Search Count: 198
All
Selected
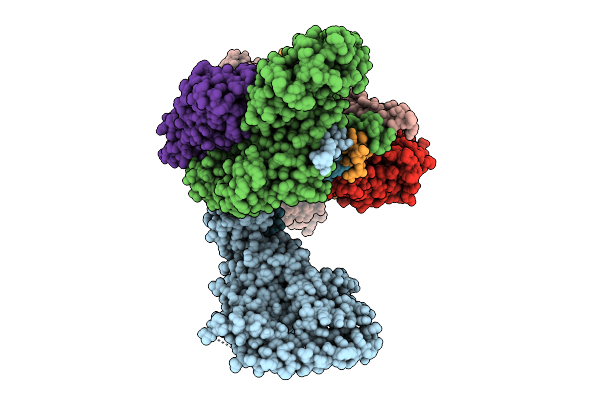 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: METAL BINDING PROTEIN Ligands: ZN |
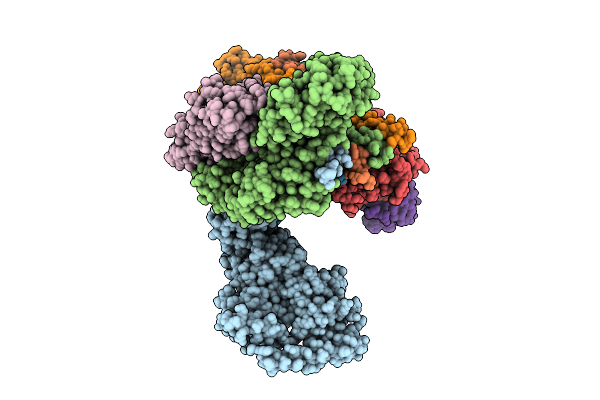 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: SIGNALING PROTEIN Ligands: 6LT, ZN |
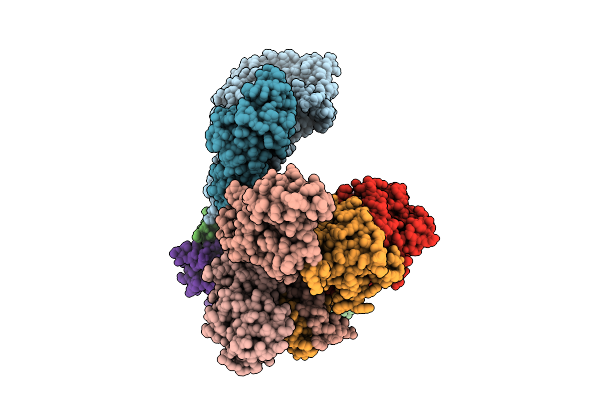 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: SIGNALING PROTEIN Ligands: 6LT, ZN |
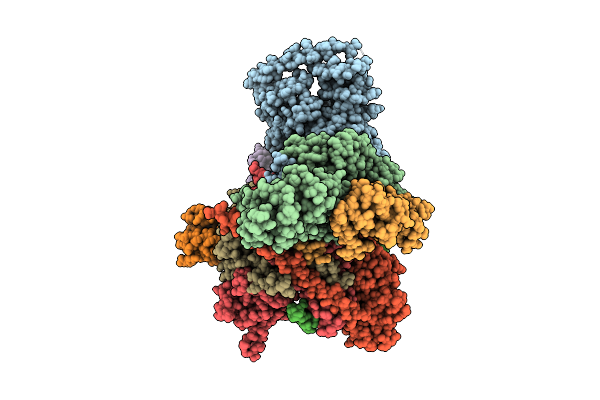 |
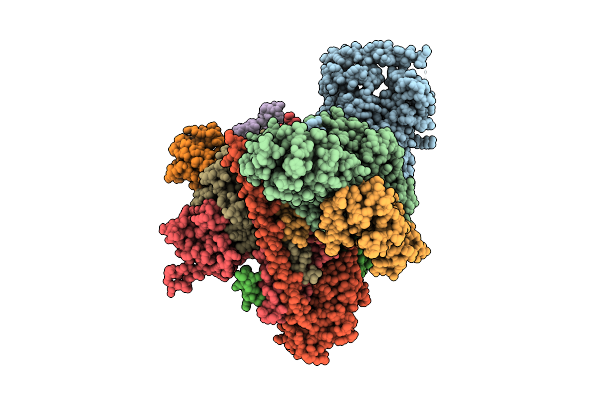
Cryo-Em Structure Of Cop9 Signalosome Precatalytic State With Neddylated Cullin-1
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: SIGNALING PROTEIN Ligands: ZN |
 |
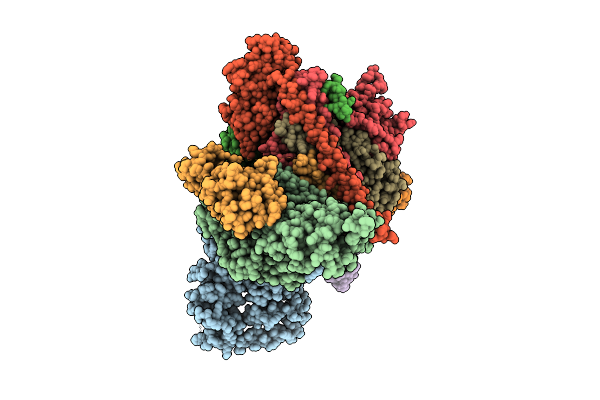
Cryo-Em Structure Of Cop9 Signalosome Precatalytic State With Neddylated Cullin-2
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2025-12-03 Classification: SIGNALING PROTEIN Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: SIGNALING PROTEIN Ligands: 6LT, ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.52 Å Release Date: 2025-12-03 Classification: SIGNALING PROTEIN Ligands: ZN |
