Search Count: 237
All
Selected
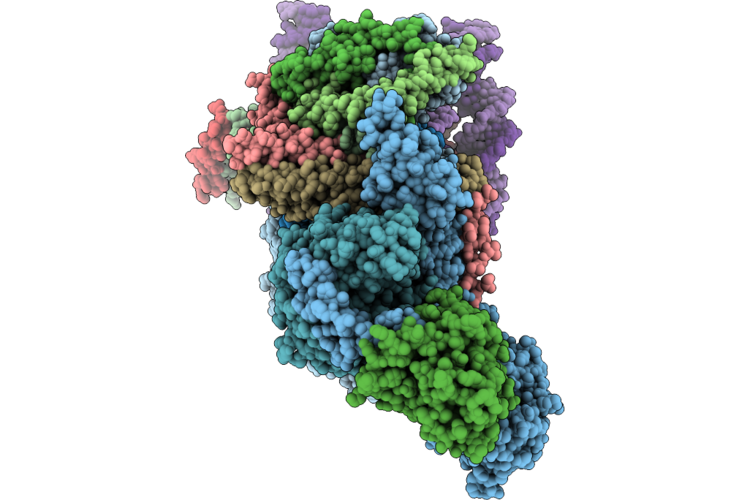 |
Cryo-Em Structure Of The Histone Deacetylase Complex Rpd3L In Complex With Mono-Nucleosome
Organism: Saccharomyces cerevisiae s288c, Xenopus laevis, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: GENE REGULATION/DNA Ligands: ZN |
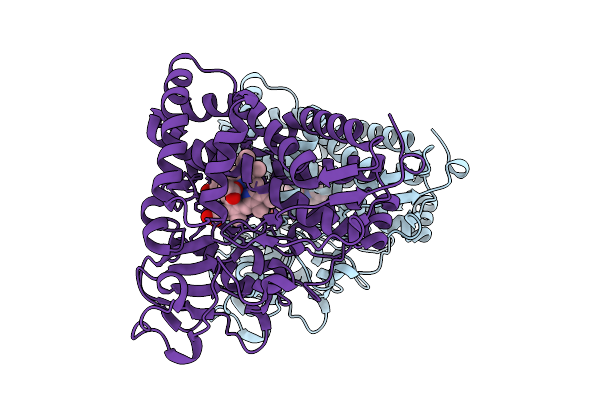 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: HEM, ACT |
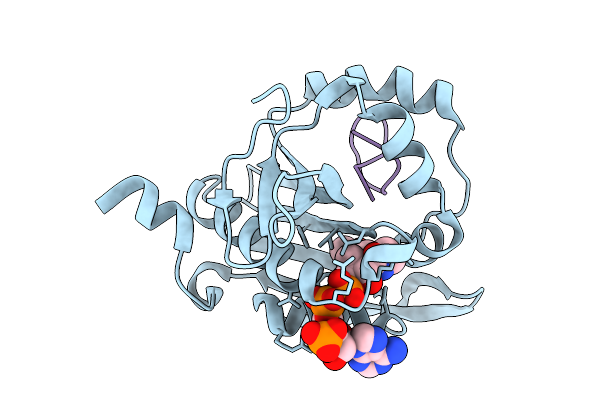 |
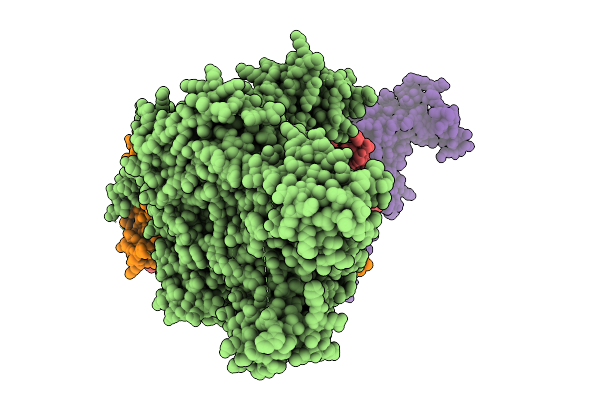
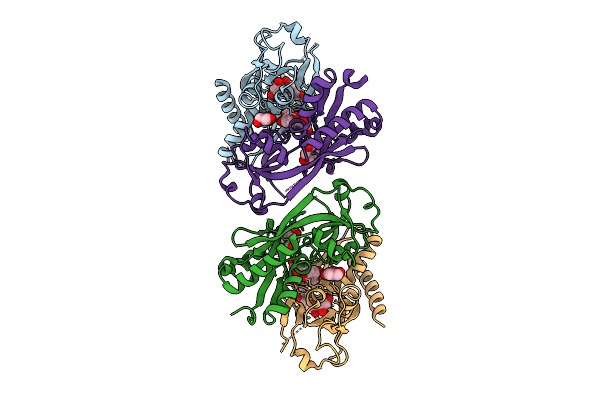
Sub-Particle Structure Of The Iterative Acetyltransferase From Actinomycetes In Complex With Accoa And Monoacetylated Lasso Peptides
Organism: Actinosynnema mirum dsm 43827
Method: ELECTRON MICROSCOPY Resolution:3.43 Å Release Date: 2026-02-25 Classification: TRANSFERASE Ligands: ACO |
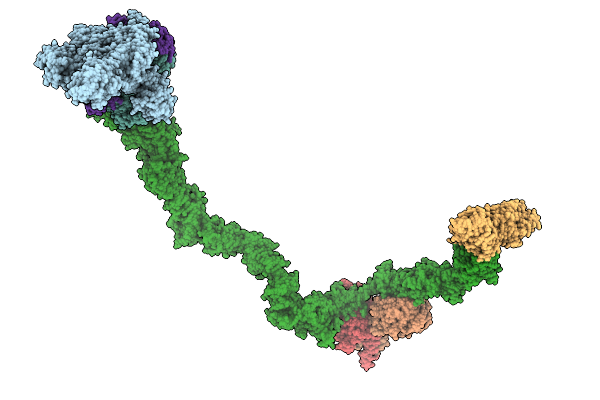 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: CELL CYCLE |
 |
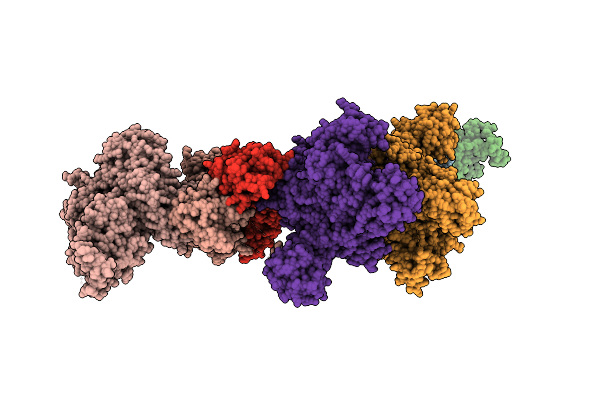
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.34 Å Release Date: 2026-01-21 Classification: CELL CYCLE |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:5.00 Å Release Date: 2026-01-21 Classification: CELL CYCLE |
 |
