Search Count: 45
All
Selected
 |
Solution Structure Of The Translation Initiation Factor If-1 From Neisseria Gonorrhoeae (Nccp11945). Seattle Structural Genomics Center For Infectious Disease Target Negoa.17902.A
Organism: Neisseria gonorrhoeae (strain atcc 700825 / fa 1090)
Method: SOLUTION NMR Release Date: 2024-10-16 Classification: RNA BINDING PROTEIN |
 |
Thermus Thermophilus Initiation Transcription Complex In The Pre-Translocated State
Organism: Thermus thermophilus hb8, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2024-07-24 Classification: TRANSCRIPTION/DNA/RNA Ligands: MG, ZN |
 |
Organism: Thermus thermophilus hb8, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2024-07-24 Classification: TRANSCRIPTION/DNA/RNA Ligands: MG, ZN |
 |
Thermus Thermophilus Initiation Transcription Complex Containing Cmpcpp In The Post-Translocated State
Organism: Thermus thermophilus hb8, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.17 Å Release Date: 2024-07-24 Classification: TRANSCRIPTION/DNA/RNA Ligands: MG, ZN, 2TM |
 |
Organism: Trichomonas vaginalis g3
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2024-05-08 Classification: LIPID BINDING PROTEIN |
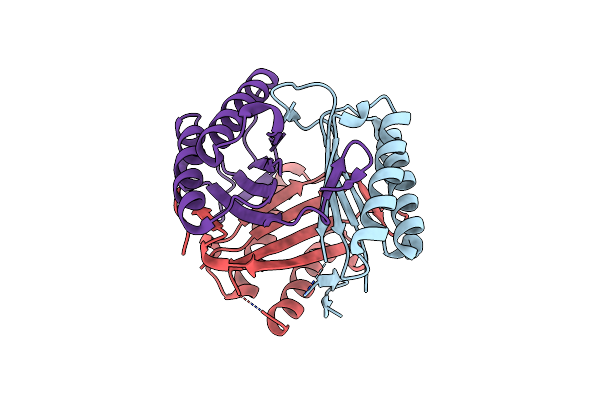 |
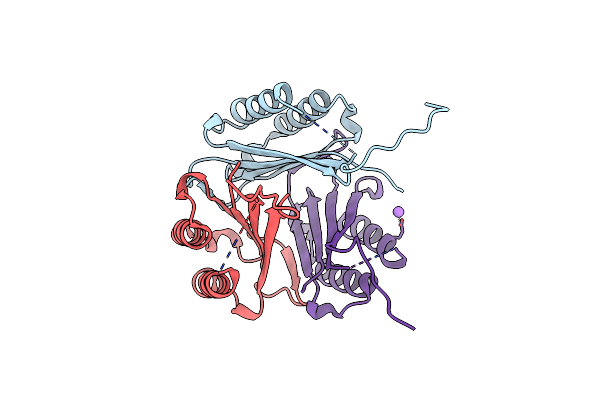
Crystal Structure Of Macrophage Migration Inhibitory Factor (Mif) From Trichomonas Vaginalis (I4122 Form)
