Search Count: 241
All
Selected
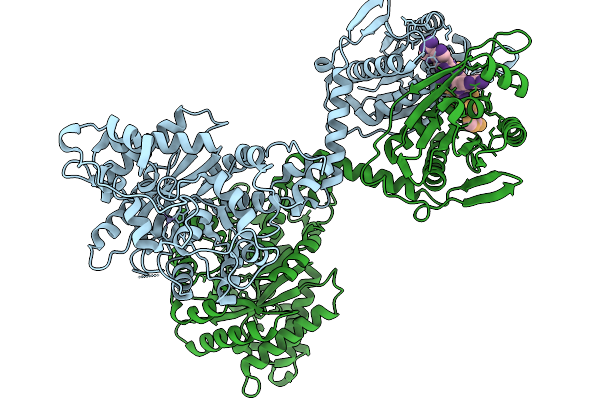 |
Crispr-Associated Deaminase Cad1 In Ca4 Bound Form, Symmetry Expanded Dimer, Refined Against A Composite Map
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: SIGNALING PROTEIN Ligands: ZN |
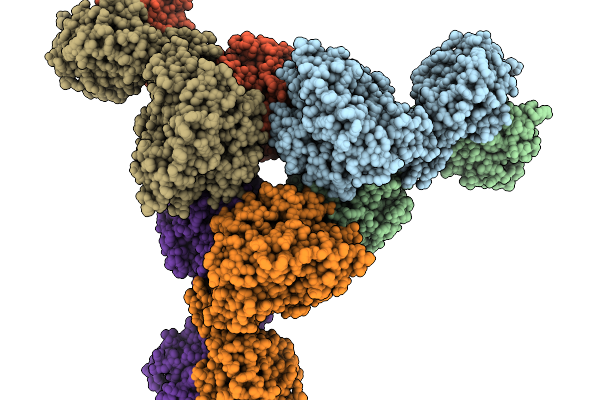 |
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: HYDROLASE Ligands: ZN |
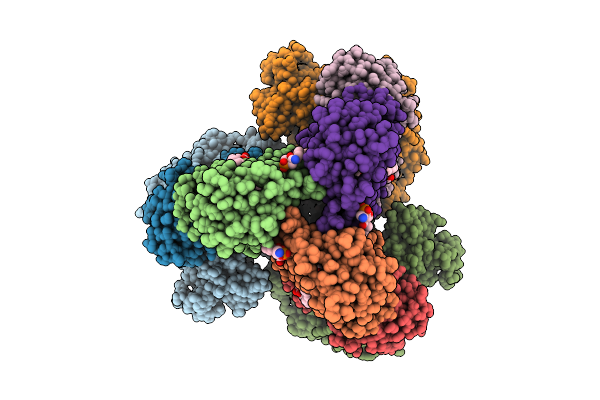 |
Cryo-Em Structure Of The Apo-Form Succinate Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SIN, SF4, FES, F3S, HEM, LMT, PGV, PEV |
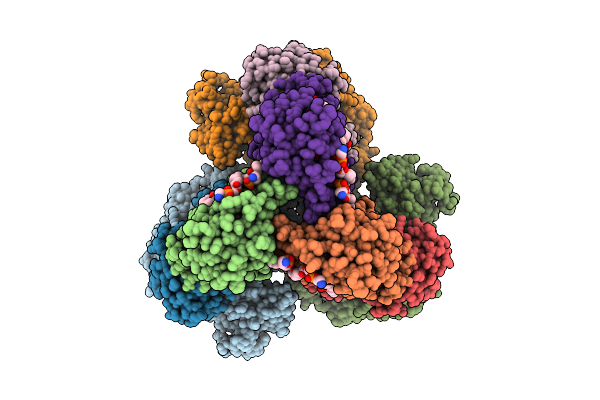 |
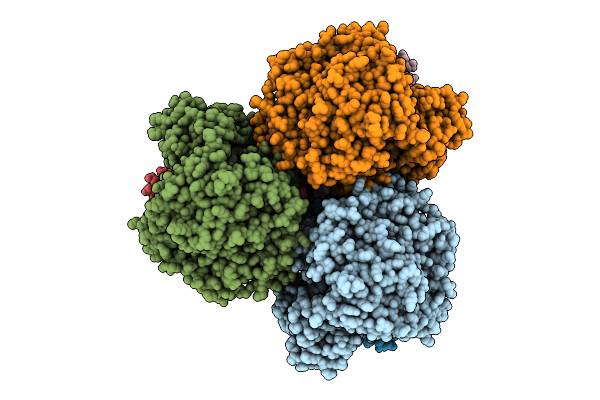
