Search Count: 107
All
Selected
 |
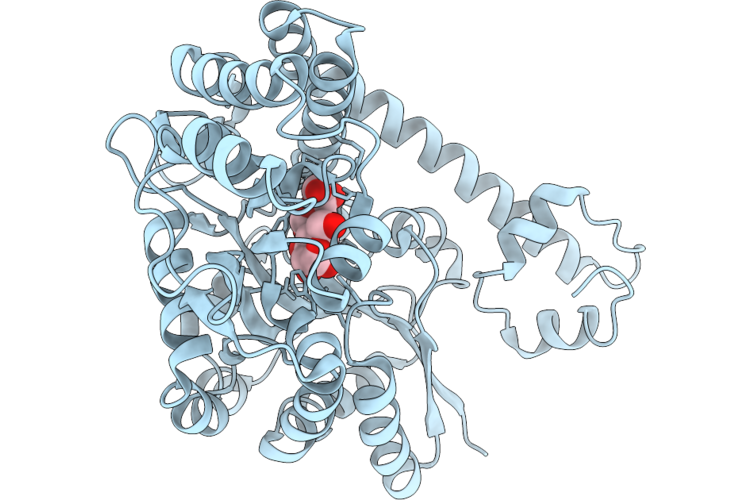
The Structure Of Egalitarian In Complex With The K10 Mrna Localization Signal Reveals A Modular Binding Surface Required For Function
Organism: Drosophila melanogaster
Method: X-RAY DIFFRACTION Resolution:3.08 Å Release Date: 2026-04-29 Classification: RNA BINDING PROTEIN Ligands: CA |
 |
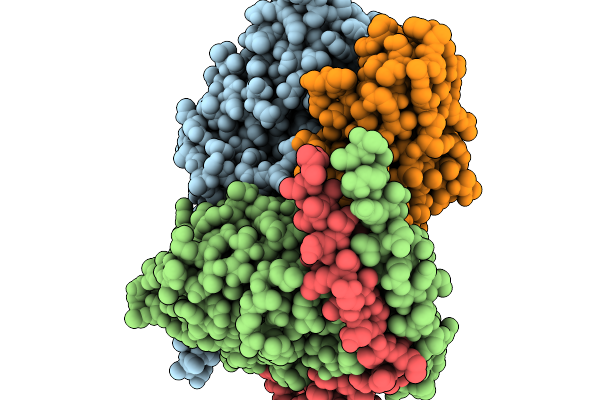
The Structure Of Egalitarian In Complex With The K10 Mrna Localization Signal Reveals A Modular Binding Surface Required For Function
Organism: Escherichia coli (strain k12), Drosophila melanogaster
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2026-04-29 Classification: RNA BINDING PROTEIN |
 |
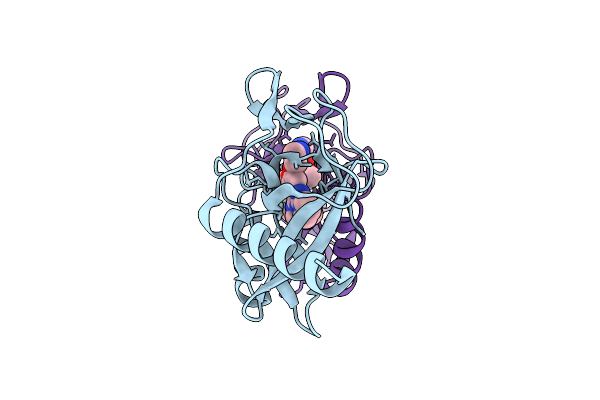
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: OR9 |
 |
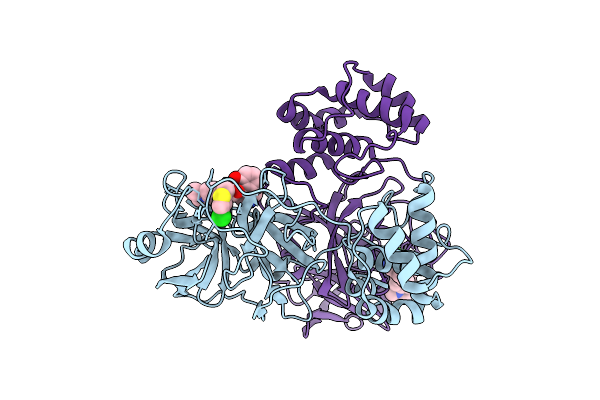
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-07-30 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: E1J, H81 |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2025-07-16 Classification: VIRAL PROTEIN, HYDROLASE Ligands: WZK |