Organism: Trichomonas vaginalis g3
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2024-03-20 Classification: CYTOKINE Ligands: CL |
 |
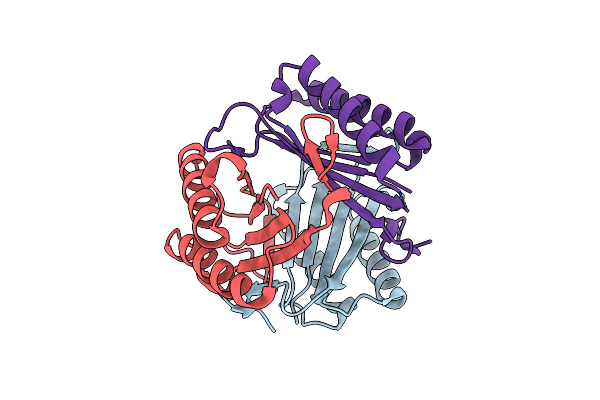
Crystal Structure Of Macrophage Migration Inhibitory Factor-1 (Mif1) From Onchocerca Volvulus
Organism: Onchocerca volvulus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2024-01-17 Classification: CYTOKINE |
 |
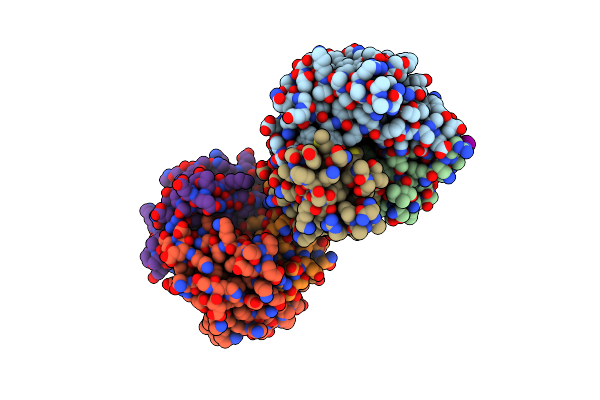
Crystal Structure Of Macrophage Migration Inhibitory Factor (Mif) From Trichomonas Vaginalis (Apo, P41212 Form)
Organism: Trichomonas vaginalis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2023-12-06 Classification: CYTOKINE |
 |
Crystal Structure Of Macrophage Migration Inhibitory Factor (Mif) From Trichomonas Vaginalis (I41 Form)
Organism: Trichomonas vaginalis g3
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2023-11-08 Classification: CYTOKINE Ligands: IOD, PYR |
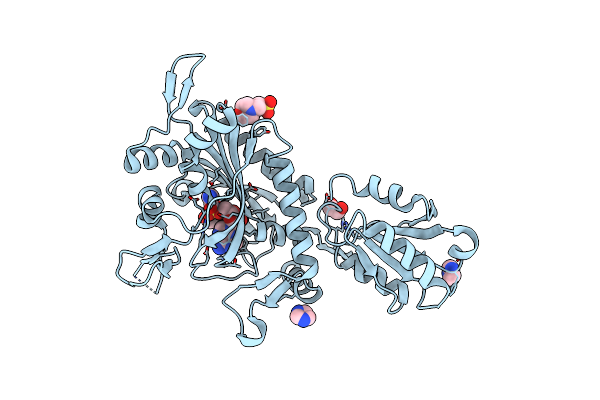 |
Crystal Structure Of Glycine--Trna Ligase From Mycobacterium Tuberculosis (G5A Bound)
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2023-09-13 Classification: TRANSFERASE Ligands: G5A, IMD, MES, EDO, MG, CL |
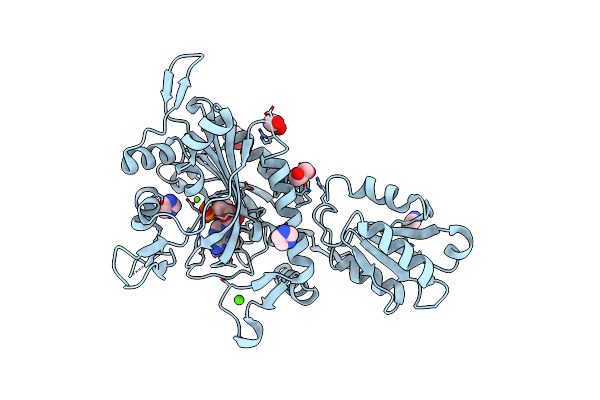 |
Crystal Structure Of Glycine--Trna Ligase From Mycobacterium Tuberculosis (Amp-Mg Bound)
