Search Count: 58
All
Selected
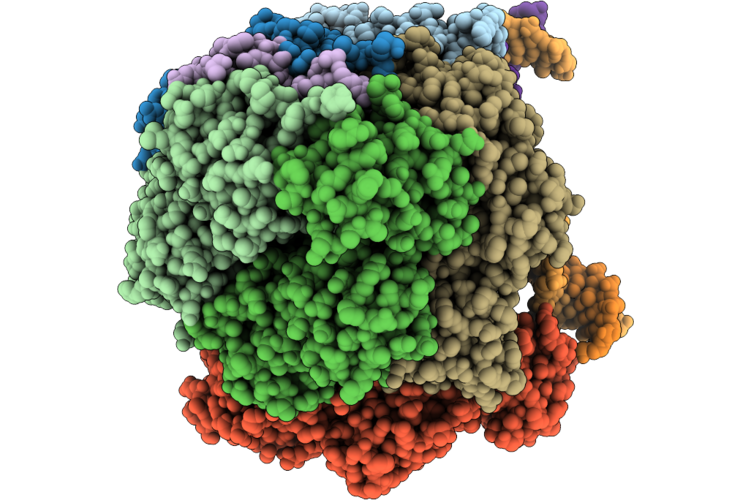 |
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 1-Nt Gapped Dna In State 1 Conformer 1 With Flexibly Bound Dna At The Shoulder
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:3.14 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
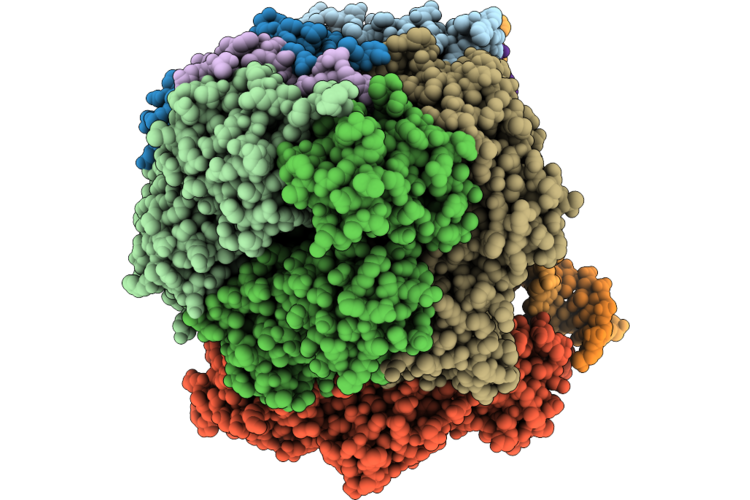 |
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 1-Nt Gapped Dna In State 1 Conformer 2 With Flexibly Bound Dna At The Shoulder
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.71 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
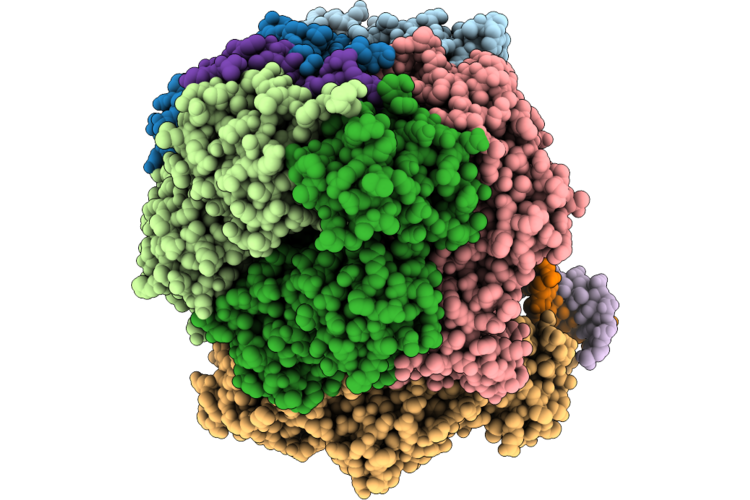 |
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 1-Nt Gapped Dna In State 1 Conformer 3 With Disordered Dna At The Shoulder
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.73 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
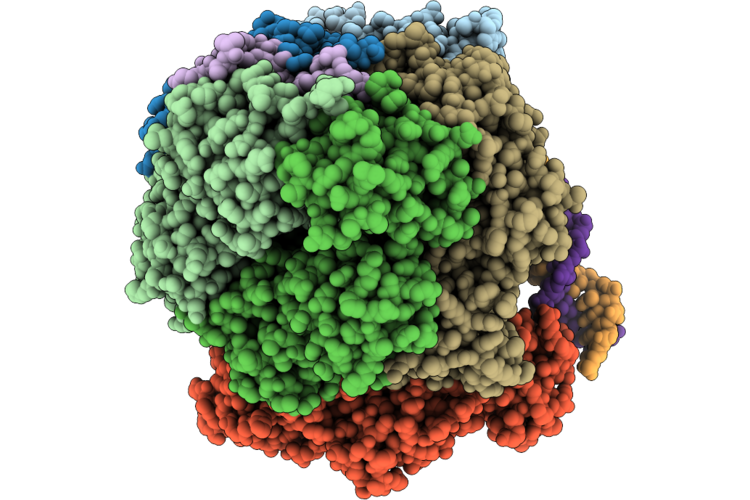 |
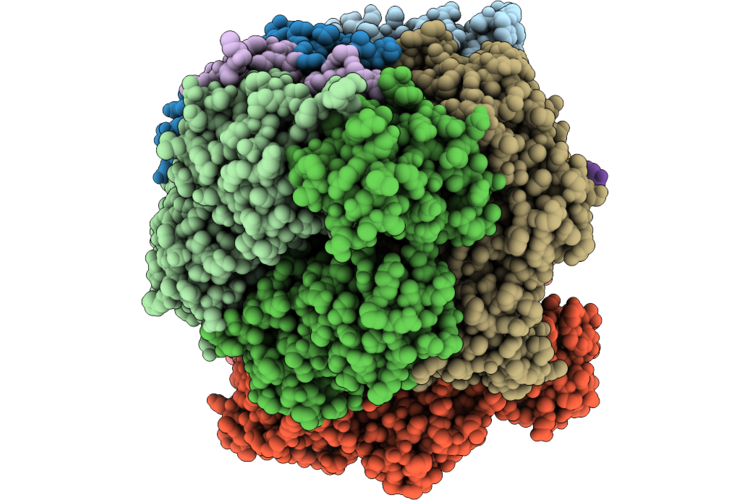
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 1-Nt Gapped Dna In State 2 Conformer 1 With Sharply Bent Dna