Cryo-Em Structure Of The Lipid-Bound Succiante Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SIN, SF4, FES, F3S, HEM, LMT, PGV, PEV |
 |
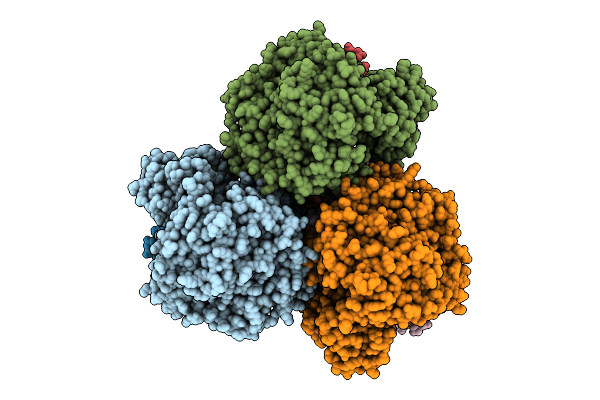
Cryo-Em Structure Of The Mk7-Bound Succinate Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SF4, FES, F3S, HEM, LMT, MQ7 |
 |
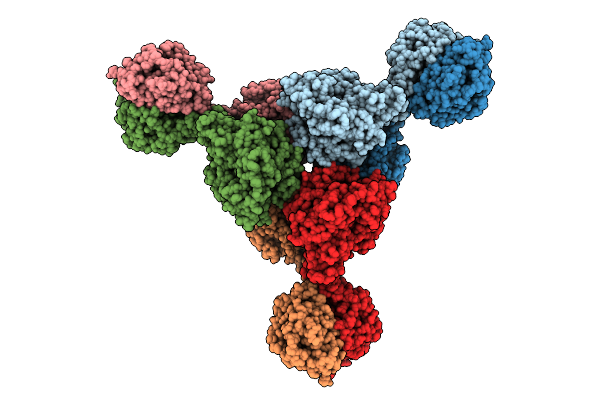
Cryo-Em Structure Of The Mk4-Bound Succinate Dehydrogenase From Chloroflexus Aurantiacus
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: MEMBRANE PROTEIN Ligands: FAD, SF4, FES, F3S, HEM, LMT, 1L3 |
 |
Crispr-Associated Deaminase Cad1 In Ca4 Bound In Hexamer Form Refined Against The Consensus Map
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: ANTIVIRAL PROTEIN/RNA Ligands: ZN |
 |
Organism: Streptomyces sp.
