Search Count: 23
All
Selected
 |
Organism: Enterococcus hirae
Method: X-RAY DIFFRACTION Resolution:3.82 Å Release Date: 2022-06-22 Classification: MOTOR PROTEIN Ligands: GOL, ADP, MG, ALF |
 |
Organism: Chimpanzee adenovirus y25
Method: ELECTRON MICROSCOPY Release Date: 2021-12-15 Classification: VIRUS |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.19 Å Release Date: 2021-09-01 Classification: MEMBRANE PROTEIN Ligands: UQ8, LHG, 3PE, HEO, HEM, CDL, CU, ZN |
 |
2.55-Angstrom Cryo-Em Structure Of Cytochrome Bo3 From Escherichia Coli In Native Membrane
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2021-08-25 Classification: OXIDOREDUCTASE Ligands: HEM, HEO, CU, 3PE, UQ8 |
 |
2.55-Angstrom Cryo-Em Structure Of Cytochrome Bo3 From Escherichia Coli In Native Membrane
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2021-08-25 Classification: OXIDOREDUCTASE Ligands: HEM, HEO, CU, 3PE, UQ8 |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2021-08-25 Classification: OXIDOREDUCTASE Ligands: HEM, HEO, CU, 3PE, UQ8 |
 |
Organism: Chimpanzee adenovirus y25
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2021-06-23 Classification: VIRAL PROTEIN Ligands: CA, EDO, TRS |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:4.10 Å Release Date: 2018-08-01 Classification: DNA BINDING PROTEIN |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2018-08-01 Classification: DNA BINDING PROTEIN |
 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.82 Å Release Date: 2017-02-08 Classification: TRANSCRIPTION |
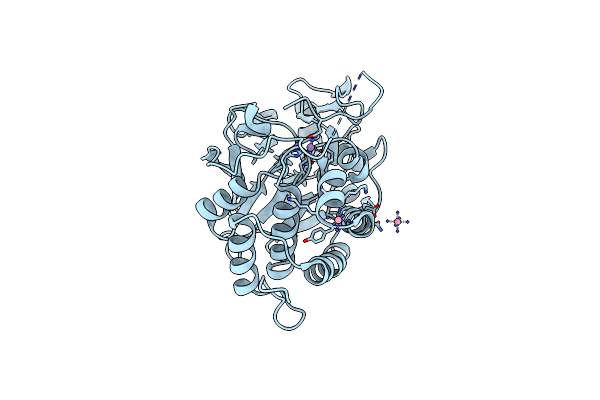 |