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.72 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 1-Nt Gapped Dna In State 2 Conformer 2 With Less Bent Dna
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:3.12 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.95 Å Release Date: 2026-04-29 Classification: TRANSFERASE Ligands: ZN, ADP, AGS, MG |
 |
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 1 The Dna Recognition State
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.54 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
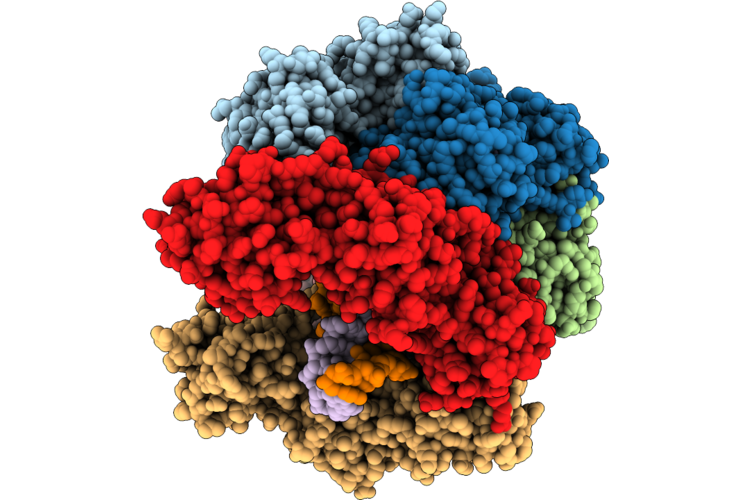
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 2 Conformer 1 With Fully Open Clamp And Unsettled Dna
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.54 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
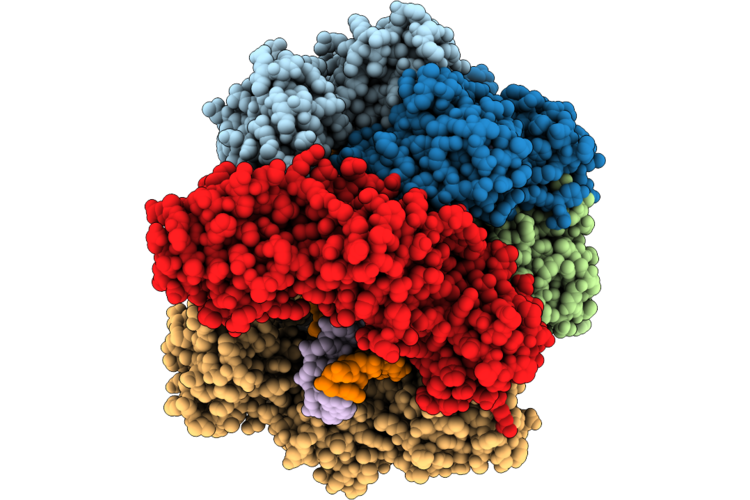
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 2 Conformer 2 With Fully Open Clamp And Settled Dna
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.53 Å Release Date: 2026-04-29 Classification: REPLICATION Ligands: ZN, AGS, MG |