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2023-06-28 Classification: LIGASE Ligands: MG, CA, EDO, GOL, IMD, AMP |
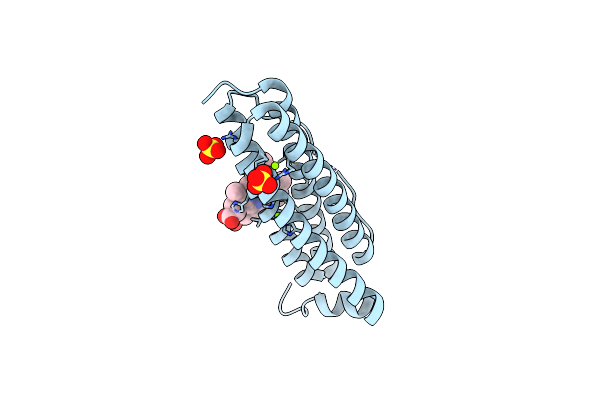 |
Crystal Structure Of Bacterioferritin (Bfr) From Brucella Abortus (Magnesium Bound, F16L Mutant)
Organism: Brucella abortus 2308
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2023-05-17 Classification: METAL BINDING PROTEIN Ligands: MG, SO4, CL, HEM |
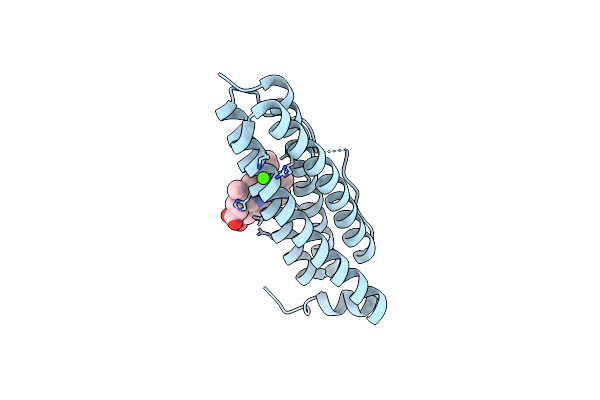 |
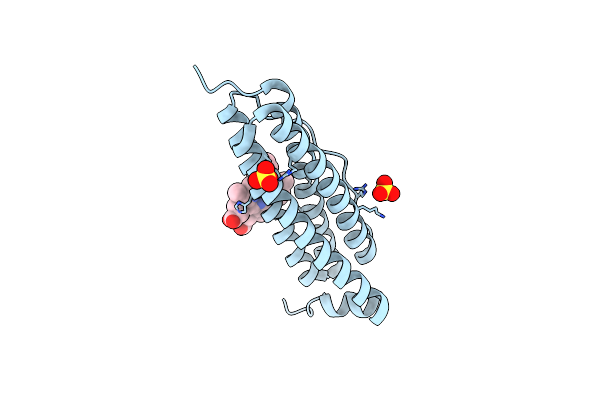
Crystal Structure Of Bacterioferritin (Bfr) From Brucella Abortus (Apo, F16L Mutant)
Organism: Brucella abortus 2308
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2023-05-17 Classification: METAL BINDING PROTEIN Ligands: CL, CA, HEM |
 |
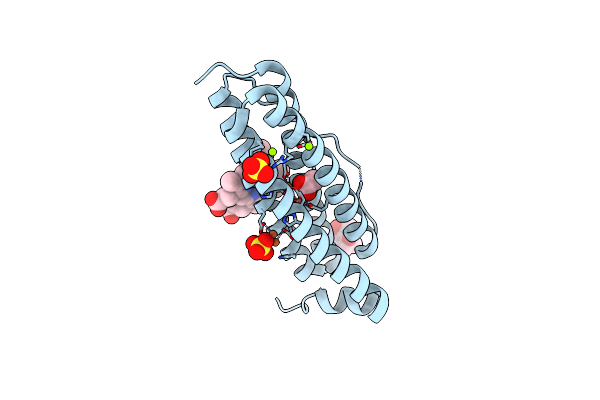
Crystal Structure Of Bacterioferritin (Bfr) From Brucella Abortus (Apo Cubic Form 2, F16L Mutant)
Organism: Brucella abortus 2308
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2023-05-17 Classification: METAL BINDING PROTEIN Ligands: CL, SO4, HEM |
 |
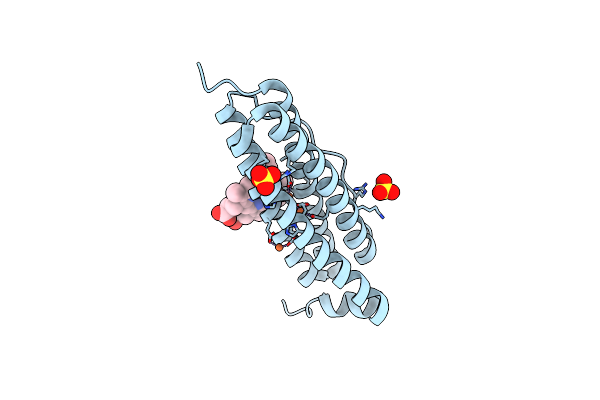
Crystal Structure Of Bacterioferritin (Bfr) From Brucella Abortus (Iron Bound, F16L Mutant)
Organism: Brucella abortus 2308
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2023-05-17 Classification: METAL BINDING PROTEIN Ligands: MG, SO4, CL, FE2, ACT, HEM, MPD |
 |
Crystal Structure Of Bacterioferritin (Bfr) From Brucella Abortus (Iron Bound, Cubic Form 2, F16L Mutant)
