Search Count: 932
All
Selected
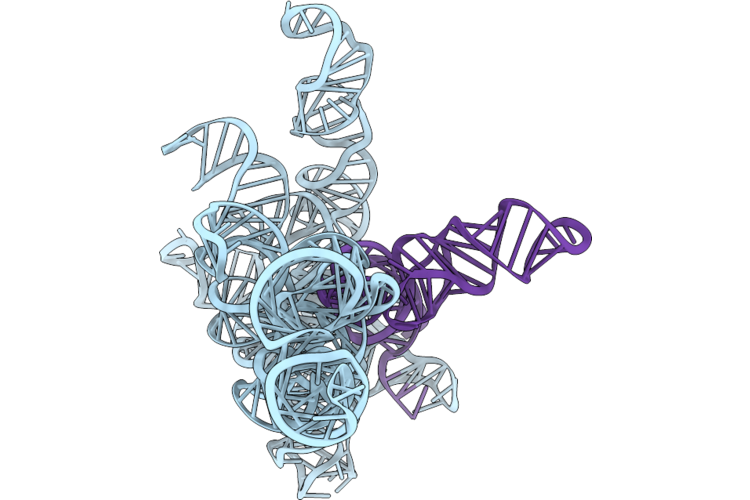 |
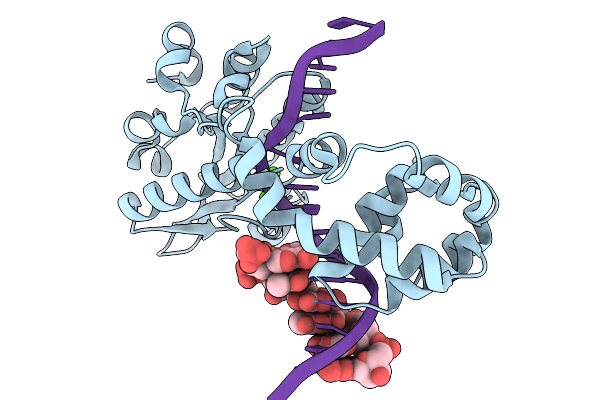
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme In Complex With Precursor Trna With Non-Complementary 5' Leader (Consensus)
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: RNA Ligands: CA |
 |
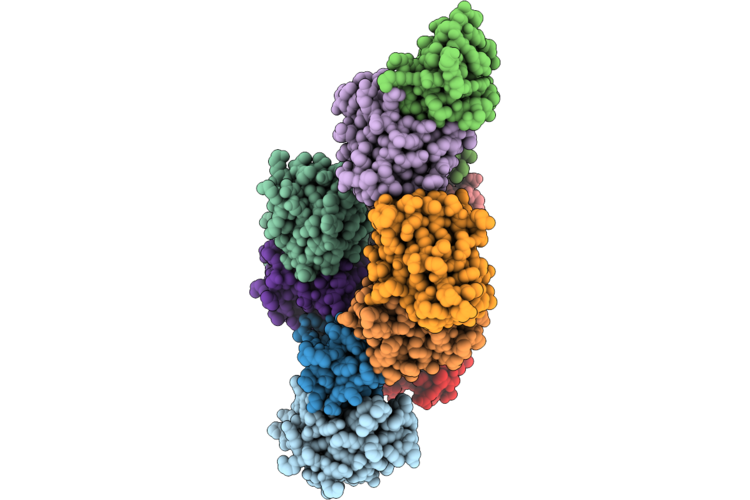
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme In Complex With Precursor Trna With Loop-Back 5' Leader
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: RNA Ligands: CA |
 |
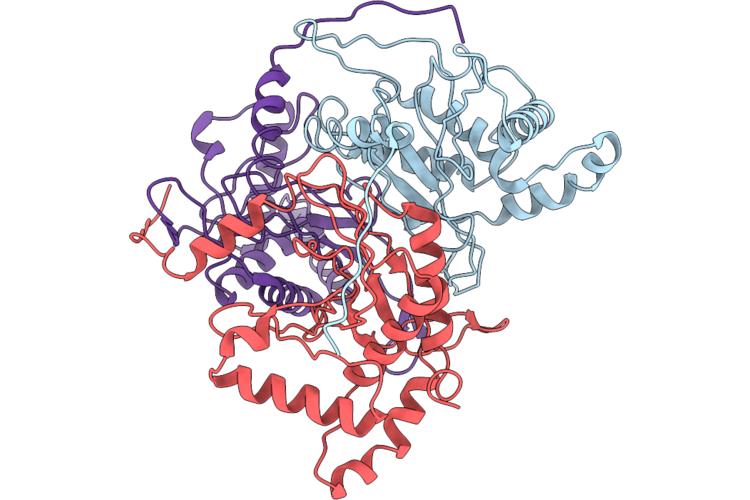
Crystal Structure Of Geocas9 Hnh Domain Bound To S78A Mutant Anti-Crispr Acriic1
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-29 Classification: DNA BINDING PROTEIN |
 |
Organism: Geobacillus stearothermophilus, Neisseria meningitidis
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-04-29 Classification: DNA BINDING PROTEIN |
 |
