Search Count: 44
All
Selected
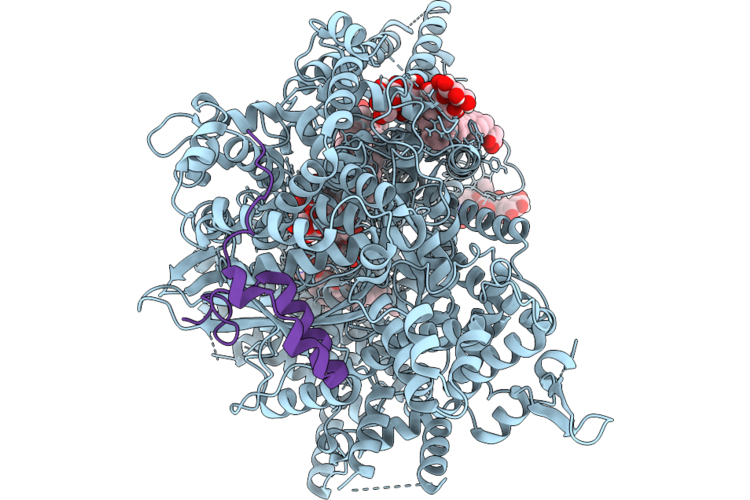 |
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) At The Catalytically Less Relevant L2 State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: Y01, 3PE, LMN |
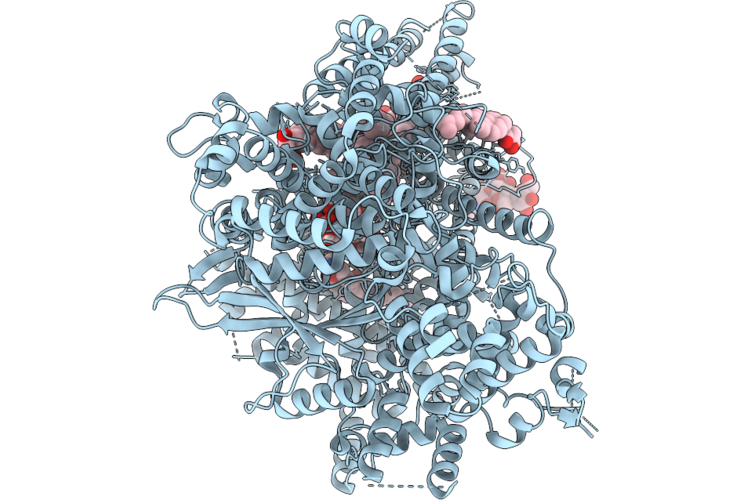 |
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) At The Catalytically Less Relevant L1 State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: Y01, 3PE, LMN |
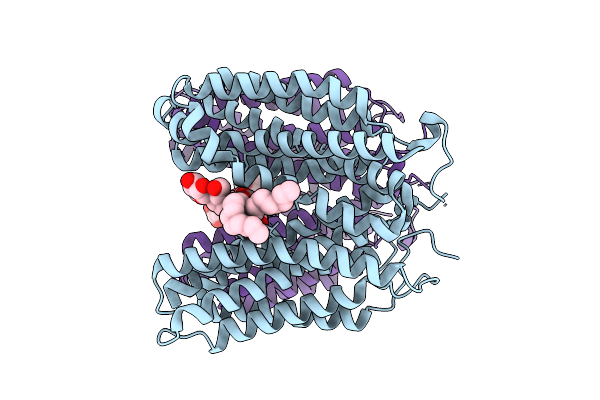 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: TRANSPORT PROTEIN Ligands: LMN |
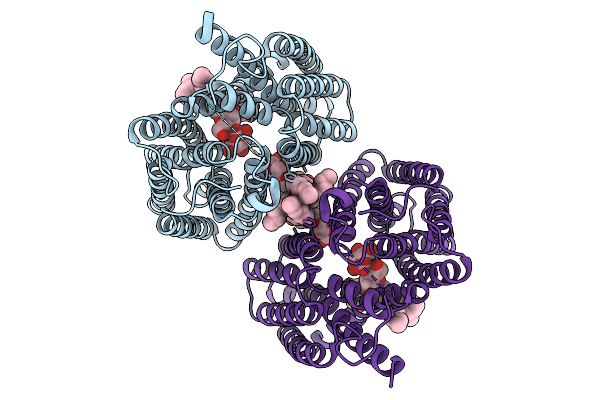 |
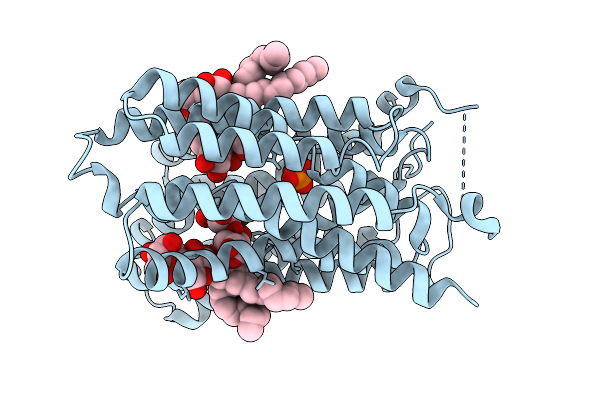
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.15 Å Release Date: 2025-12-31 Classification: TRANSPORT PROTEIN Ligands: LMN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: TRANSPORT PROTEIN Ligands: LMN, PO4 |
 |
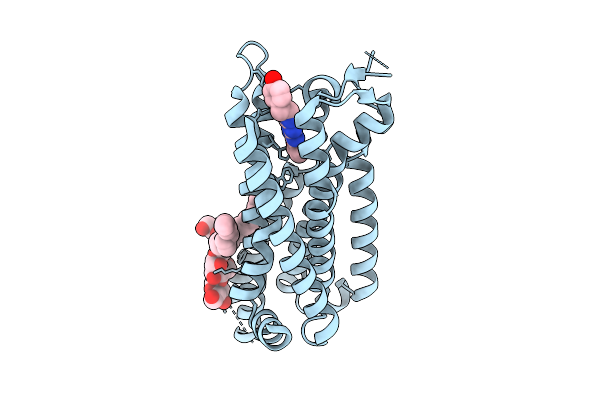
