Search Count: 622
All
Selected
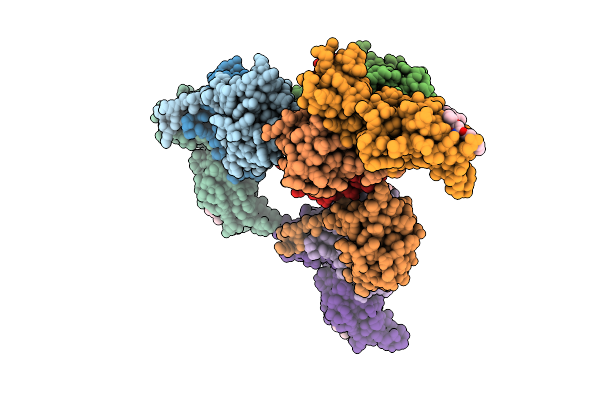 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-01 Classification: LIGASE Ligands: A1JO9 |
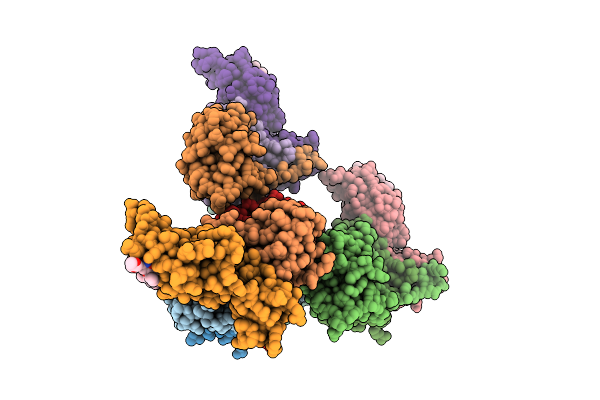 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-04-01 Classification: LIGASE Ligands: A1JO8 |
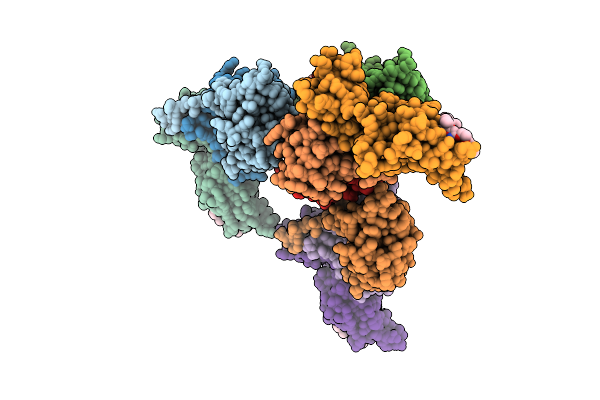 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-04-01 Classification: LIGASE Ligands: A1JO1 |
 |
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-01 Classification: ANTITOXIN |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: NAG |
 |
Cryo-Em Structure Of Human Atp Citrate Lyase In Complex With Inhibitor Evt0185-Coa
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: LYASE Ligands: ADP, W1K |
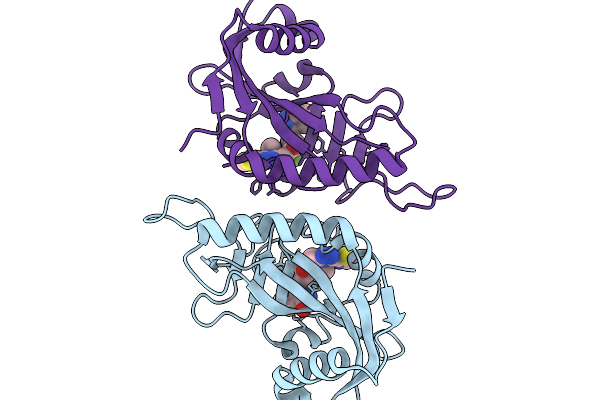 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-01-14 Classification: NUCLEAR PROTEIN Ligands: A1IFB, CL |
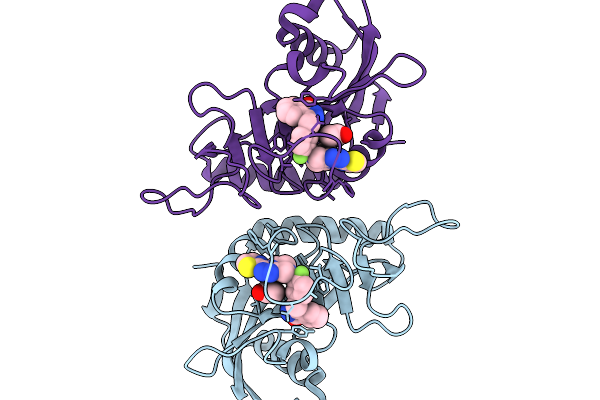 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2026-01-14 Classification: NUCLEAR PROTEIN Ligands: A1IFC, ACT, CL |
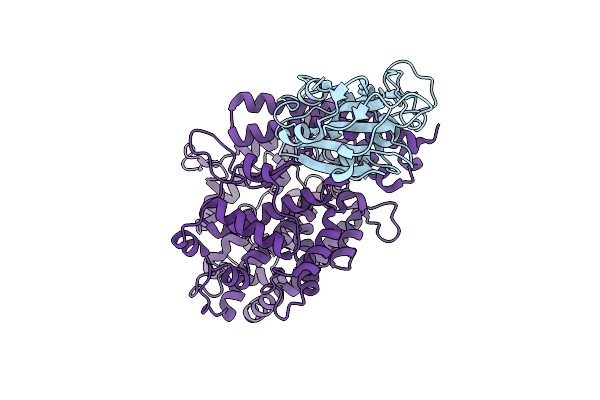 |
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: VIRAL PROTEIN/HYDROLASE |
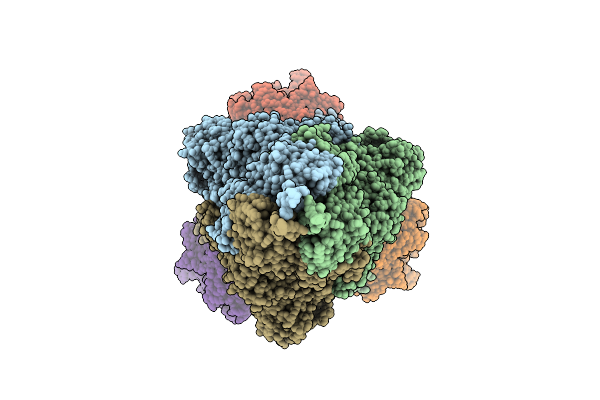 |
Organism: Homo sapiens, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: VIRAL PROTEIN/HYDROLASE |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: MEMBRANE PROTEIN Ligands: CLR |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: MEMBRANE PROTEIN Ligands: Y01, CLR |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: MEMBRANE PROTEIN |
