Search Count: 418
All
Selected
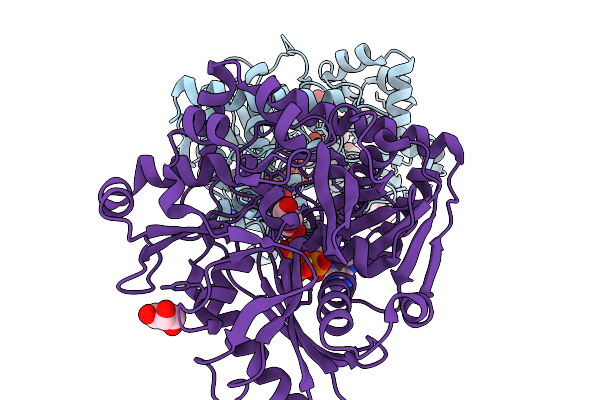 |
Crystal Structure Of Glycerol Kinase From Entamoeba Histolytica Complexed With Amp-Pnp And Glycerol.
Organism: Entamoeba histolytica
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-03-04 Classification: TRANSFERASE Ligands: ANP, GOL, CAC, MG, P6G, PG4 |
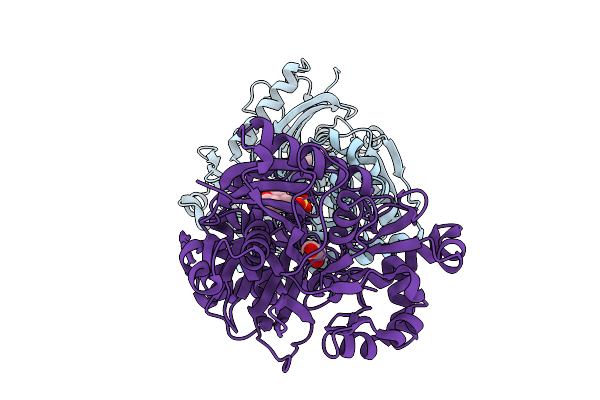 |
Crystal Structure Of Glycerol Kinase From Entamoeba Histolytica (Ligand-Free Form)
Organism: Entamoeba histolytica
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-25 Classification: TRANSFERASE Ligands: MES, EDO |
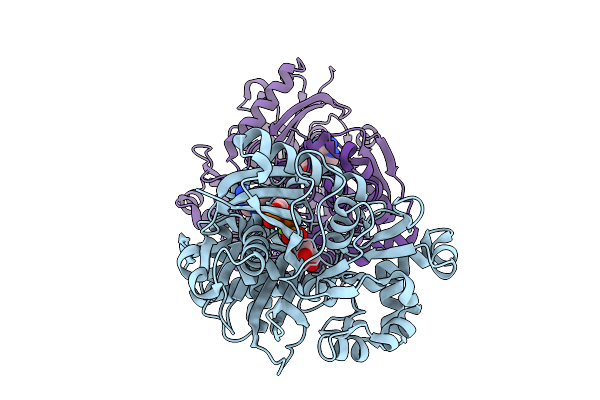 |
Crystal Structure Of Glycerol Kinase From Entamoeba Histolytica Complexed With Adp And G3P.
Organism: Entamoeba histolytica
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-02-25 Classification: TRANSFERASE Ligands: ADP, G3P, MG, PG4 |
 |
Crystal Structure Of Glycerol Kinase From Entamoeba Histolytica Complexed With Daphnetin.
Organism: Entamoeba histolytica
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-02-25 Classification: TRANSFERASE Ligands: GOL, PO4, A1L8C |
 |
Crystal Structure Of Glycerol Kinase From Entamoeba Histolytica Complexed With Gk-Butyl.
