Search Count: 40
All
Selected
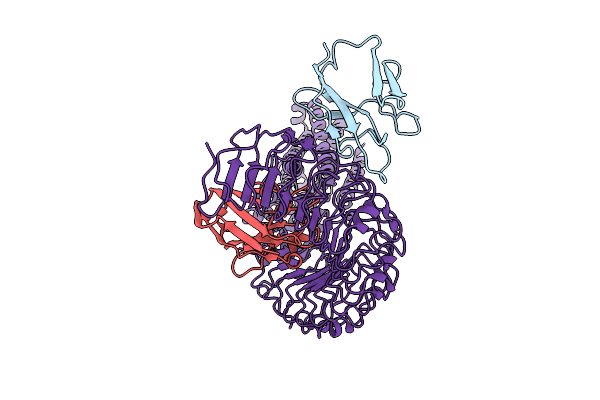 |
Organism: Camelus bactrianus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN |
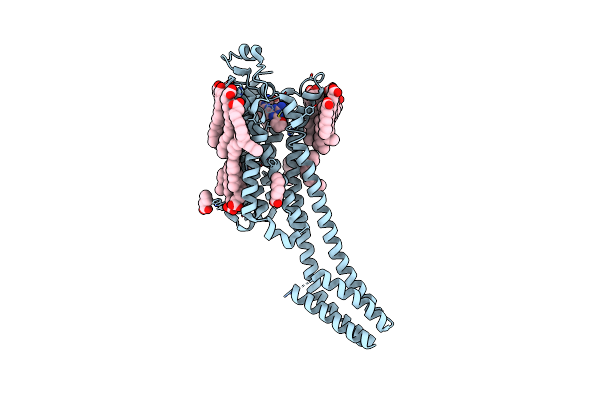 |
Crystal Structure Of Stabilized A2A Adenosine Receptor A2Ar-Star2-Bril In Complex With Compound 7, A Novel Nanomolar A2A Receptor Antagonist From Modern Hit-Finding With Structure-Guided De Novo Design
Organism: Homo sapiens, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-06-18 Classification: MEMBRANE PROTEIN |
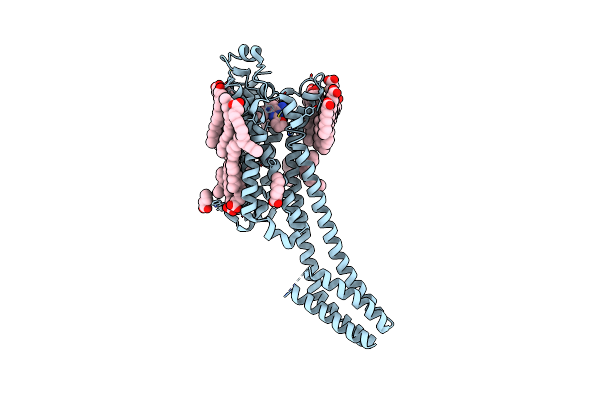 |
Crystal Structure Of Stabilized A2A Adenosine Receptor A2Ar-Star2-Bril In Complex With Compound 9, A Novel Nanomolar A2A Receptor Antagonist From Modern Hit-Finding With Structure-Guided De Novo Design
Organism: Homo sapiens, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2025-06-18 Classification: MEMBRANE PROTEIN |
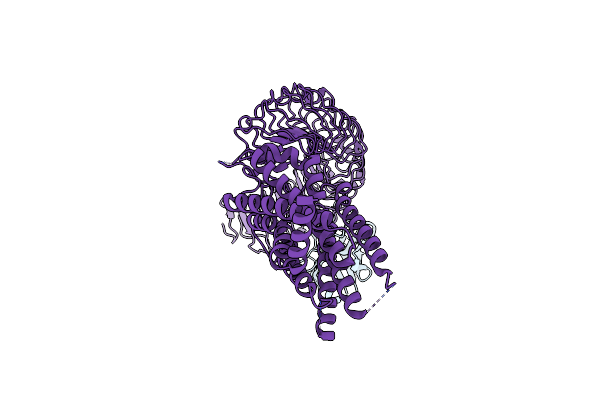 |
Organism: Homo sapiens, Camelus bactrianus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-15 Classification: MEMBRANE PROTEIN |
 |
Organism: Camelus bactrianus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-01-15 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens, Camelus bactrianus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens, Camelus bactrianus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens, Camelus bactrianus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens, Camelus bactrianus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: MEMBRANE PROTEIN Ligands: CLR |
 |
Crystal Structure Of Sars-Cov-2 Main Protease In Orthorhombic Space Group P212121
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2023-03-22 Classification: HYDROLASE Ligands: CL, DMS, MLI |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2023-01-25 Classification: VIRAL PROTEIN Ligands: DMS |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2023-01-18 Classification: VIRAL PROTEIN Ligands: CL |
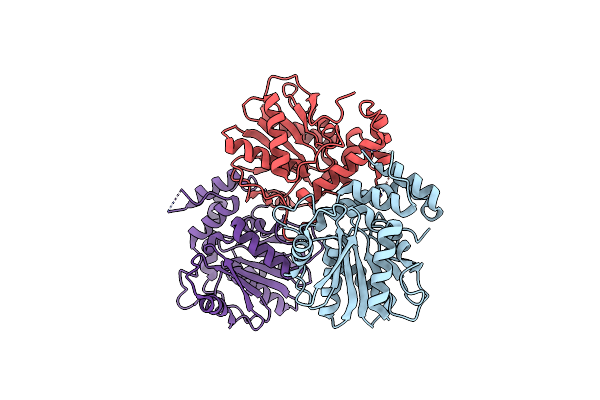 |
The Mutant Of Bifunctional Thioesterase Catalyzing Epimerization And Cyclization