Organism: Brucella abortus 2308
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2023-05-17 Classification: METAL BINDING PROTEIN Ligands: SO4, CL, FE2, HEM |
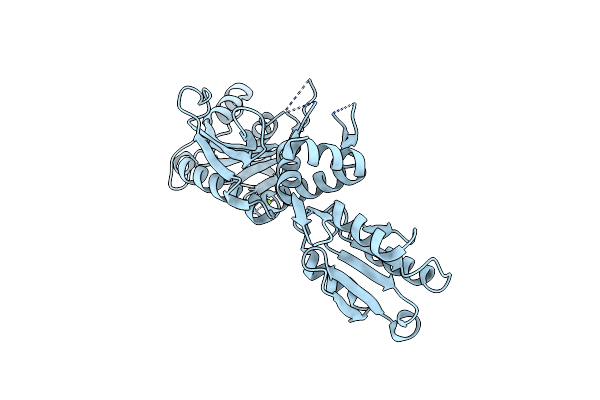 |
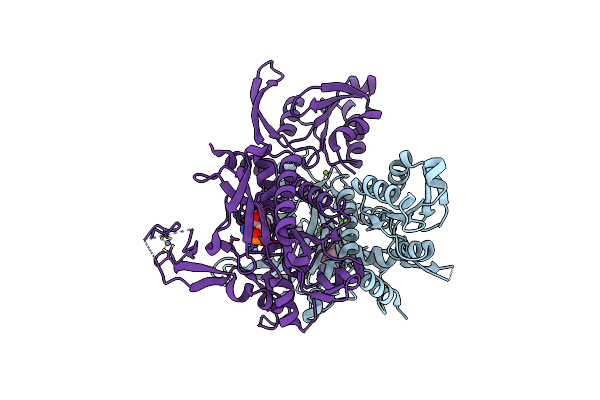
Crystal Structure Of Glycine Trna Ligase From Mycobacterium Thermoresistibile (Apo)
Organism: Mycolicibacterium thermoresistibile atcc 19527
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2023-05-03 Classification: LIGASE Ligands: MG |
 |
Crystal Structure Of Glycine Trna Ligase From Mycobacterium Thermoresistibile (Amp Bound)
Organism: Mycolicibacterium thermoresistibile atcc 19527
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2023-05-03 Classification: LIGASE Ligands: AMP, MG, ZN |
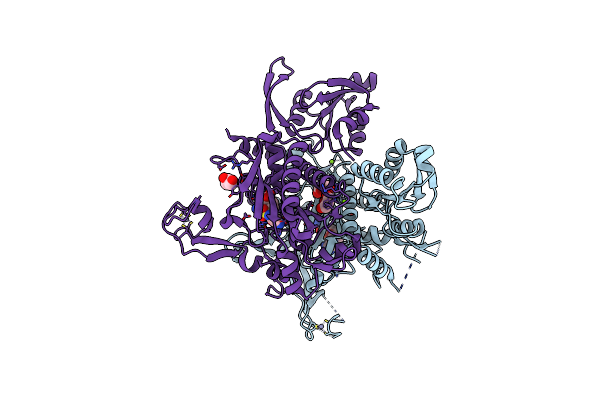 |
Crystal Structure Of Glycine Trna Ligase From Mycobacterium Thermoresistibile (Glycyl Adenylate Bound)
Organism: Mycolicibacterium thermoresistibile atcc 19527
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2023-05-03 Classification: LIGASE Ligands: G5A, ZN, MG, GOL |
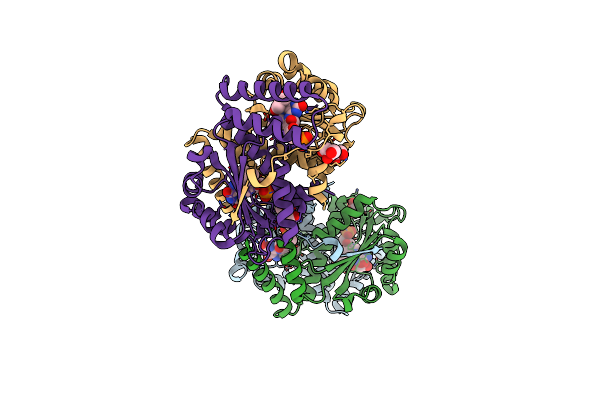 |
Crystal Structure Of Dihydropteridine Reductase/Oxygen-Insensitive Nad(P)H Nitroreductase From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2022-07-27 Classification: OXIDOREDUCTASE Ligands: FMN, PO4, GOL |