Organism: Entamoeba histolytica
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2026-02-25 Classification: TRANSFERASE Ligands: GOL, PO4, 6Y0 |
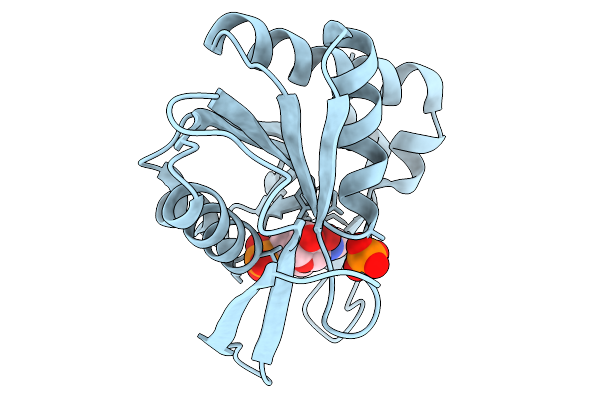 |
Structural Studies On The Conformation Changes Induced By Ligand Binding In An Adenine Phosphoribosyltransferase (Fnaprt) From Fusobacterium Nucleatum
Organism: Fusobacterium nucleatum
Method: X-RAY DIFFRACTION Resolution:1.61 Å Release Date: 2026-02-11 Classification: TRANSFERASE Ligands: PO4, AMP |
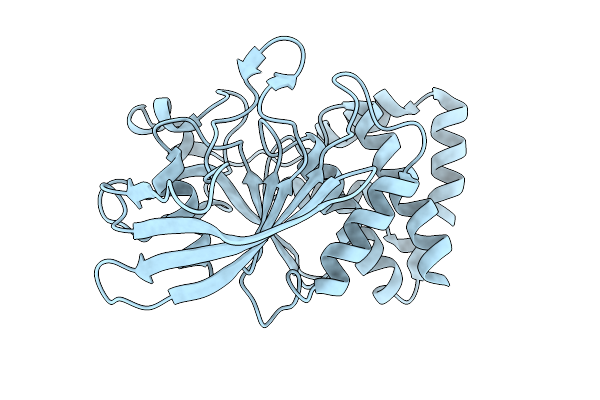 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-01-28 Classification: HYDROLASE |
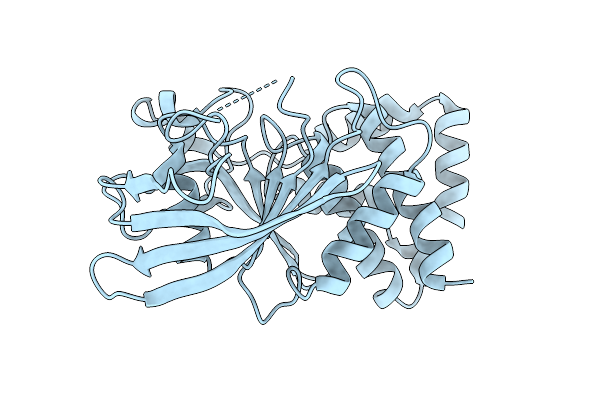 |
Crystal Structure Of Oxidized Form (C92-C121) Of Protein Tyrosine Phosphatase 1B (Ptp1B)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-01-28 Classification: HYDROLASE |
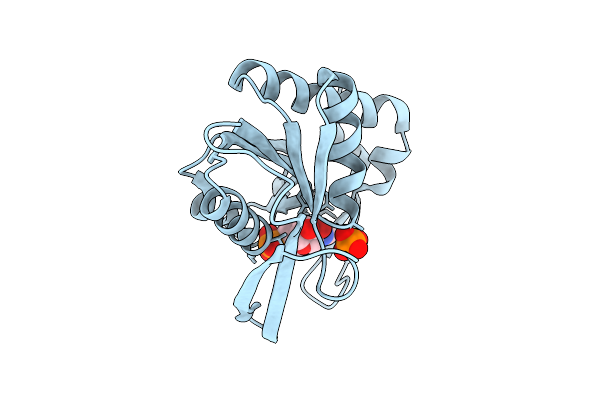 |
Structural Studies On The Conformation Changes Induced By Ligand Binding In An Adenine Phosphoribosyltransferase (Fnaprt) From Fusobacterium Nucleatum
Organism: Fusobacterium nucleatum
Method: X-RAY DIFFRACTION Resolution:1.44 Å Release Date: 2025-12-24 Classification: TRANSFERASE Ligands: PO4, AMP |
 |
Crystal Structure Of Ice Binding Protein From Candidatus Cryosericum Odellii Smc5
Organism: Candidatus cryosericum odellii
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2025-09-03 Classification: RECOMBINATION |
 |