Organism: Streptomyces abietis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2022-01-26 Classification: BIOSYNTHETIC PROTEIN |
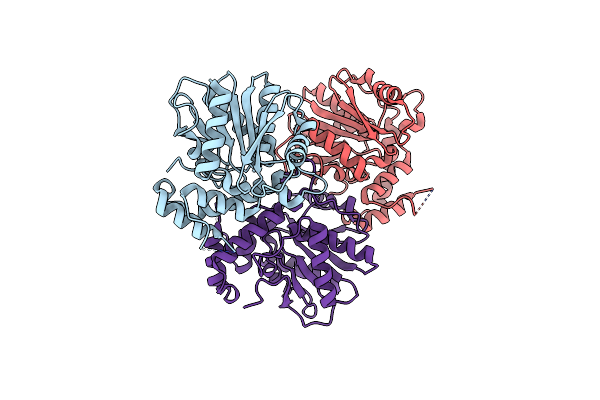 |
The Functional Characterization And Crystal Structure Of The Bifunctional Thioesterase Catalyzing Epimerization And Cyclization
Organism: Streptomyces abietis
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2021-08-18 Classification: BIOSYNTHETIC PROTEIN |
 |
Structure Of The A2A Adenosine Receptor Determined At Swissfel Using Native-Sad At 4.57 Kev From All Available Diffraction Patterns
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2020-07-15 Classification: SIGNALING PROTEIN Ligands: ZMA, OLA, OLB, OLC, CLR |
 |
Structure Of The A2A Adenosine Receptor Determined At Swissfel Using Native-Sad At 4.57 Kev From 50,000 Diffraction Patterns
Organism: Homo sapiens, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2020-07-15 Classification: MEMBRANE PROTEIN Ligands: ZMA, OLA, OLB, OLC, CLR |
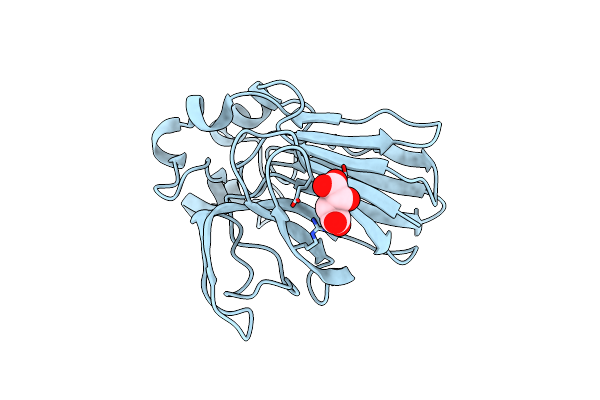 |
Structure Of Thaumatin Determined At Swissfel Using Native-Sad At 4.57 Kev From All Available Diffraction Patterns
Organism: Thaumatococcus daniellii
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2020-07-15 Classification: PLANT PROTEIN Ligands: TLA |
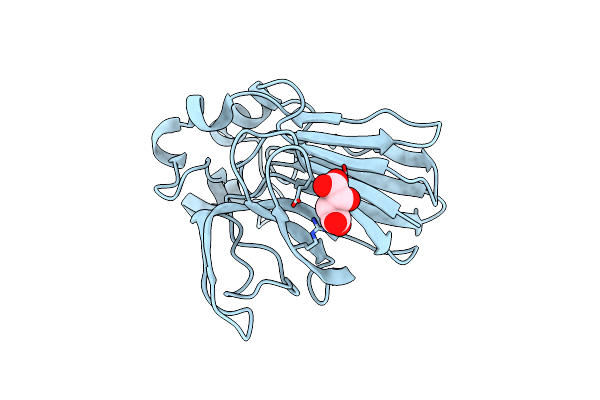 |
Structure Of Thaumatin Determined At Swissfel Using Native-Sad At 4.57 Kev From 20,000 Diffraction Patterns
Organism: Thaumatococcus daniellii
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2020-07-15 Classification: PLANT PROTEIN Ligands: TLA |
 |
Structure Of Thaumatin Determined At Swissfel Using Native-Sad At 6.06 Kev From All Available Diffraction Patterns
Organism: Thaumatococcus daniellii
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2020-07-15 Classification: PLANT PROTEIN Ligands: TLA |
 |
Structure Of Thaumatin Determined At Swissfel Using Native-Sad At 6.06 Kev From 50,000 Diffraction Patterns.
Organism: Thaumatococcus daniellii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2020-07-15 Classification: PLANT PROTEIN Ligands: TLA |

