Search Count: 278
All
Selected
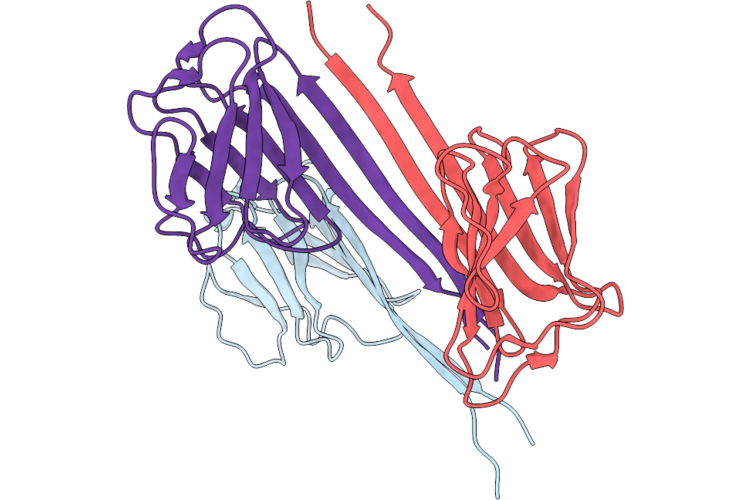 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-29 Classification: NUCLEAR PROTEIN |
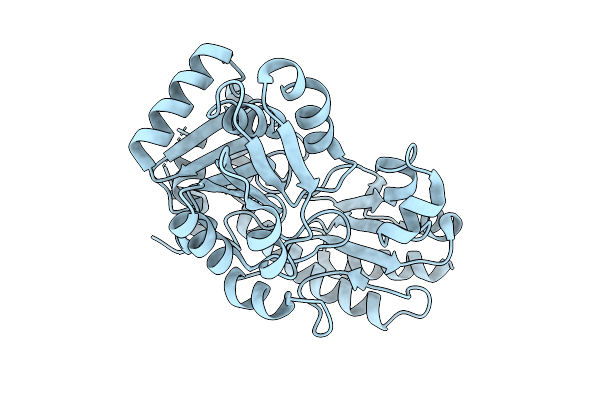 |
Organism: Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:2.33 Å Release Date: 2026-04-15 Classification: LYASE |
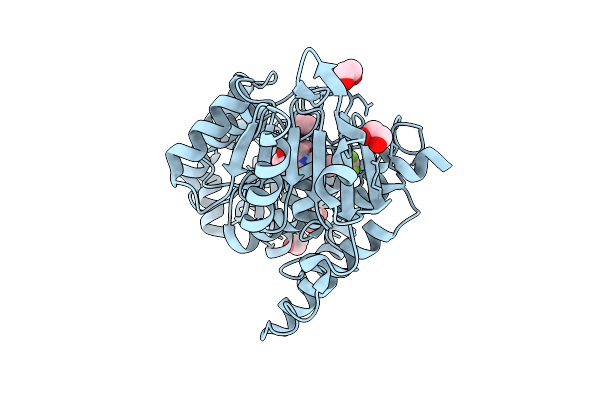 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-03-11 Classification: TRANSFERASE Ligands: A1I5F, EDO, ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:4.74 Å Release Date: 2026-01-28 Classification: PROTEIN FIBRIL |
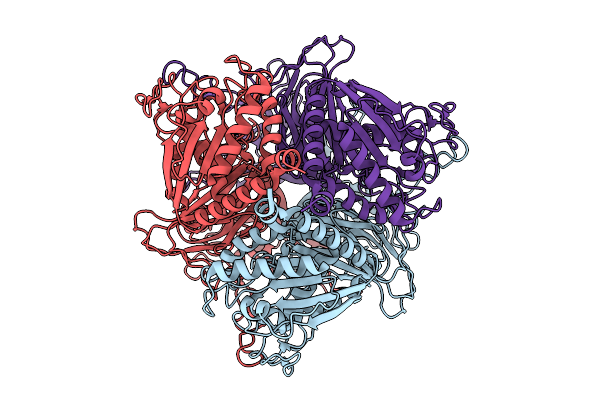 |
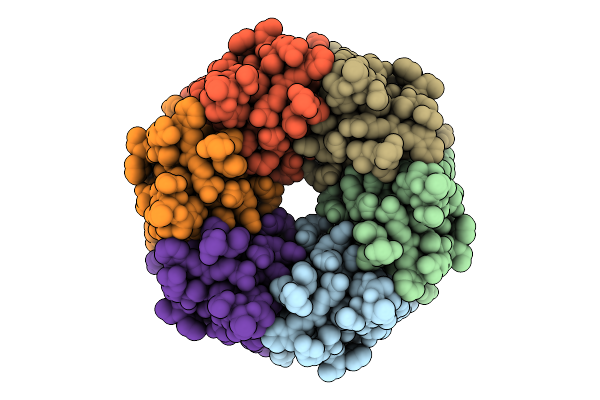
Organism: Shigella phage sfp21a
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-01-28 Classification: VIRAL PROTEIN |
 |
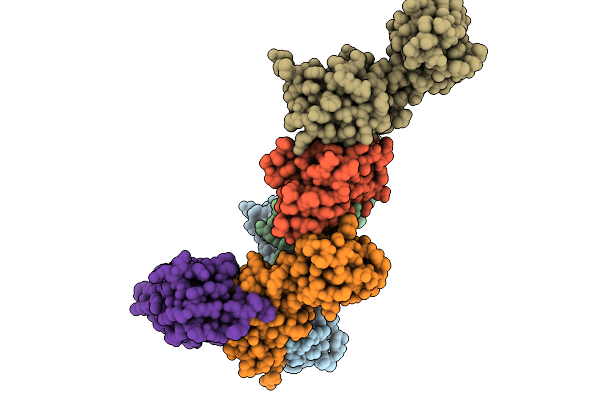
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: ELECTRON MICROSCOPY Resolution:3.05 Å Release Date: 2025-11-12 Classification: STRUCTURAL PROTEIN |
 |
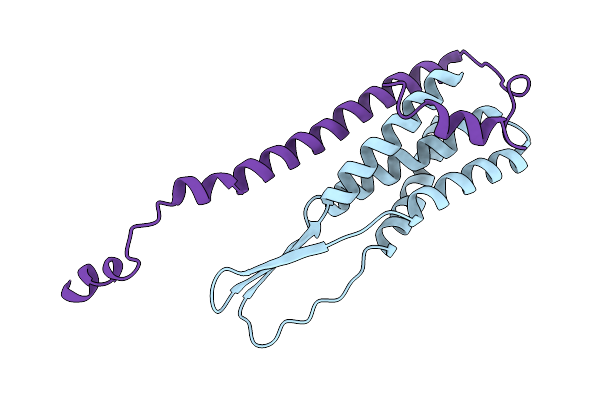
Organism: Homo sapiens, Vicugna pacos
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2025-10-01 Classification: IMMUNE SYSTEM Ligands: EDO, SO4 |
 |
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: ELECTRON MICROSCOPY Resolution:3.32 Å Release Date: 2025-09-24 Classification: STRUCTURAL PROTEIN |
 |
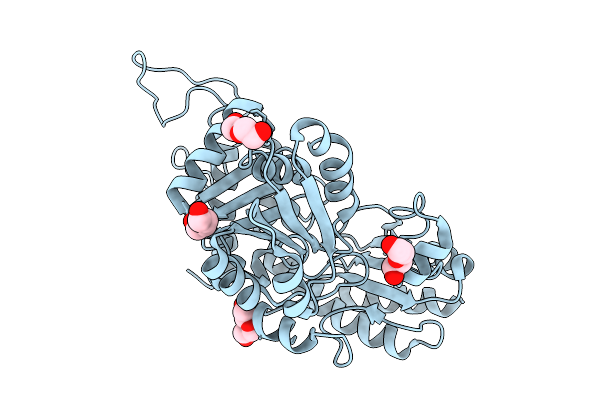
Organism: Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2025-09-03 Classification: LYASE Ligands: SO4 |
 |
Organism: Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2025-09-03 Classification: LYASE Ligands: PEG |
 |
Organism: Staphylococcus aureus subsp. aureus mu50
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-09-03 Classification: LYASE Ligands: PGU |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN Ligands: A1IIE |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN Ligands: ZN, A1IIF |
 |
Organism: Danio rerio
Method: ELECTRON MICROSCOPY Release Date: 2025-07-09 Classification: MEMBRANE PROTEIN Ligands: A1IGW |
 |
Organism: Danio rerio
Method: ELECTRON MICROSCOPY Release Date: 2025-07-09 Classification: MEMBRANE PROTEIN Ligands: A1IGY |
 |
Organism: Proteus vulgaris
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-07-02 Classification: ANTITOXIN/DNA BINDING PROTEIN/DNA Ligands: MG, CL |
 |
Organism: Proteus vulgaris
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-07-02 Classification: ANTITOXIN/DNA BINDING PROTEIN/DNA Ligands: MG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-01-15 Classification: PROTEIN FIBRIL |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-01-15 Classification: PROTEIN FIBRIL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.98 Å Release Date: 2025-01-01 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: A1AFR, GOL, SO4 |

