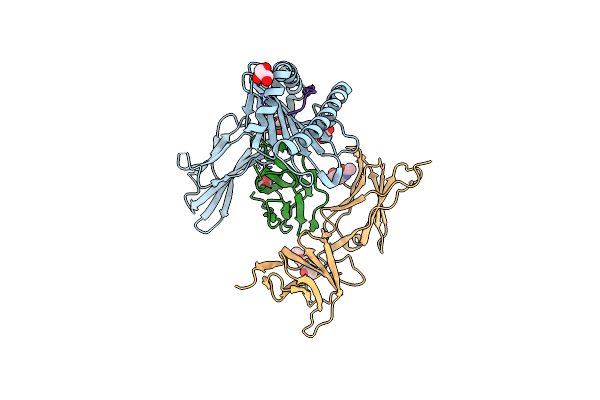 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2024-12-18 Classification: IMMUNE SYSTEM Ligands: GOL, PEG, NAG |
 |
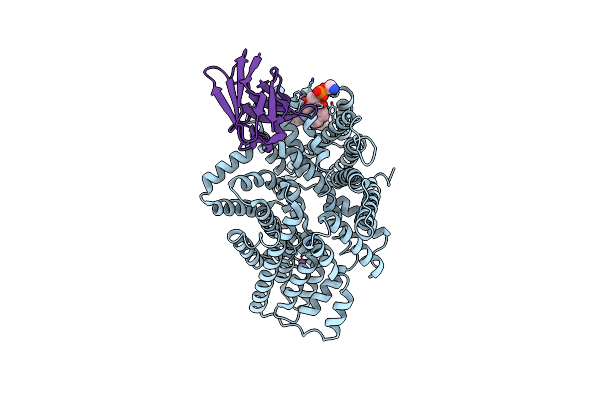
Sialic Acid Bound Form Of Tripartite Atp-Independent Periplasmic (Trap) Transporter From Fusobacterium Nucleatum.
Organism: Fusobacterium nucleatum, Vicugna pacos
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM |
 |
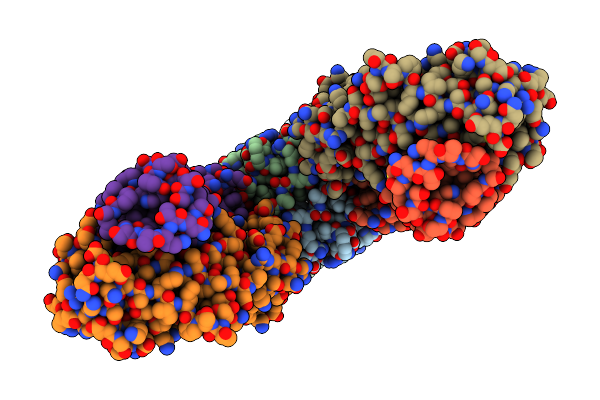
Apo Form Of Tripartite Atp-Independent Periplasmic (Trap) Transporter From Fusobacterium Nucleatum.
Organism: Fusobacterium nucleatum, Vicugna pacos
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM |
 |
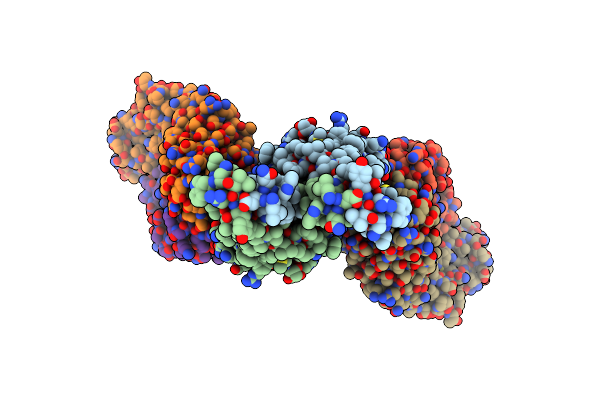
Organism: Mus musculus, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: MEMBRANE PROTEIN Ligands: 1PX, K |
 |
Cryo-Em Structure Of Sars-Cov-2 M (Long Conformation) In The Presence Of C1P
Organism: Mus musculus, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: MEMBRANE PROTEIN |
 |
Organism: Neisseria gonorrhoeae
Method: SOLID-STATE NMR Release Date: 2024-07-24 Classification: CELL ADHESION |
 |
Organism: Severe acute respiratory syndrome coronavirus 2, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.40 Å Release Date: 2024-07-10 Classification: ANTIVIRAL PROTEIN/IMMUNE SYSTEM |
 |
Organism: Synthetic construct, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2024-07-10 Classification: ANTIVIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-04 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-04 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Diploptera punctata
Method: X-RAY DIFFRACTION Resolution:2.95 Å Release Date: 2023-08-23 Classification: LIPID BINDING PROTEIN Ligands: NAG, ZN |
 |
Organism: Diploptera punctata
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2023-08-23 Classification: LIPID BINDING PROTEIN Ligands: NAG |