 |
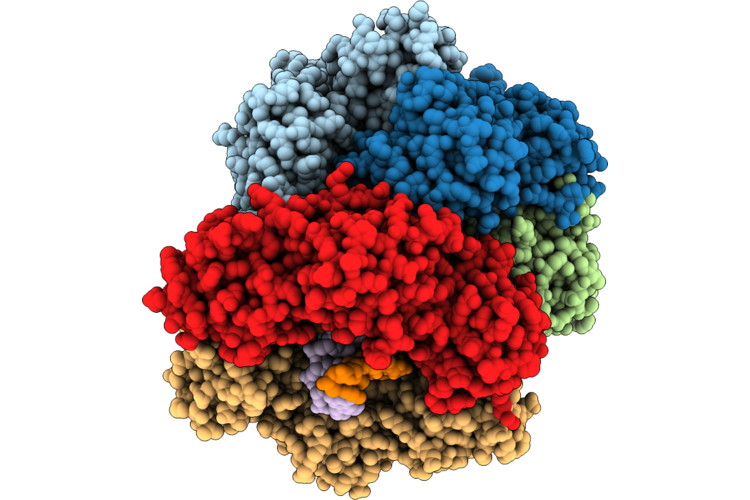
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 2 Conformer 3 With Partially Closed Clamp
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
 |
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 2 Conformer 4 With Fully Closed Clamp
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.88 Å Release Date: 2026-04-29 Classification: REPLICATION Ligands: ZN, AGS, MG |
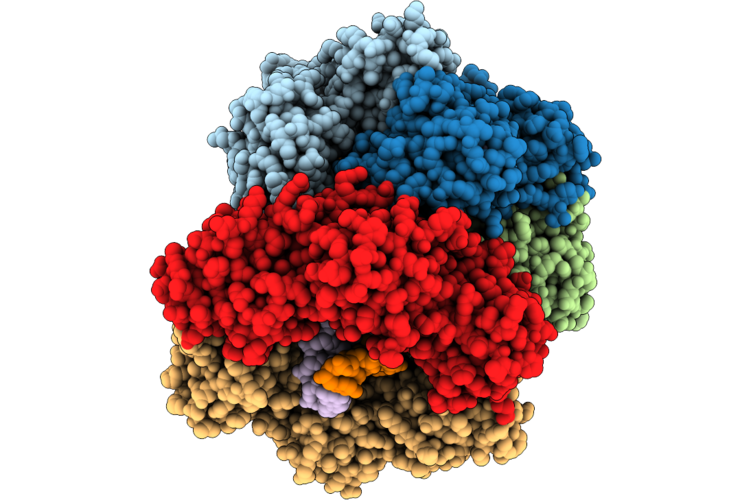 |
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 2 Conformer 5 With Fully Closed Clamp
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
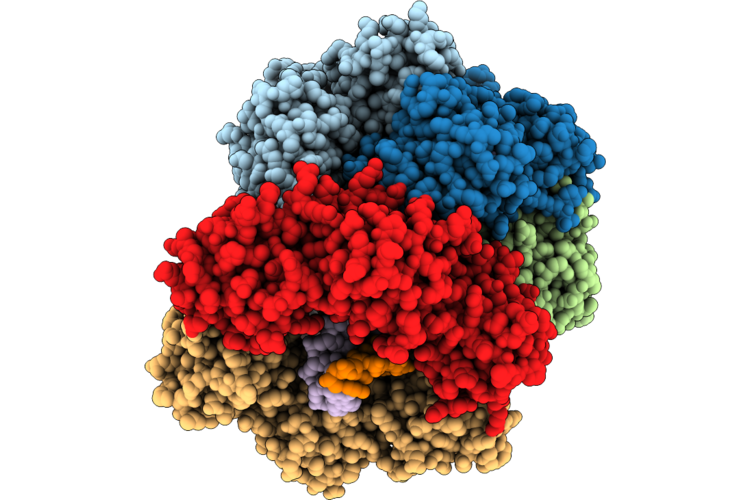 |
Structure Of The E. Coli Clamp Loader Dnax-Complex Loading Beta-Clamp Onto 10-Nt Gapped Dna In State 2 Conformer 6 With Fully Closed Clamp
Organism: Escherichia coli, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-04-29 Classification: REPLICATION/DNA Ligands: ZN, AGS, MG |
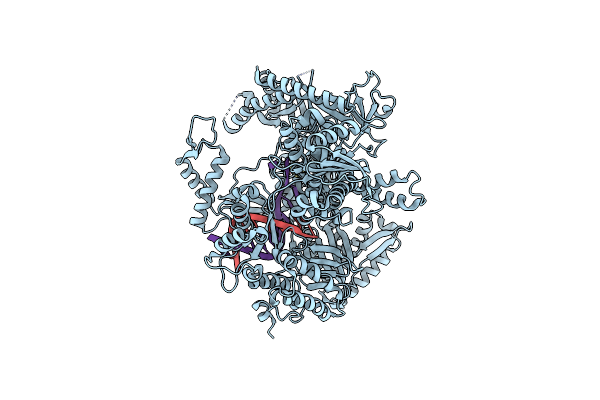 |
Organism: Saccharomyces cerevisiae, Dna molecule
Method: ELECTRON MICROSCOPY Release Date: 2024-10-09 Classification: DNA BINDING PROTEIN/DNA Ligands: SF4 |