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.34 Å Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: LMN, ZMA |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.15 Å Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: LMN, ZMA |
 |
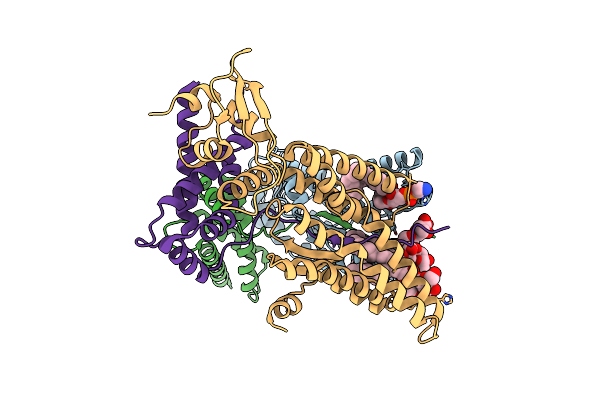
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: ELECTRON MICROSCOPY Release Date: 2024-10-02 Classification: MEMBRANE PROTEIN Ligands: LMN, PEF |
 |
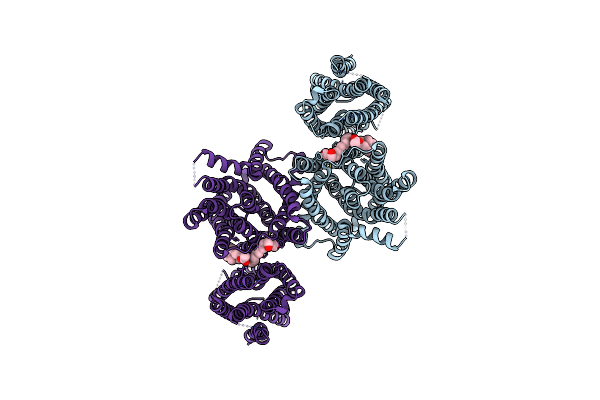
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: ELECTRON MICROSCOPY Release Date: 2024-10-02 Classification: MEMBRANE PROTEIN Ligands: LMN, PEF |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-09-18 Classification: MEMBRANE PROTEIN Ligands: LMN, Y01 |
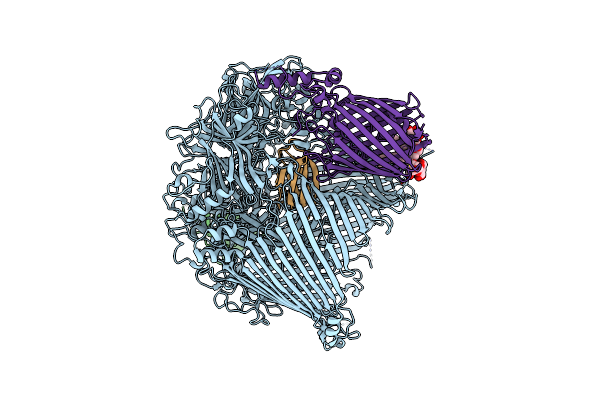 |
The Type 9 Secretion System In Vitro Assembled, Rema-Ctd Substrate Bound Complex
Organism: Flavobacterium johnsoniae
Method: ELECTRON MICROSCOPY Release Date: 2024-01-31 Classification: MEMBRANE PROTEIN Ligands: LMN |
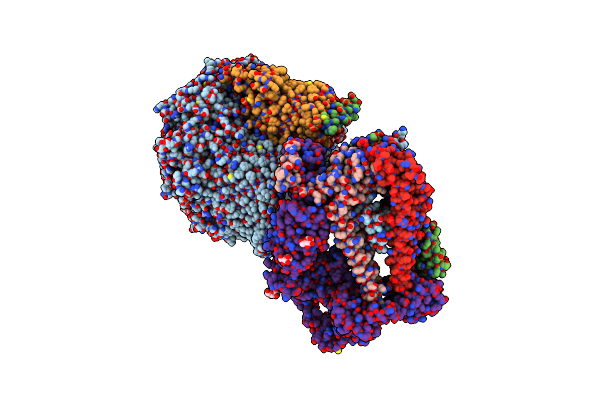 |
The Type 9 Secretion System Extended Translocon - Spra-Porv-Ppi-Remz-Skpa-Spre Complex
