Search Count: 5
All
Selected
 |
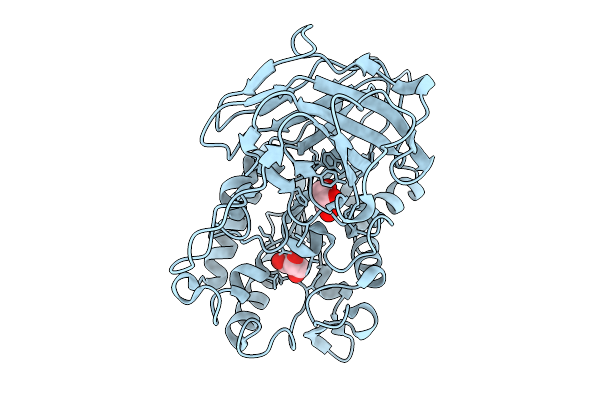
Structural Basis For The Polymer-Protein Binding Mechanism Of Polyvinyl Alcohol Esterase
Organism: Comamonas sp. nyz500
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2025-10-08 Classification: HYDROLASE Ligands: SO4, ACT, GOL, CL, NA |
 |
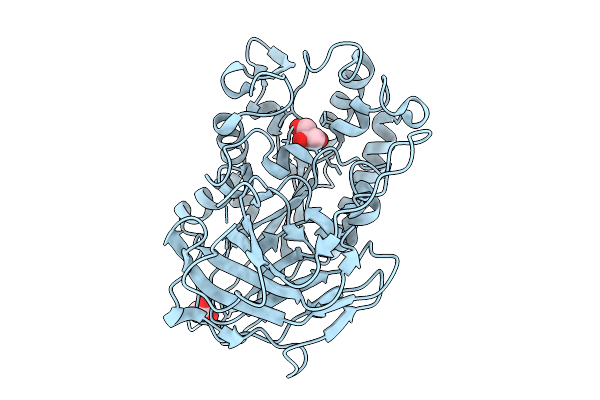
Structural Basis For The Polymer-Protein Binding Mechanism Of Polyvinyl Alcohol Esterase
Organism: Comamonas sp. nyz500
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-10-08 Classification: HYDROLASE Ligands: GOL |
 |
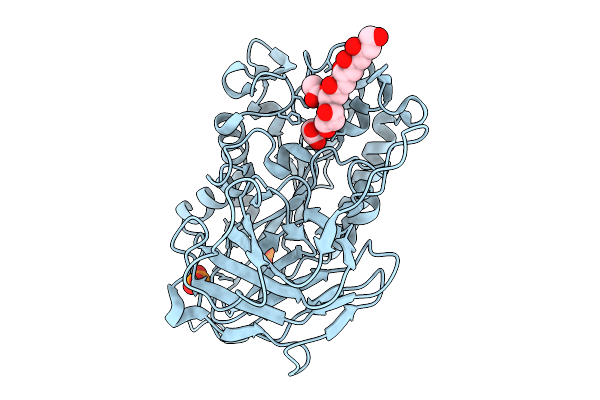
Structural Basis For The Polymer-Protein Binding Mechanism Of Polyvinyl Alcohol Esterase
Organism: Comamonas sp. nyz500
Method: X-RAY DIFFRACTION Resolution:2.84 Å Release Date: 2025-10-08 Classification: HYDROLASE Ligands: CL, GOL, ACT |
 |
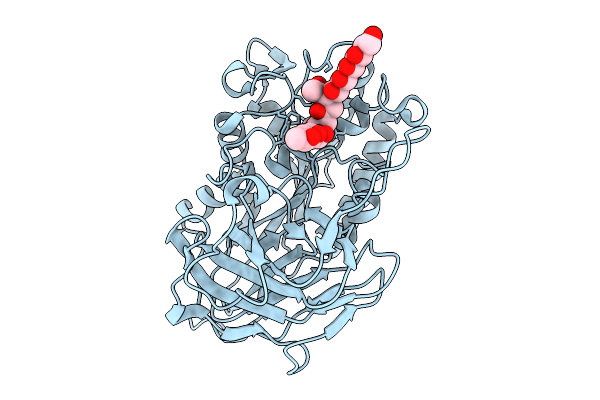
Structural Basis For The Polymer-Protein Binding Mechanism Of Polyvinyl Alcohol Esterase
Organism: Comamonas sp. nyz500
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2025-10-08 Classification: HYDROLASE Ligands: CL, A1EGH, ACT, PO4 |
 |
Structural Basis For The Polymer-Protein Binding Mechanism Of Polyvinyl Alcohol Esterase
Organism: Comamonas sp. nyz500
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2025-10-08 Classification: HYDROLASE Ligands: A1EGL, CL |

