Search Count: 146
All
Selected
 |
Crystal Structure Of The Complex Of The Gh16 Carrageenase Sfgh16 From Saccharicrinis Fermentans With An Oligotetrasaccharide Of Kappa-Carrageenan
Organism: Saccharicrinis fermentans dsm 9555 = jcm 21142
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-03-11 Classification: HYDROLASE |
 |
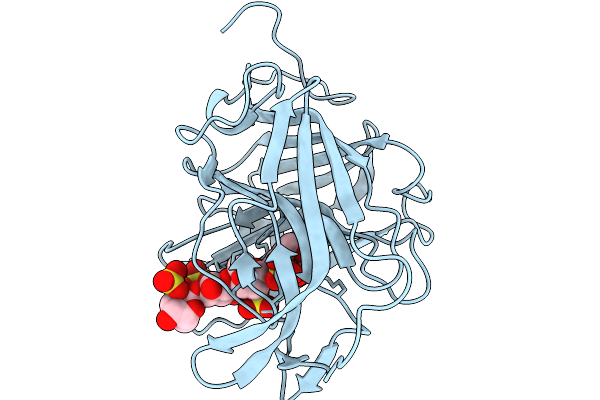
Crystal Structure Of The Gh16 Carrageenase Sfgh16 From Saccharicrinis Fermentans
Organism: Saccharicrinis fermentans dsm 9555 = jcm 21142
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-11 Classification: HYDROLASE |
 |
Crystal Structure Of The Complex Of The Gh16 Carrageenase Sfgh16 From Saccharicrinis Fermentans With An Oligotetrasaccharide Of Iota-Carrageenan
Organism: Saccharicrinis fermentans dsm 9555 = jcm 21142
Method: X-RAY DIFFRACTION Resolution:1.44 Å Release Date: 2026-03-04 Classification: HYDROLASE |
 |
Crystal Structure Of Retro-Aldolase Ra95.5-8 Mutant With A Covalently Bound Cyclohexenone Derivative
Organism: Saccharolobus solfataricus
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: SO4, EDO |
 |
Crystal Structure Of Retro-Aldolase T53L/K210H Ra95.5-8 With A Covalently Bound Cyclohexenone Derivative
Organism: Saccharolobus solfataricus
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: SO4, EDO |
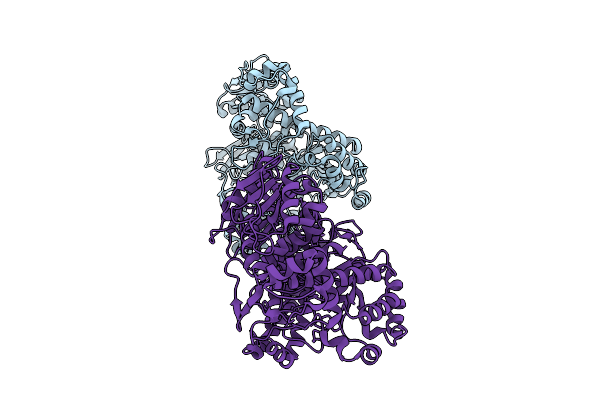 |
Organism: Wenyingzhuangia aestuarii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-10-01 Classification: HYDROLASE |
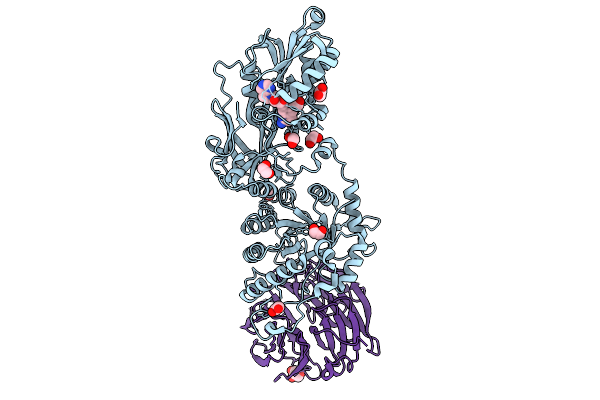 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-09-10 Classification: TRANSLOCASE Ligands: A1CHO, EDO, GOL |
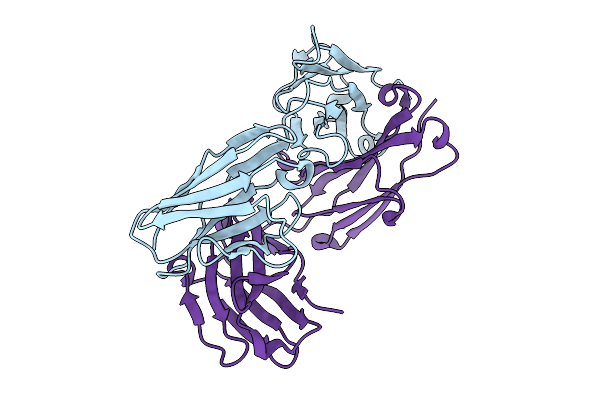 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2025-08-27 Classification: IMMUNE SYSTEM |
 |
Organism: Amycolatopsis thermoflava
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2025-07-23 Classification: OXIDOREDUCTASE Ligands: NAP, A1ECB |
 |
Organism: Amycolatopsis thermoflava
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2025-07-23 Classification: OXIDOREDUCTASE Ligands: PGE, PEG, CL, CA, NDP |
 |
Organism: Homo sapiens, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2025-06-25 Classification: TRANSCRIPTION Ligands: ZN, MN, MG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.69 Å Release Date: 2025-05-07 Classification: GENE REGULATION Ligands: UNX |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2025-05-07 Classification: GENE REGULATION Ligands: UNX |
 |
Organism: Chaetomium thermophilum (strain dsm 1495 / cbs 144.50 / imi 039719)
Method: ELECTRON MICROSCOPY Release Date: 2025-03-12 Classification: SPLICING/RNA Ligands: M7M, MG, GTP, IHP, ZN |
 |
Organism: Chaetomium thermophilum (strain dsm 1495 / cbs 144.50 / imi 039719)
Method: ELECTRON MICROSCOPY Release Date: 2025-03-12 Classification: SPLICING/RNA Ligands: M7M, MG, GTP, ZN |
 |
Organism: Chaetomium thermophilum (strain dsm 1495 / cbs 144.50 / imi 039719)
Method: ELECTRON MICROSCOPY Release Date: 2025-03-12 Classification: SPLICING/RNA Ligands: M7M, GTP, ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: TRANSCRIPTION Ligands: ZN, MN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-02-12 Classification: RIBOSOME Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-02-12 Classification: RIBOSOME Ligands: ZN |
 |
Cryo-Em Structure Of The Human Ubiquitylated Pre-40S Ribosome With Riok3 (Without Nob1)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-02-12 Classification: RIBOSOME Ligands: ZN |