 |
Organism: Paenibacillus sp. fsl e2-0178
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2025-11-26 Classification: RNA Ligands: GUN, MG |
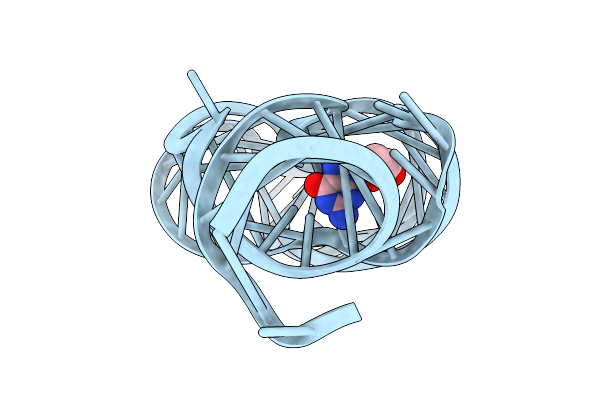 |
Crystal Structure Of Guanine-Ii Riboswitch In Complex With 2'-Deoxyguanosine
Organism: Paenibacillus sp. fsl e2-0178
Method: X-RAY DIFFRACTION Resolution:2.61 Å Release Date: 2025-11-26 Classification: RNA Ligands: MG, GNG |
 |
Organism: Paenibacillus sp. fsl e2-0178
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2025-11-26 Classification: RNA Ligands: HPA, GTP, MG |
 |
Organism: Paenibacillus sp. fsl e2-0178
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2025-11-26 Classification: RNA Ligands: GMP, MG |
 |
Crystal Structure Of Guanine-Ii Riboswitch In Complex With Guanine Soaked With Mn2+
Organism: Paenibacillus sp. fsl e2-0178
Method: X-RAY DIFFRACTION Resolution:2.97 Å Release Date: 2025-11-26 Classification: RNA Ligands: GUN, MN, GTP, MG |
 |
Crystal Structure Of Guanine-Ii Riboswitch In Complex With 7,8-Dihydroneopterin
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.54 Å Release Date: 2025-11-26 Classification: RNA Ligands: NPR, MG |
 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:3.12 Å Release Date: 2025-11-26 Classification: RNA Ligands: 7PD, MG |