Organism: Flavobacterium johnsoniae
Method: ELECTRON MICROSCOPY Release Date: 2024-01-31 Classification: MEMBRANE PROTEIN Ligands: LMN, MAN |
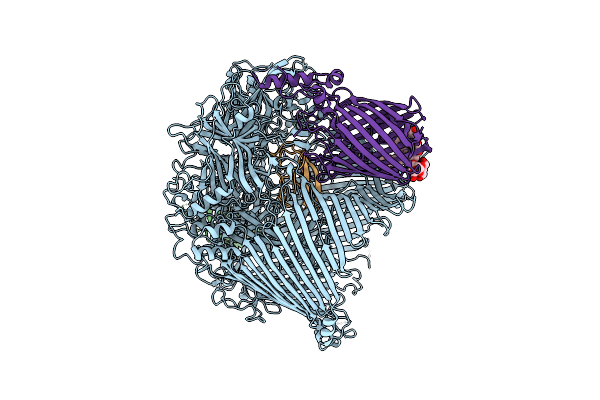 |
The Type 9 Secretion System In Vitro Assembled, Fspa-Ctd Substrate Bound Complex
Organism: Flavobacterium johnsoniae
Method: ELECTRON MICROSCOPY Release Date: 2024-01-31 Classification: MEMBRANE PROTEIN Ligands: LMN |
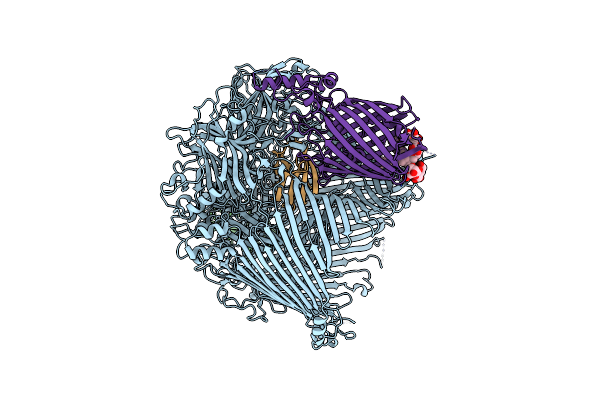 |
Organism: Flavobacterium johnsoniae
Method: ELECTRON MICROSCOPY Release Date: 2024-01-31 Classification: MEMBRANE PROTEIN Ligands: LMN |
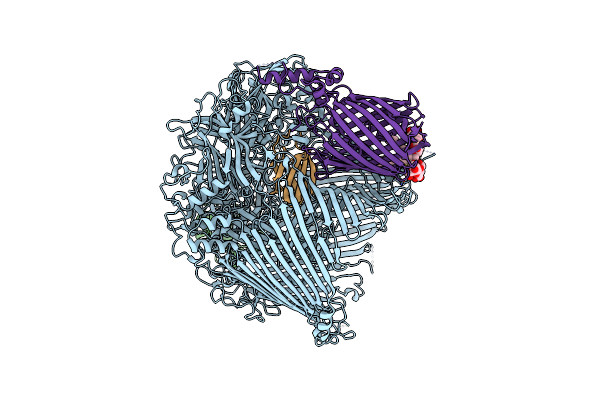 |
The Type 9 Secretion System In Vivo Assembled, Remz Substrate Bound Complex - Conformation 1
Organism: Flavobacterium johnsoniae
Method: ELECTRON MICROSCOPY Release Date: 2024-01-31 Classification: MEMBRANE PROTEIN Ligands: LMN |
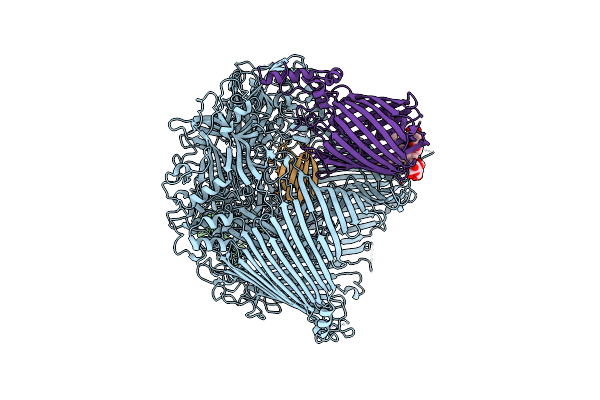 |
The Type 9 Secretion System In Vivo Assembled, Remz Substrate Bound Complex - Conformation 2