Phosphatase Domain Of Pp5 Bound To A Phosphomimetic Cdc37 Substrate Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2016-07-27 Classification: HYDROLASE Ligands: MN, NCO |
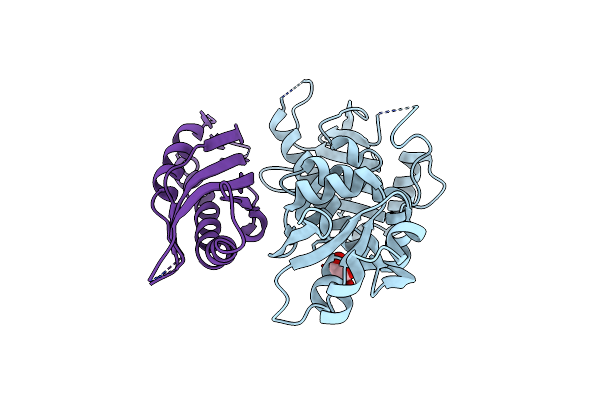 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2004-01-29 Classification: CHAPERONE Ligands: GOL |
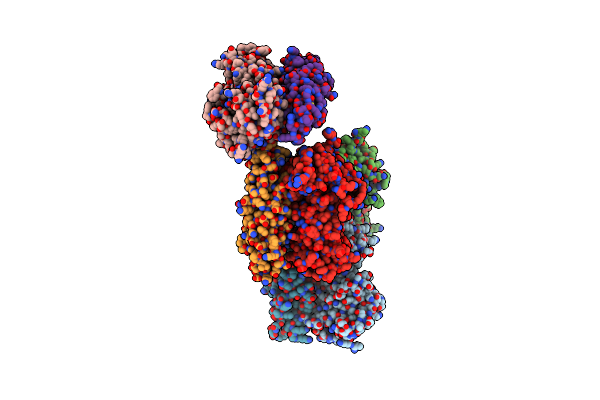 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2004-01-29 Classification: CHAPERONE |
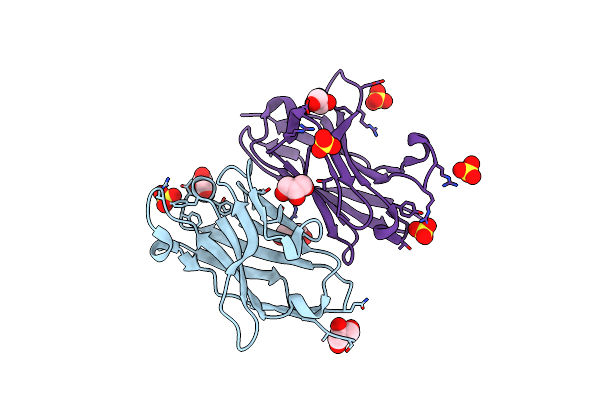 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2004-01-22 Classification: IMMUNOGLOBULIN Ligands: SO4, GOL |
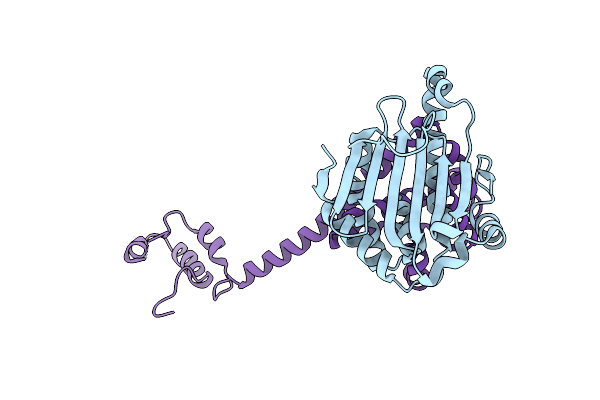 |
Organism: Saccharomyces cerevisiae, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2004-01-15 Classification: CHAPERONE |
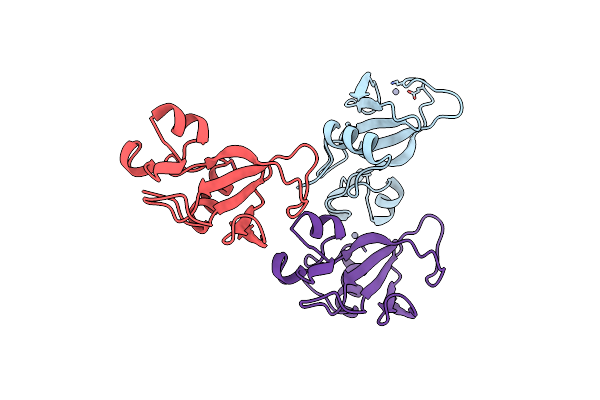 |
Organism: Bacillus amyloliquefaciens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 1998-12-09 Classification: HYDROLASE Ligands: ZN |
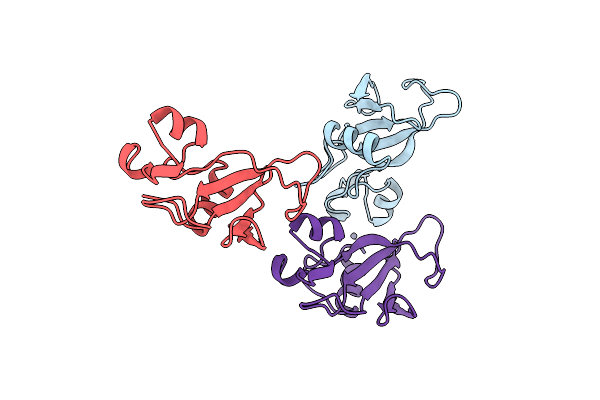 |
Organism: Bacillus amyloliquefaciens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 1998-12-09 Classification: HYDROLASE Ligands: ZN |
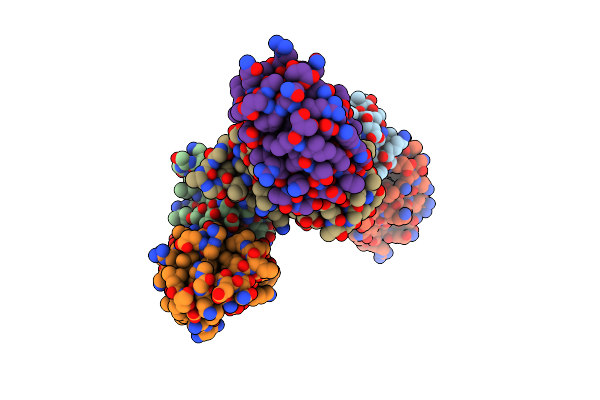 |
Organism: Bacillus amyloliquefaciens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 1998-12-09 Classification: HYDROLASE/HYDROLASE INHIBITOR |
 |
Organism: Bacillus amyloliquefaciens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 1998-12-09 Classification: HYDROLASE/HYDROLASE INHIBITOR |
 |
Organism: Bacillus amyloliquefaciens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 1998-12-09 Classification: HYDROLASE Ligands: ZN |