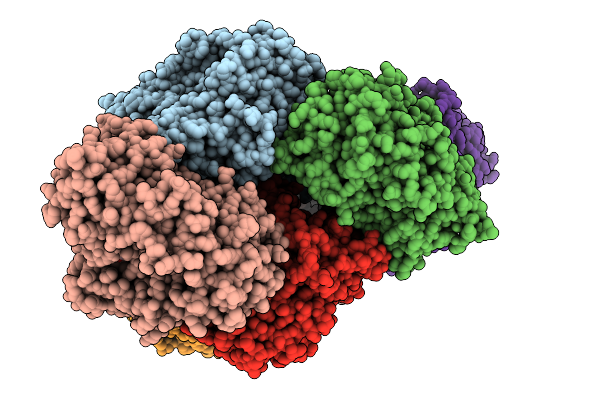
Organism: Priestia megaterium
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: HEM, A1EI5 |
 |
Organism: Priestia megaterium
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: HEM, EDO |
 |
Organism: Priestia megaterium
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: A1EI5, HEM |
 |
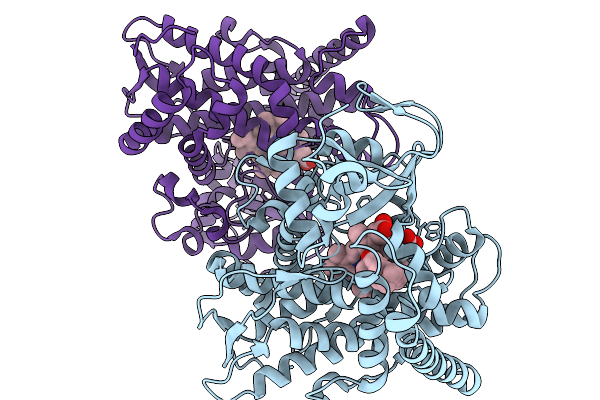
Crystal Structure Of P450 Bm3 F87A/V78S/L75N Mutant Complex With (-)-Ambroxide
Organism: Priestia megaterium
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: HEM, A1EI5 |
 |
Organism: Priestia megaterium
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: HEM, A1EI5 |
 |
Crystal Structure Of Nodd-Ebd (Effector Binding Domain) In Complex With Hesperetin From Rhizobium Leguminosarum Bv. Vicae 3841
Organism: Rhizobium leguminosarum
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2025-12-24 Classification: TRANSCRIPTION Ligands: GOL, 6JP |
 |
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-12-24 Classification: PLANT PROTEIN |
 |
Crystal Structure Of Nodd-Ebd (Effector Binding Domain) From Rhizobium Leguminosarum Bv. Vicae 3841
Organism: Rhizobium leguminosarum