Organism: Flavobacterium johnsoniae
Method: ELECTRON MICROSCOPY Release Date: 2024-01-31 Classification: MEMBRANE PROTEIN Ligands: LMN |
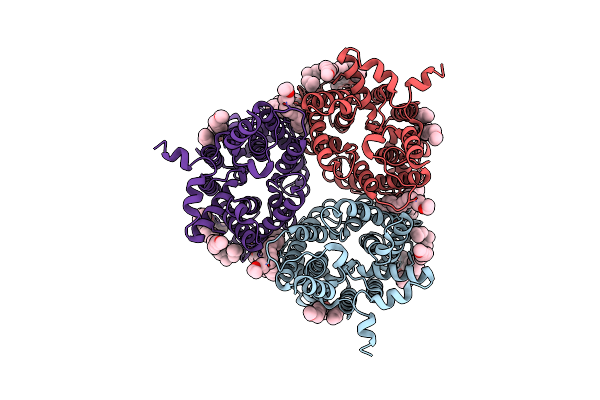 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-10-04 Classification: TRANSPORT PROTEIN Ligands: PLD, LMN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2023-04-26 Classification: MEMBRANE PROTEIN Ligands: JIZ, ATP, MG, LMN, Y01 |
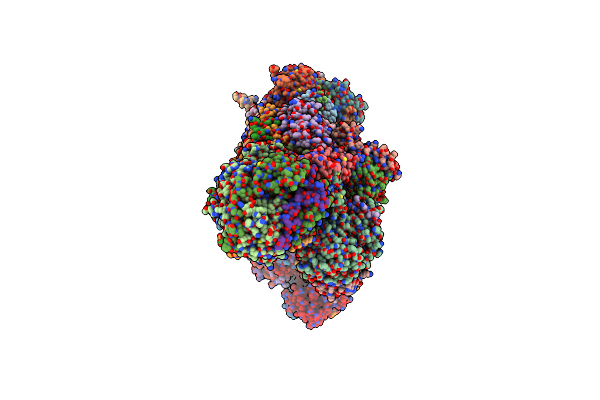 |
Cryoem Structure Of Mitochondrial Complex I From Chaetomium Thermophilum (State 2)
Organism: Chaetomium thermophilum var. thermophilum dsm 1495
Method: ELECTRON MICROSCOPY Release Date: 2022-11-30 Classification: OXIDOREDUCTASE Ligands: 3PE, PC1, CDL, LMN, FES, SF4, FMN, NDP, ZN, ZMP |
 |
Cryoem Structure Of Mitochondrial Complex I From Chaetomium Thermophilum (State 2) - Membrane Arm
Organism: Chaetomium thermophilum var. thermophilum dsm 1495
Method: ELECTRON MICROSCOPY Release Date: 2022-11-30 Classification: OXIDOREDUCTASE Ligands: 3PE, PC1, CDL, LMN, ZMP |