 |
Cryo-Em Structure Of Cop9 Signalosome Precatalytic State With Neddylated Cullin-4A
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.39 Å Release Date: 2025-12-03 Classification: SIGNALING PROTEIN Ligands: ZN |
 |
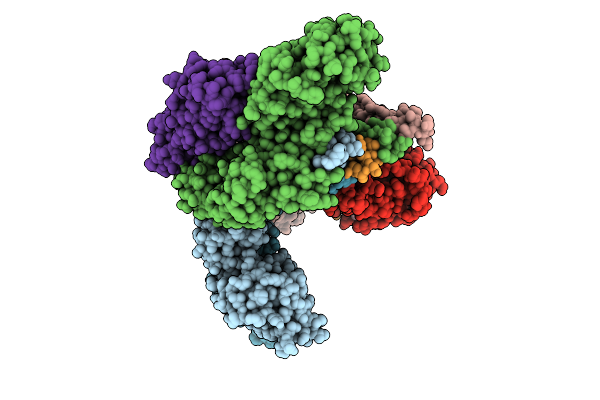
Cryo-Em Structure Of Cop9 Signalosome Precatalytic State With Neddylated Cullin-3
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.93 Å Release Date: 2025-12-03 Classification: SIGNALING PROTEIN Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2025-12-03 Classification: SIGNALING PROTEIN Ligands: ZN, A1CH3 |
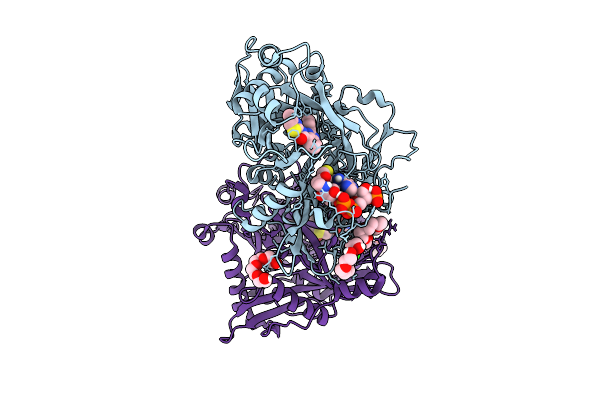 |
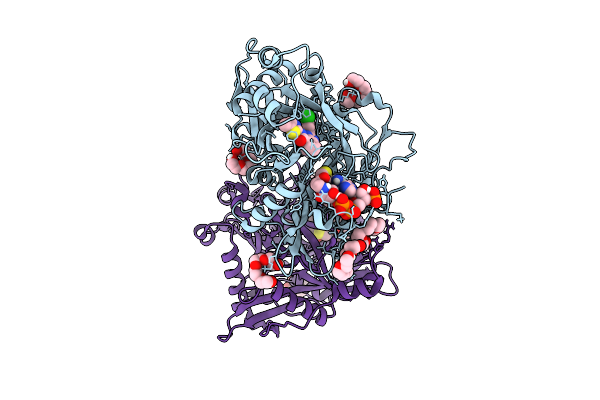
Crystal Structure Of Cryptosporidium Parvum N-Myristoyltransferase With Bound Myristoyl-Coa And Inhibitor 20045
Organism: Cryptosporidium parvum iowa ii
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-11-26 Classification: TRANSFERASE/INHIBITOR Ligands: MYA, A1BHP, CL, PG4, 1PE, PGE |
 |
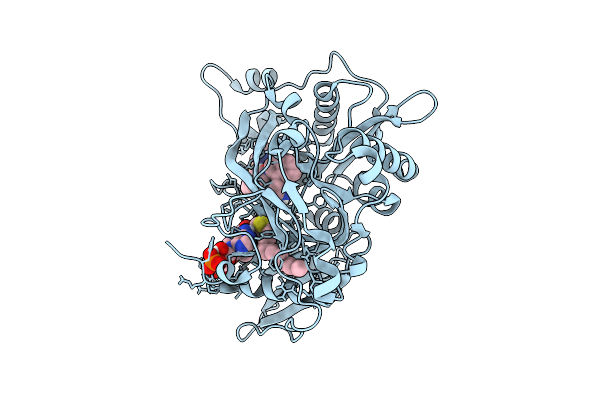