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: NAD |
 |
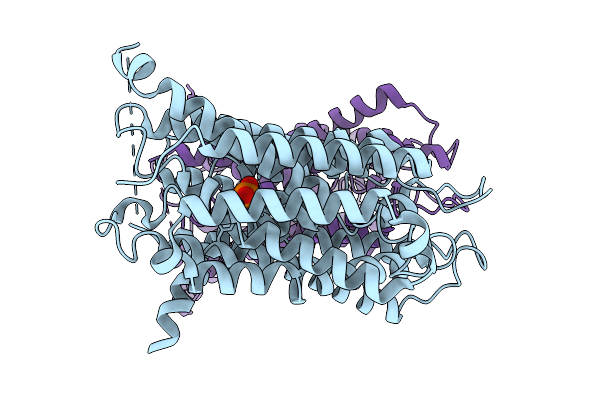
Organism: Oryza sativa subsp. japonica
Method: ELECTRON MICROSCOPY Release Date: 2025-10-08 Classification: TRANSPORT PROTEIN |
 |
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Release Date: 2025-10-08 Classification: TRANSPORT PROTEIN Ligands: PO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.16 Å Release Date: 2025-09-10 Classification: TRANSFERASE Ligands: A1INI |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2025-09-10 Classification: TRANSFERASE Ligands: A1IOI |
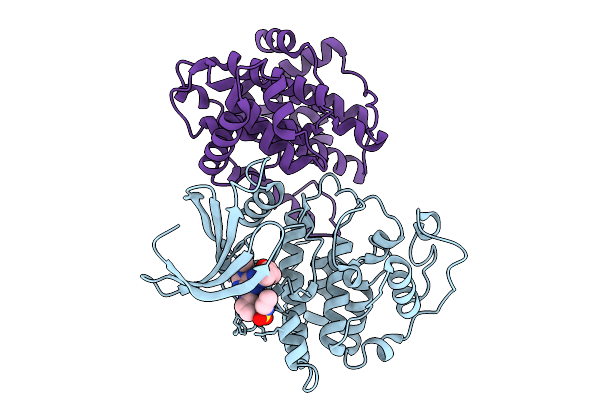 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2025-09-10 Classification: TRANSFERASE Ligands: A1IOP |
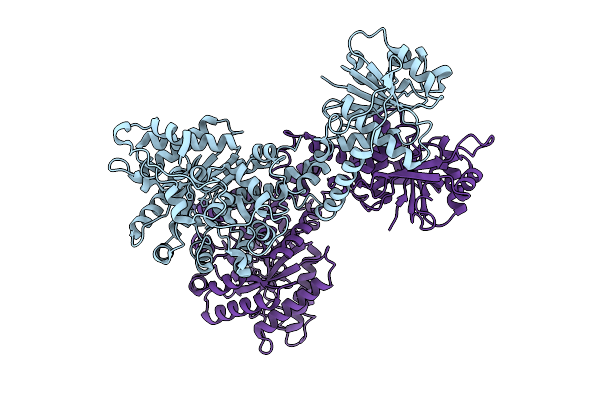 |
Cryoem Structure Of Cad1 In Apo Form, Symmetry Expanded Dimer, Refined Against A Composite Map
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: IMMUNE SYSTEM, HYDROLASE Ligands: ZN |
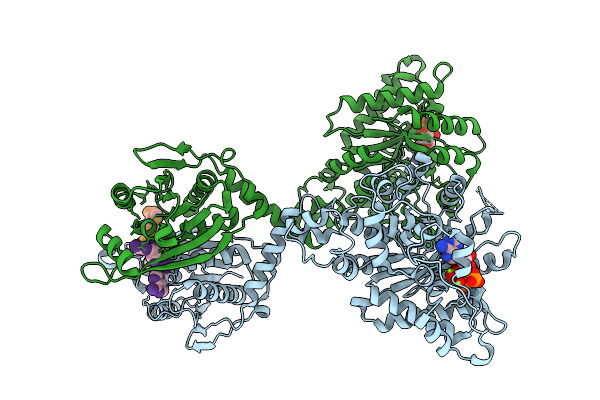 |
Cryoem Structure Of Cad1 Bound With Ca4 And Atp, Symmetry Expanded Dimer Refined Against A Composite Map