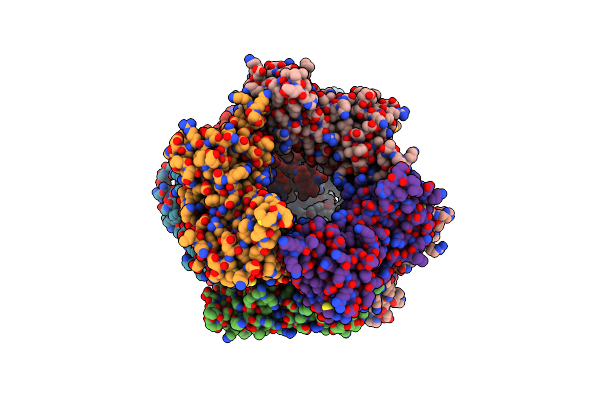 |
Cryo-Em Structure Of S. Cerevisiae Ctf18-Rfc-Pcna Complex In Apo State Conformation I
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2024-09-04 Classification: DNA BINDING PROTEIN/DNA Ligands: ADP, AGS, MG |
 |
Cryo-Em Structure Of S. Cerevisiae Ctf18-Rfc-Pcna Complex In Apo State Conformation I
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2024-09-04 Classification: DNA BINDING PROTEIN/DNA Ligands: AGS, MG, ADP |
 |
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-09-04 Classification: DNA BINDING PROTEIN/DNA Ligands: SF4 |
 |
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-09-04 Classification: DNA BINDING PROTEIN/DNA Ligands: SF4, AGS, MG, ADP |
 |
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-09-04 Classification: DNA BINDING PROTEIN/DNA Ligands: AGS, MG, ADP |
 |
Structure Of The Saccharomyces Cerevisiae Pcna Clamp Unloader Elg1-Rfc Complex
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2024-05-22 Classification: REPLICATION Ligands: AGS, MG, ADP |