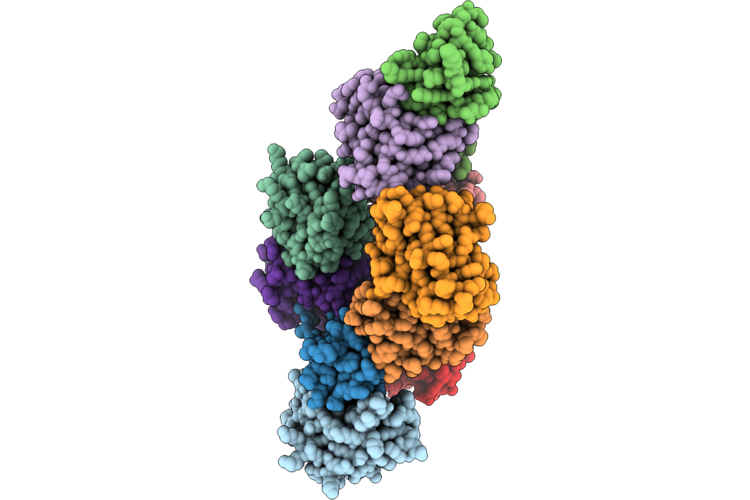
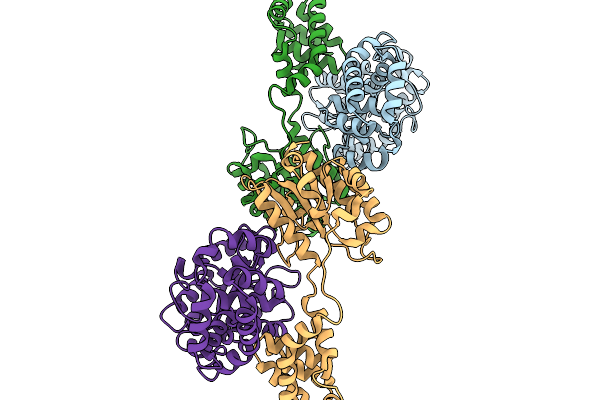
Dihydrolipoyl Acetyl Transferase (E2) Inner Core Of The Pyruvate Dehydrogenase Complex
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: STRUCTURAL PROTEIN |
 |
Crystal Structure Of The V165A/S219E/A225P Mutant Of Alanine Dehyrogenase From Geobacillus Stearothermophilus In Complex With Nicotinamide Cytosine Dinucleotide
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:2.56 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: A1EP0 |
 |
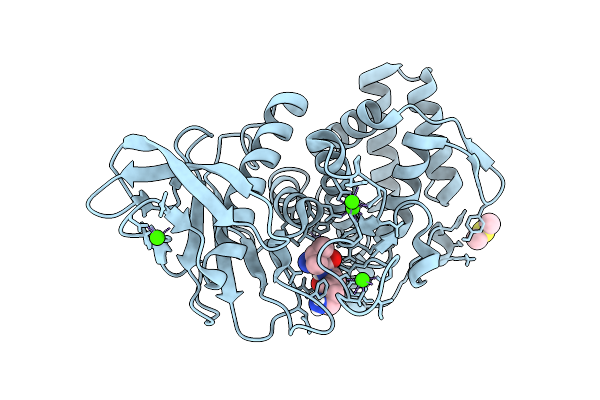
Organism: Geobacillus stearothermophilus, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2026-03-04 Classification: HYDROLASE Ligands: CA |
 |
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:2.72 Å Release Date: 2026-03-04 Classification: HYDROLASE |
 |
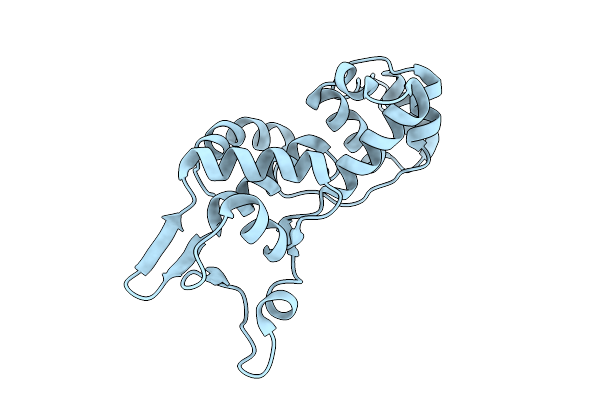
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-25 Classification: DNA BINDING PROTEIN Ligands: EDO, PGE, CL |