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2025-12-24 Classification: TRANSCRIPTION |
 |
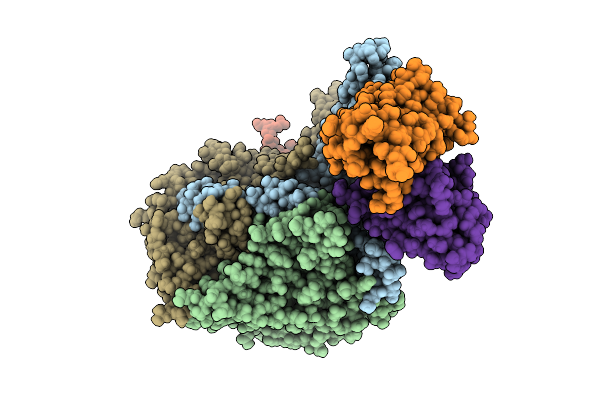
Organism: Poliovirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.65 Å Release Date: 2025-12-17 Classification: VIRUS Ligands: PLM |
 |
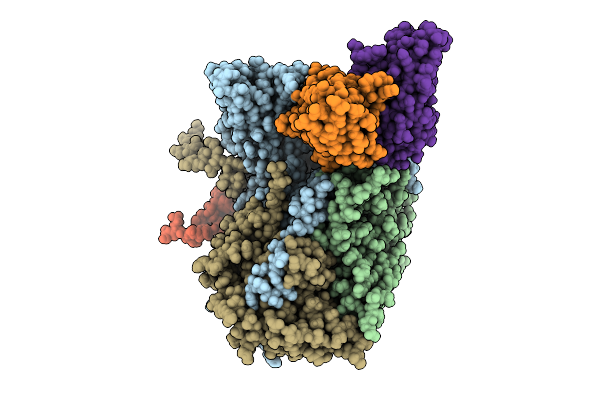
Organism: Poliovirus 3, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.95 Å Release Date: 2025-12-17 Classification: VIRUS Ligands: PLM |
 |
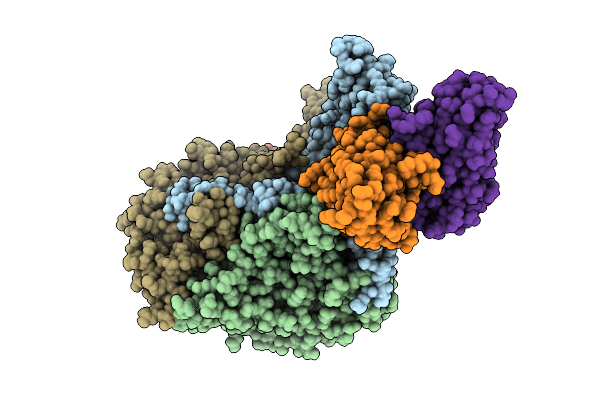
Organism: Poliovirus 3, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2025-12-17 Classification: VIRUS Ligands: PLM |
 |
Organism: Human poliovirus 1 strain sabin, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2025-12-17 Classification: VIRUS Ligands: PLM |
 |
Atomic Structure Of Vibrio Effector Fragment Vopv Bound To Beta-Cytoplasmic/Gamma1-Cytoplasmic F-Actin
Organism: Homo sapiens, Vibrio parahaemolyticus
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: STRUCTURAL PROTEIN Ligands: MG, ANP |
 |
Cryo-Em Structure Of Vibrio Effector Vopv Fragment Bound To Skeletal Alpha F-Actin
Organism: Oryctolagus cuniculus, Vibrio cholerae
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: STRUCTURAL PROTEIN Ligands: MG, ANP |