Crystal Structure Of Cryptosporidium Parvum N-Myristoyltransferase With Bound Myristoyl-Coa And Inhibitor 20057
Organism: Cryptosporidium parvum iowa ii
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-11-26 Classification: TRANSFERASE/INHIBITOR Ligands: MYA, A1BHQ, CL, 1PE, PG4 |
 |
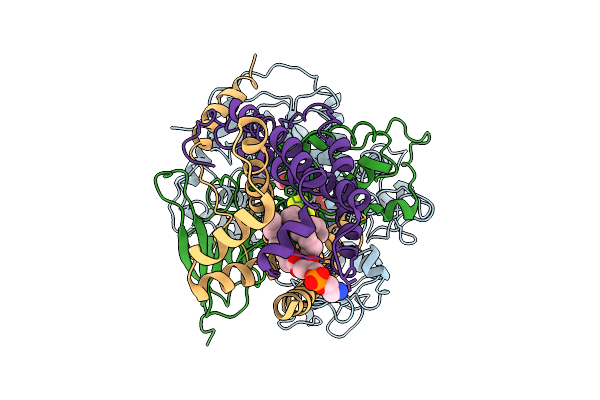
Crystal Structure Of Cryptosporidium Parvum N-Myristoyltransferase With Bound Myristoyl-Coa And Inhibitor 20084
Organism: Cryptosporidium parvum iowa ii
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2025-11-26 Classification: TRANSFERASE/INHIBITOR Ligands: MYA, A1BHR, CL |
 |
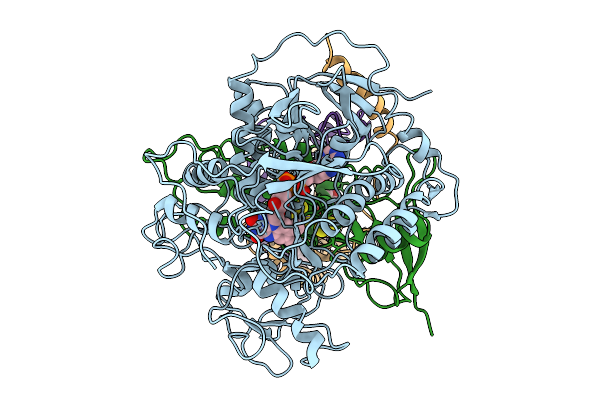
Cryo-Em Structure Of Saccharomyces Cerevisiae Mitochondrial Respiratory Complex Ii
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:3.36 Å Release Date: 2025-09-03 Classification: MEMBRANE PROTEIN Ligands: FAD, FES, SF4, F3S, PEE |
 |
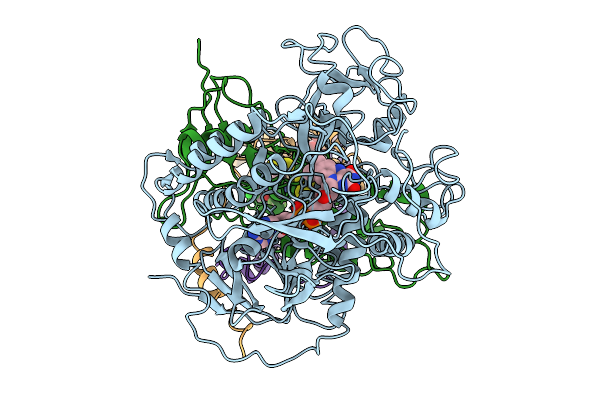
Cryo-Em Structure Of Saccharomyces Cerevisiae Mitochondrial Respiratory Complex Ii In Uq1-Bound State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: MEMBRANE PROTEIN Ligands: FAD, FES, SF4, F3S, UQ1, PEE |