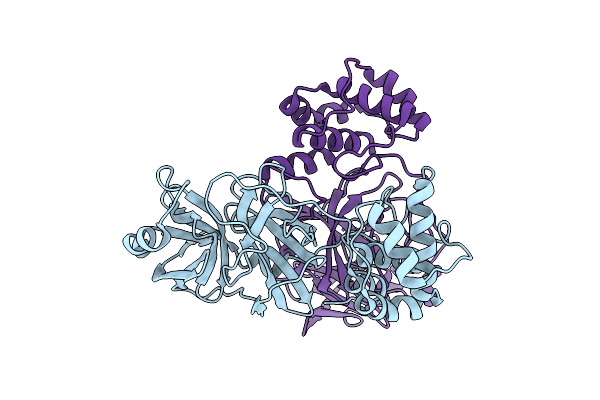 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2025-07-16 Classification: VIRAL PROTEIN, HYDROLASE |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:3.11 Å Release Date: 2025-04-30 Classification: STRUCTURAL PROTEIN |
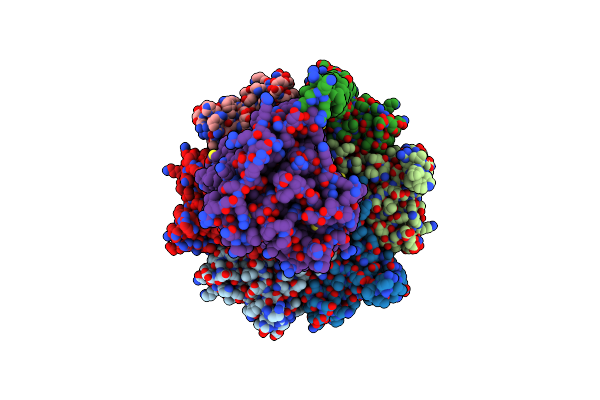 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-04-30 Classification: STRUCTURAL PROTEIN |
 |
Organism: Autographa californica nucleopolyhedrovirus
Method: ELECTRON MICROSCOPY Release Date: 2024-09-11 Classification: STRUCTURAL PROTEIN Ligands: NAG |
 |
Organism: Autographa californica nucleopolyhedrovirus
Method: ELECTRON MICROSCOPY Release Date: 2024-09-11 Classification: STRUCTURAL PROTEIN Ligands: NAG |
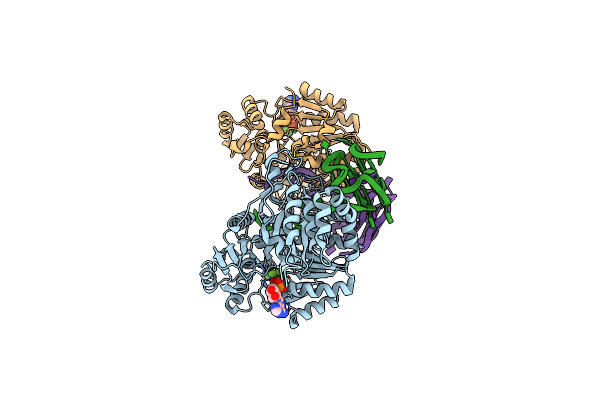 |
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2024-07-31 Classification: HYDROLASE/DNA Ligands: ADP, ALF, MG, K |
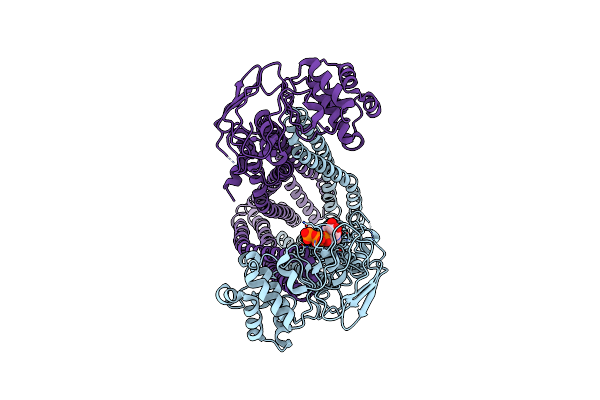 |
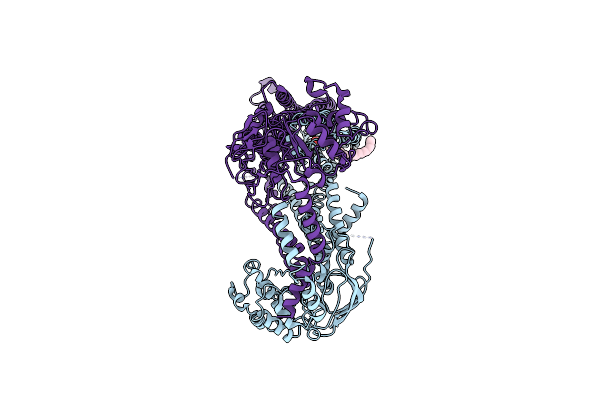
Cryo-Em Structure Of Human Abcd1 E630Q In The Presence Of Atp In Inward-Facing State 2
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-03-29 Classification: STRUCTURAL PROTEIN Ligands: ATP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-03-22 Classification: STRUCTURAL PROTEIN Ligands: 7PO |
 |