Crystal Structure Of Ice Binding Protein From Candidatus Cryosericum Odellii Smc5
Organism: Candidatus cryosericum odellii
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-09-03 Classification: RECOMBINATION |
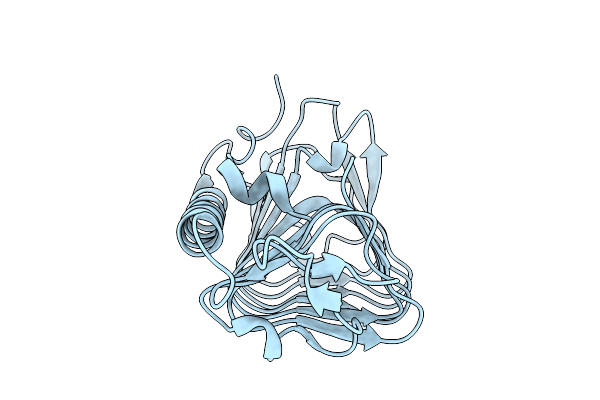 |
Crystal Structure Of An Ice-Binding Protein From Candidatus Cryosericum Odellii
Organism: Candidatus cryosericum odellii
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2025-09-03 Classification: PROTEIN BINDING |
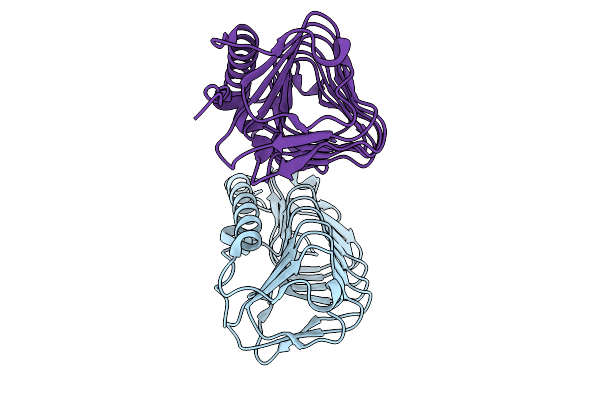 |
Crystal Structure Of Engineered Ice-Binding Protein (M74I And A97V) From Candidatus Cryosericum Odellii Strain Smc5_169
Organism: Candidatus cryosericum odellii
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2025-09-03 Classification: PROTEIN BINDING |
 |
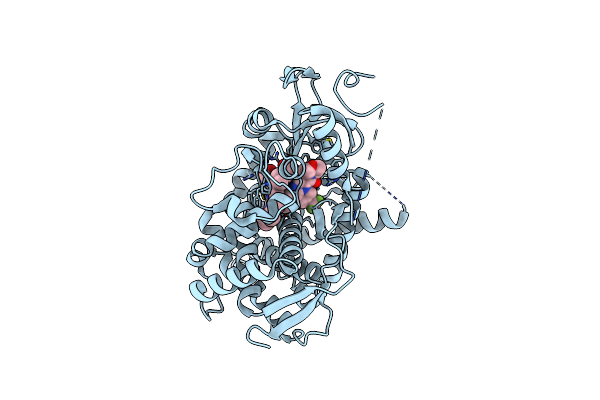
Crystal Structure Of Cyp46A1 With [(1R,5S)-3-Oxa-8-Azabicyclo[3.2.1]Octan-8-Yl][(4R,8M)-8-(1,3-Oxazol-5-Yl)-6-(Trifluoromethyl)Imidazo[1,2-A]Pyridin-3-Yl]Methanone (Compound 3K)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2025-07-02 Classification: HYDROLASE/INHIBITOR Ligands: HEM, A1BZE |
 |
Crystal Structure Of Cyp46A1 With (Morpholin-4-Yl)[(4R,8M)-8-(1,3-Oxazol-5-Yl)-6-(Trifluoromethyl)Imidazo[1,2-A]Pyridin-3-Yl]Methanone (Compound 2H)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-07-02 Classification: HYDROLASE/INHIBITOR Ligands: HEM, A1BZD |
 |
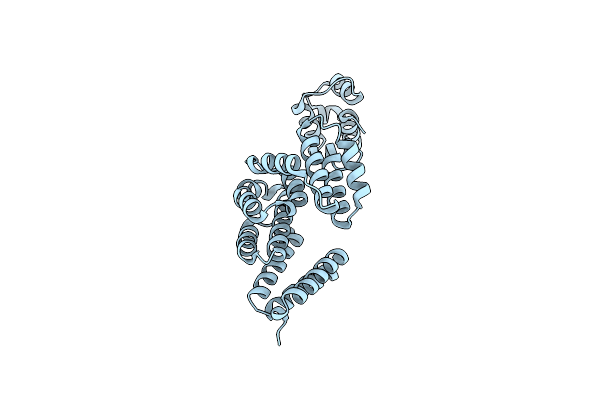