 |
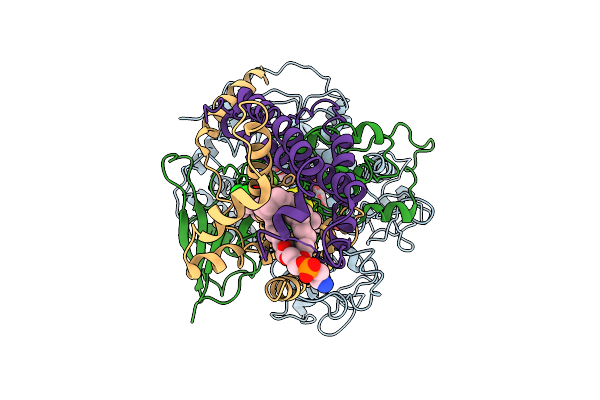
Cryo-Em Structure Of Saccharomyces Cerevisiae Mitochondrial Respiratory Complex Ii In Pydiflumetofen-Bound State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: MEMBRANE PROTEIN Ligands: FAD, FES, SF4, F3S, A1EE4, PEE |
 |
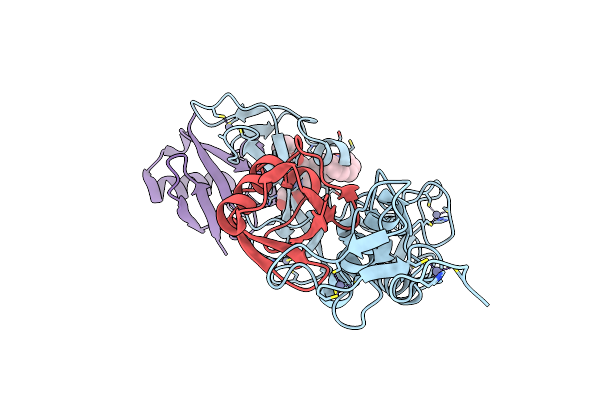
Cryo-Em Structure Of Saccharomyces Cerevisiae Mitochondrial Respiratory Complex Ii In Y19315-Bound State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: MEMBRANE PROTEIN Ligands: FAD, FES, SF4, F3S, A1EKV, PEE |
 |
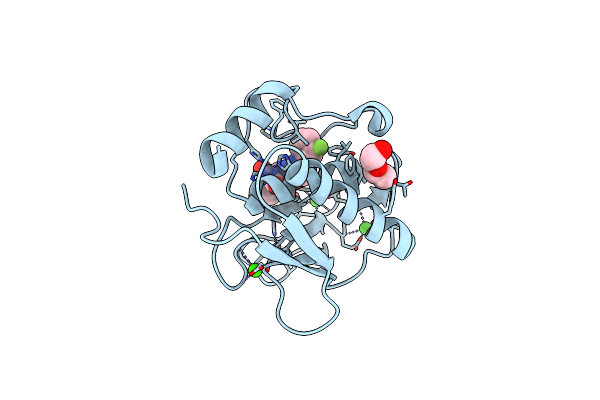
Organism: Rattus norvegicus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2024-08-21 Classification: LIGASE Ligands: ZN, A1AE9, GOL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2024-02-21 Classification: HYDROLASE Ligands: ZN, CA, A1H07, PEG, CL |
 |
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2024-02-21 Classification: MEMBRANE PROTEIN Ligands: CLR |