Cryo-Em Structure Of Human Abcd1 E630Q In The Presence Of C26:0-Coa And Atp
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-03-22 Classification: STRUCTURAL PROTEIN Ligands: CO8, ATP |
 |
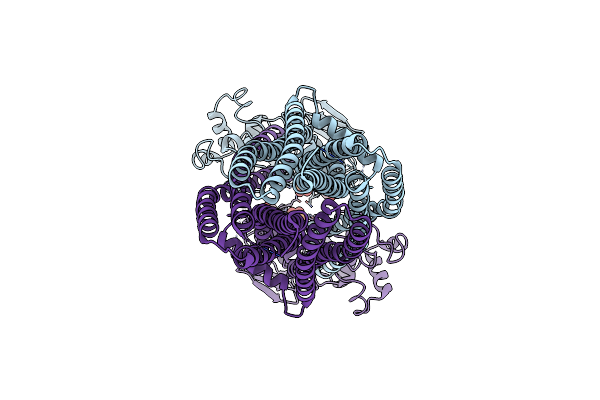
Cryo-Em Structure Of Human Abcd1 E630Q In The Presence Of Atp And Magnesium In Outward-Facing State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-03-22 Classification: STRUCTURAL PROTEIN Ligands: MG, ATP |
 |
Cryo-Em Structure Of Human Abcd1 E630Q In The Presence Of Atp In Inward-Facing State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-03-22 Classification: STRUCTURAL PROTEIN Ligands: ATP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-03-22 Classification: STRUCTURAL PROTEIN Ligands: K9G |
 |
Organism: Saccharomyces cerevisiae s288c
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2023-01-25 Classification: LIPID BINDING PROTEIN Ligands: KO0 |
 |
Organism: Saccharomyces cerevisiae s288c
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2023-01-25 Classification: LIPID BINDING PROTEIN Ligands: IUF |
 |
Organism: Saccharomyces cerevisiae s288c
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2023-01-25 Classification: LIPID BINDING PROTEIN Ligands: IUJ |

