Search Count: 274
All
Selected
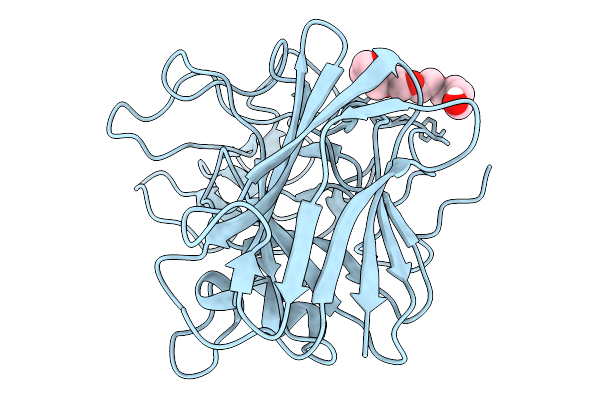 |
Organism: Saccharomyces cerevisiae s288c
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2026-02-18 Classification: STRUCTURAL PROTEIN Ligands: P6G |
 |
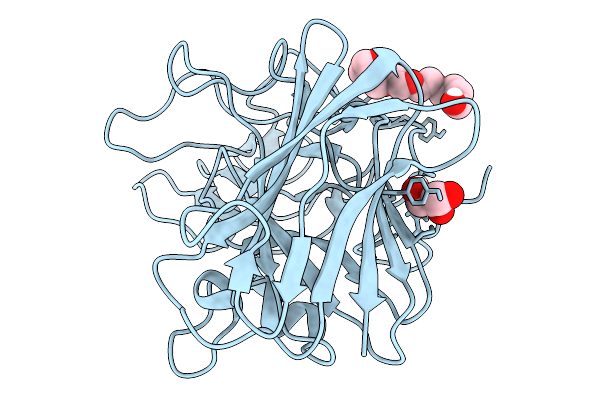
Structure Of The Q318A Mutant Of The Sun4 Domain From Saccharomyces Cerevisiae
Organism: Saccharomyces cerevisiae s288c
Method: X-RAY DIFFRACTION Resolution:1.28 Å Release Date: 2026-02-18 Classification: STRUCTURAL PROTEIN Ligands: P6G, CL |
 |
Structure Of The E283A Mutant Of The Sun4 Domain From Saccharomyces Cerevisiae
Organism: Saccharomyces cerevisiae s288c
Method: X-RAY DIFFRACTION Resolution:1.24 Å Release Date: 2026-02-18 Classification: CARBOHYDRATE Ligands: P6G, GOL, CL |
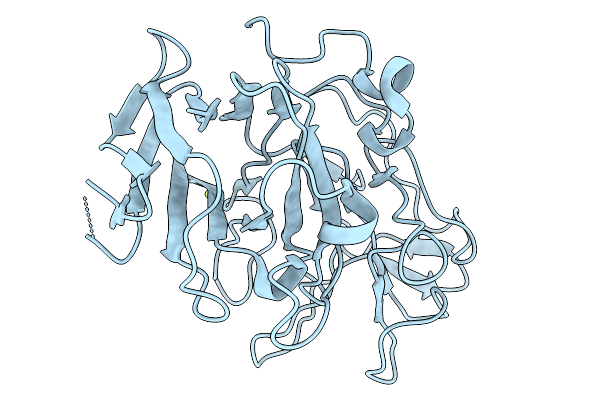 |
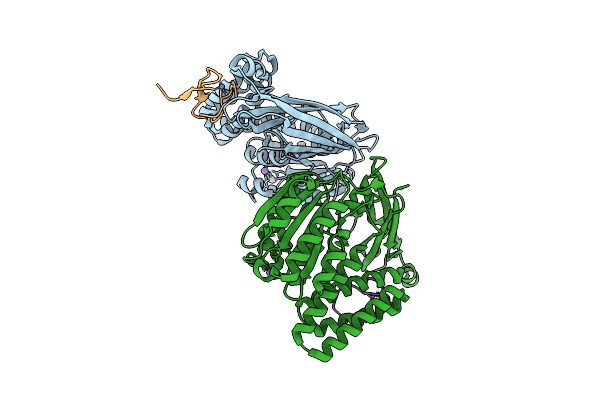
Organism: Saccharomyces cerevisiae s288c
Method: X-RAY DIFFRACTION Resolution:1.23 Å Release Date: 2026-02-18 Classification: CARBOHYDRATE Ligands: MG |
 |
Organism: Sulfolobus acidocaldarius dsm 639
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2025-10-29 Classification: PROTEIN BINDING |
 |
Organism: Thermochaetoides thermophila dsm 1495
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: GOL, FMT, NA |
 |
Crystal Structure Of Ctgh76 From Chaetomium Thermophilum In Complex With Mannose
Organism: Thermochaetoides thermophila dsm 1495
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: GOL, FMT, MAN |
 |
Crystal Structure Of Ctgh76 From Chaetomium Thermophilum In Complex With Mannose Products Form
Organism: Thermochaetoides thermophila dsm 1495
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: MAN, GOL, FMT |
 |
Organism: Thermochaetoides thermophila dsm 1495
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: MAN, GOL, FMT |
 |
Crystal Structure Of Inactive (D163N) Variant Ctgh76 From Chaetomium Thermophilum In Complex With Alpha-1,6-Mannobiose
Organism: Thermochaetoides thermophila dsm 1495
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: MAN, GOL |
 |
Crystal Structure Of Inactive (D164N) Variant Ctgh76 From Chaetomium Thermophilum In Complex With Alpha-1,6-Mannobiose
Organism: Thermochaetoides thermophila dsm 1495
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: MAN |
 |
Crystal Structure Of Ctgh76 From Chaetomium Thermophilum In Complex With Alpha-1,3-1,6-Mannotriose
Organism: Thermochaetoides thermophila dsm 1495
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: GOL, FMT |
 |
Crystal Structure Of Ctgh76 From Chaetomium Thermophilum In Complex With Glucosamine
Organism: Thermochaetoides thermophila dsm 1495
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: PA1, GOL, FMT |
 |
Crystal Structure Of Ctgh76 From Chaetomium Thermophilum In Complex With Laminaribiose
Organism: Thermochaetoides thermophila dsm 1495
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: BGC, GOL, FMT |
 |
Crystal Structure Of Ctgh76 From Chaetomium Thermophilum In Complex With Glucose
Organism: Thermochaetoides thermophila dsm 1495
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: GOL, BGC |
 |
Crystal Structure Of Ctgh76 From Chaetomium Thermophilum In Complex With Isomaltose
Organism: Thermochaetoides thermophila dsm 1495
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2025-07-16 Classification: HYDROLASE Ligands: BGC, GOL, FMT |
 |
Organism: Chlamydomonas reinhardtii cc3269
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2025-05-14 Classification: FLAVOPROTEIN Ligands: FAD, CL |
 |
Sfx Structure Of Cracry 10 Ns After Photoexcitation Of The Oxidized Protein
Organism: Chlamydomonas reinhardtii
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-05-14 Classification: FLAVOPROTEIN Ligands: FAD, CL |
 |
Sfx Structure Of Cracry 30 Ns After Photoexcitation Of The Oxidized Protein
Organism: Chlamydomonas reinhardtii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-05-14 Classification: FLAVOPROTEIN Ligands: FAD, CL |
 |
Sfx Structure Of Cracry 100 Ns After Photoexcitation Of The Oxidized Protein
Organism: Chlamydomonas reinhardtii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-05-14 Classification: FLAVOPROTEIN Ligands: FAD, CL |