 |
Gradient Equilibration Of Hexagonal Thermolysin To Low Salt Over 15 Minutes
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-10-15 Classification: HYDROLASE Ligands: VAL, LYS, CA, ZN, DMS |
 |
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2025-08-27 Classification: RNA BINDING PROTEIN |
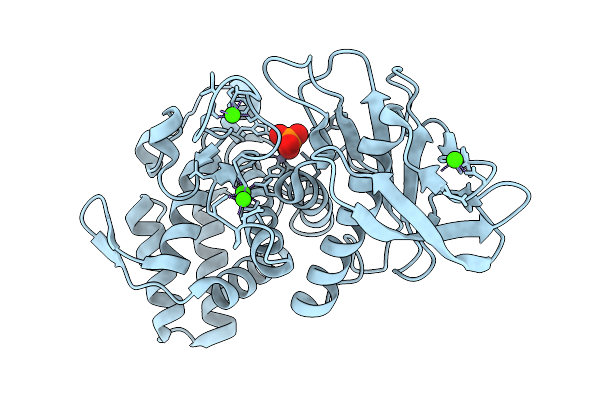 |
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-08-27 Classification: HYDROLASE Ligands: ZN, CA, PO4 |
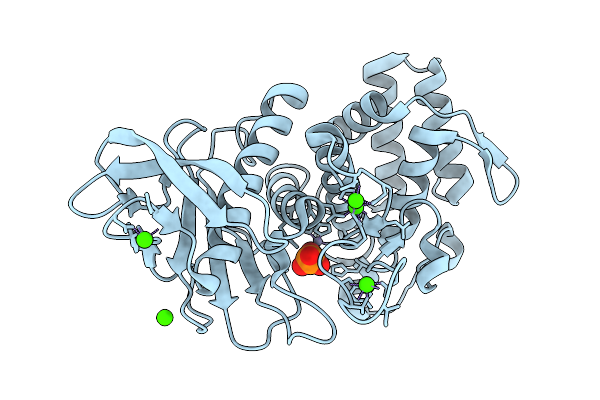 |
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2025-08-27 Classification: HYDROLASE Ligands: ZN, CA, PO4 |
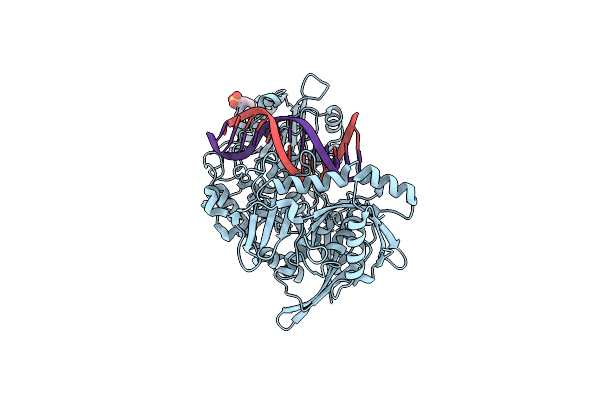 |
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-04-02 Classification: HYDROLASE Ligands: CL, MES |
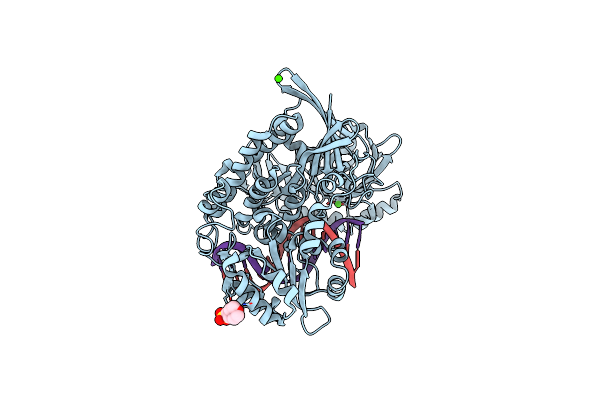 |
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2025-04-02 Classification: HYDROLASE Ligands: CL, MES, CA |