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: IMMUNE SYSTEM, HYDROLASE Ligands: ATP, ZN, MG |
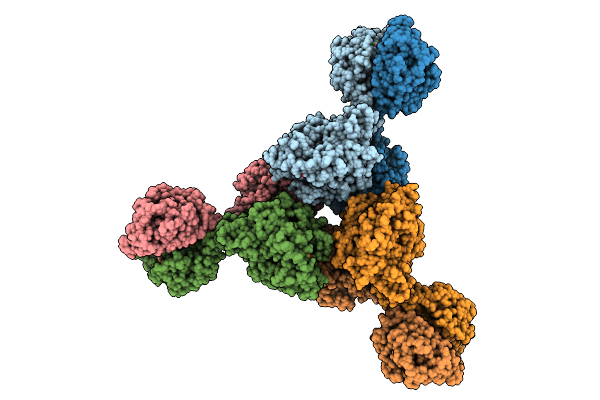 |
Cryoem Structure Of Cad1 Bound With Ca4 And Atp, Hexamer With Three Intact Dimers
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: IMMUNE SYSTEM, HYDROLASE Ligands: ATP, ZN, MG |
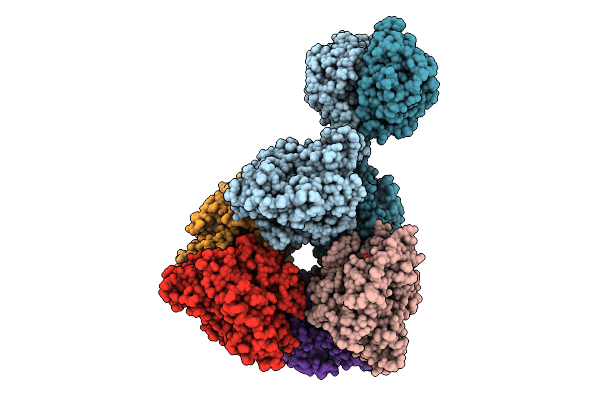 |
Cryoem Structure Of Cad1 Bound With Ca4 And Atp, Hexamer With One Intact Dimer
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: IMMUNE SYSTEM, HYDROLASE Ligands: ATP, ZN, MG |
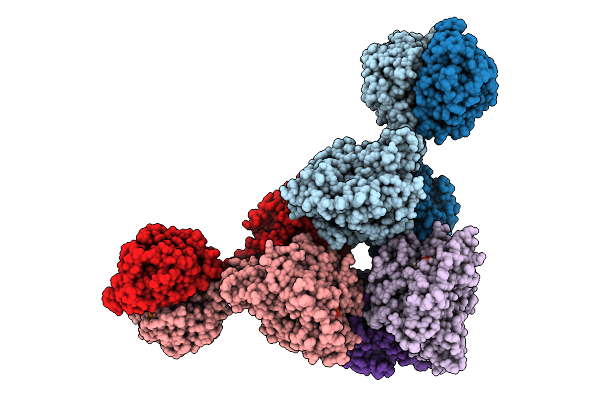 |
Cryoem Structure Of Cad1 Bound With Ca4 And Atp, Hexamer With Two Intact Dimers
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: IMMUNE SYSTEM, HYDROLASE Ligands: ATP, ZN, MG |
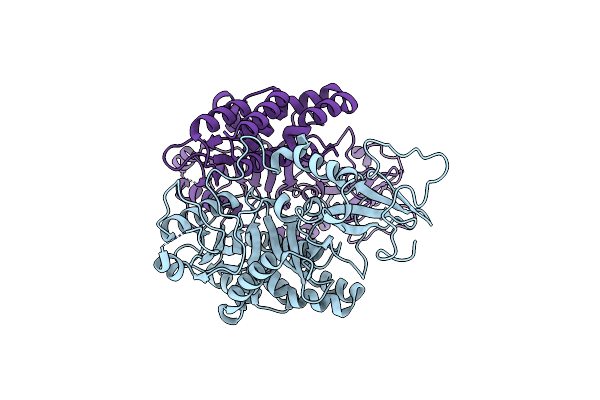 |
Organism: Pseudomonas phage vb_pa32_gums
Method: ELECTRON MICROSCOPY Release Date: 2025-07-02 Classification: UNKNOWN FUNCTION |
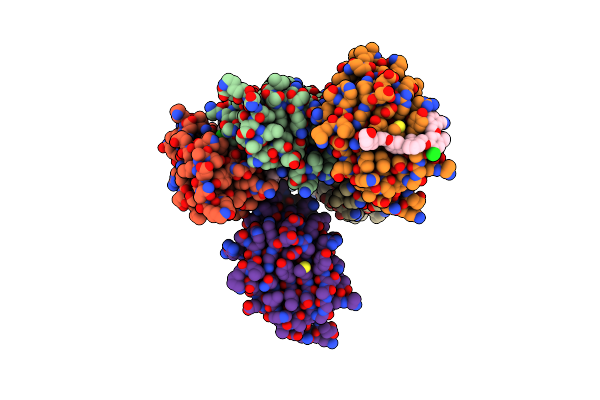 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.15 Å Release Date: 2025-06-25 Classification: IMMUNE SYSTEM Ligands: A1EJ6 |

