Search Count: 151
All
Selected
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:4.30 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: NAG |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.37 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: NAG, GLY, GLU |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.37 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: NAG, GLY, GLU, JC9 |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:6.37 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: NAG, JC9 |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: NAG, GLY, GLU, JC9 |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.55 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: NAG, GLY, GLU |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.27 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: NAG, GLY, GLU, JC9 |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: NAG, GLY, GLU |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.98 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: NAG, GLY, GLU, JC9 |
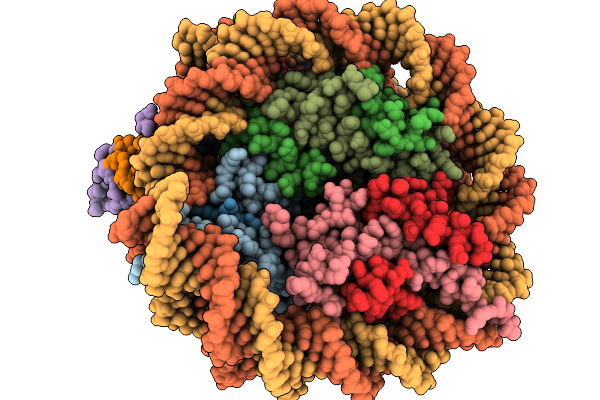 |
The 1:1 Cryo-Em Structure Of Bap1/Asxl1-K351Ub In Complex With H2Ak119Ub Nucleosome
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: NUCLEAR PROTEIN/DNA |
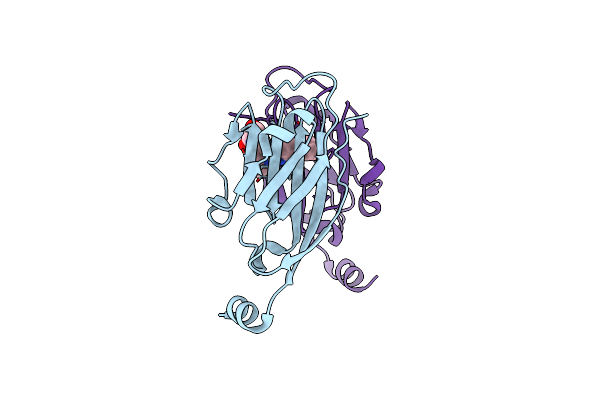 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2025-03-12 Classification: IMMUNE SYSTEM Ligands: A1D9R |
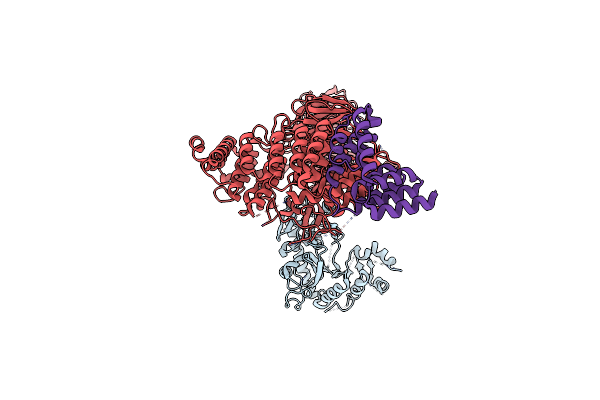 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-08-14 Classification: PROTEIN TRANSPORT |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-08-14 Classification: PROTEIN TRANSPORT |
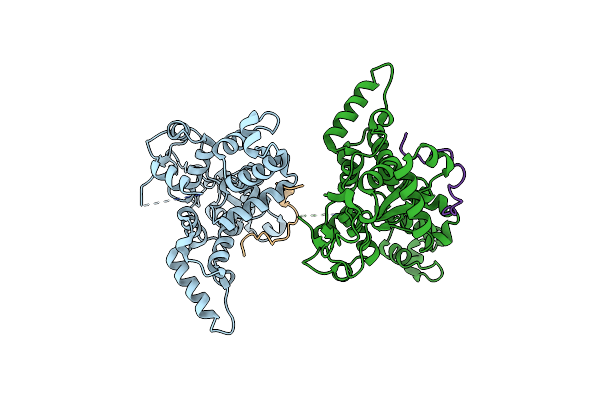 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2024-04-24 Classification: TRANSPORT PROTEIN/UNKNOWN FUNCTION |
 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-01-24 Classification: HYDROLASE Ligands: MN |
 |
Crystal Structure Of Yeast Cytosine Deaminase Mutant Ycd-Rq-1/8Sah In Complex With (R)-4-Hydroxy-3,4-Dihydropyrimidin-2(1H)-One
Organism: Saccharomyces cerevisiae s288c
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2024-01-24 Classification: HYDROLASE Ligands: OS0, ZN, EDO |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-09-27 Classification: PROTEIN TRANSPORT Ligands: NAG |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-09-27 Classification: PROTEIN TRANSPORT Ligands: NAG, CLR |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-09-27 Classification: PROTEIN TRANSPORT Ligands: NAG |
 |
Organism: African swine fever virus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2023-09-13 Classification: DNA BINDING PROTEIN |