Crystal Structure Of Cyp46A1 With Cyclopropyl[(4M)-4-(1,3-Oxazol-5-Yl)-6-(Trifluoromethyl)-1H-Indol-1-Yl]Methanone (Compound 2B)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-07-02 Classification: HYDROLASE/INHIBITOR Ligands: HEM, A1BZC, EDO |
 |
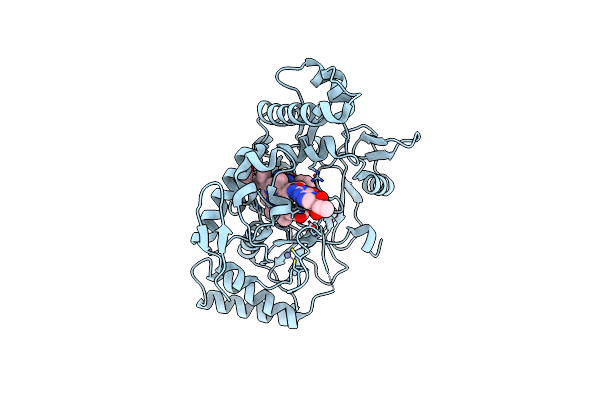
Crystal Structure Of Transcriptional Regulator (Nrpr) From Streptococcus Salivarius K12
Organism: Streptococcus salivarius k12
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2025-05-21 Classification: TRANSCRIPTION |
 |
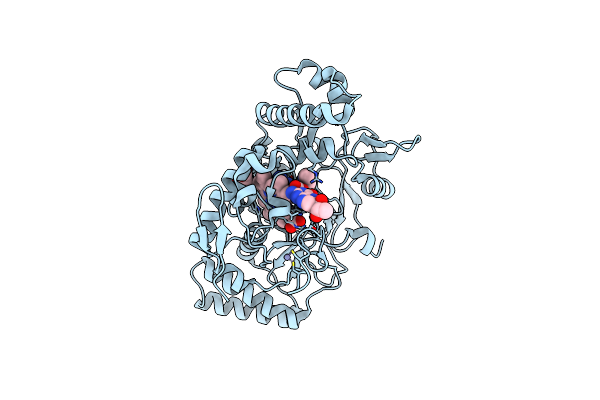
Structure Of Rat Neuronal Nitric Oxide Synthase R349A Mutant Heme Domain Bound With 4-Methyl-6-(3-((Methylamino)Methyl)Phenyl)Pyridin-2-Amine Dihydrochloride
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2025-04-30 Classification: OXIDOREDUCTASE Ligands: HEM, H4B, KMI, ACT, ZN |
 |
Structure Of Rat Nnos R349A Mutant Heme Domain Bound With 4-Methyl-6-(3-((Methylamino)Methyl)-5-(Trifluoromethyl)Phenyl)Pyridin-2-Amine Dihydrochloride
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2025-04-30 Classification: OXIDOREDUCTASE Ligands: HEM, H4B, ACT, ZN, A1A0F |
 |
Structure Of Rat Neuronal Nitric Oxide Synthase R349A Mutant Heme Domain Bound With 6-(3-Chloro-5-((Methylamino)Methyl)Phenyl)-4-Methylpyridin-2-Amine Dihydrochloride
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-04-30 Classification: OXIDOREDUCTASE Ligands: HEM, H4B, A1A0G, ACT, ZN |