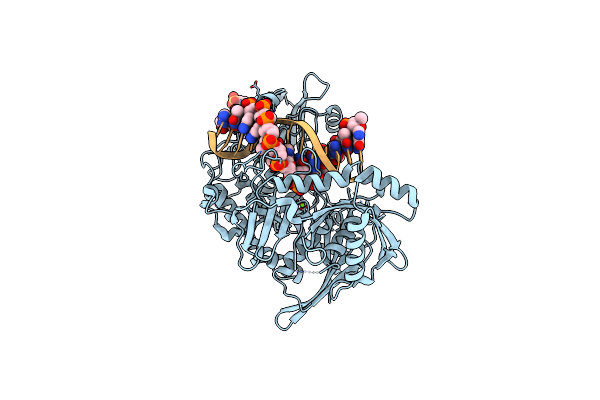 |
Bsmi (Bottom Nicking Mutant) Crystallized With Mg2+ And Cognate Dsdna (Post-Reactive Complex)
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:2.64 Å Release Date: 2025-04-02 Classification: HYDROLASE Ligands: MG, CL, MES |
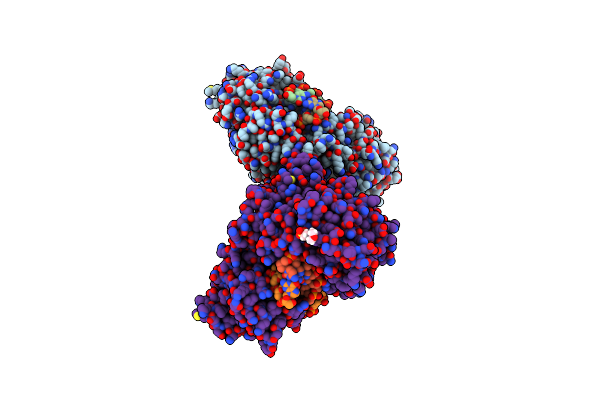 |
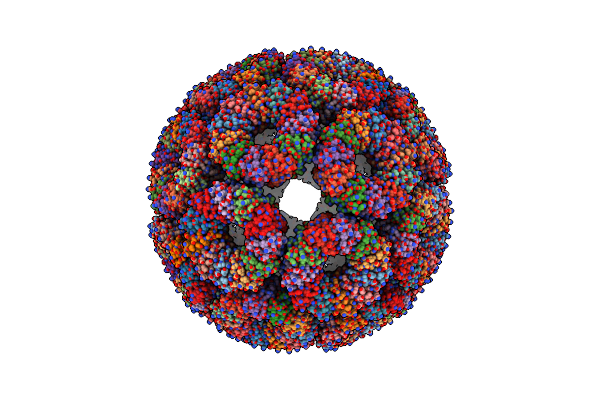
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2025-04-02 Classification: HYDROLASE Ligands: CA, MES, CL |
 |
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-25 Classification: VIRUS LIKE PARTICLE Ligands: CO |
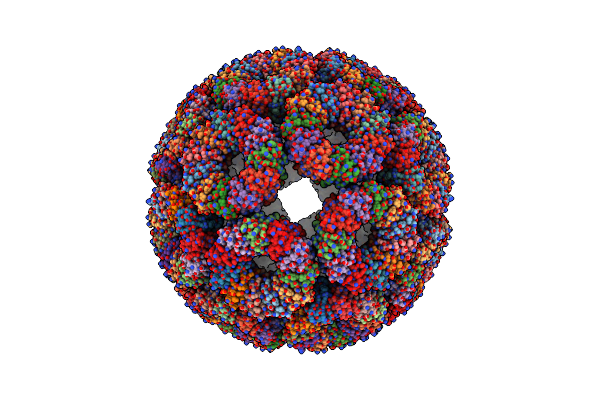 |
Structure Of The Co(Ii) Triggered Trap (S33Hk35H) Protein Cage (Dextro Form)
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-25 Classification: VIRUS LIKE PARTICLE Ligands: CO |
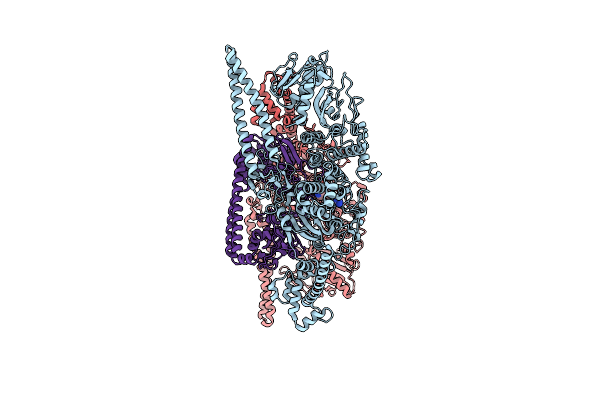 |
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2024-10-30 Classification: ANTIMICROBIAL PROTEIN Ligands: SAM |

