Search Count: 26
All
Selected
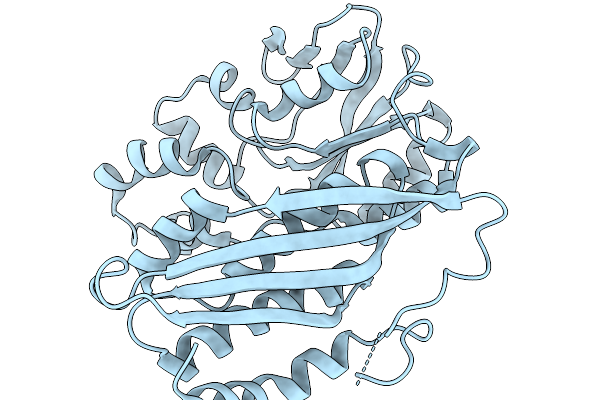 |
Crystal Structure Of Chimeric Air Synthetase Constructed From E. Coli (Residues 1-168) And Pyrococcus Abyssi (Residues 163-334)
Organism: Escherichia coli, Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-11 Classification: LIGASE |
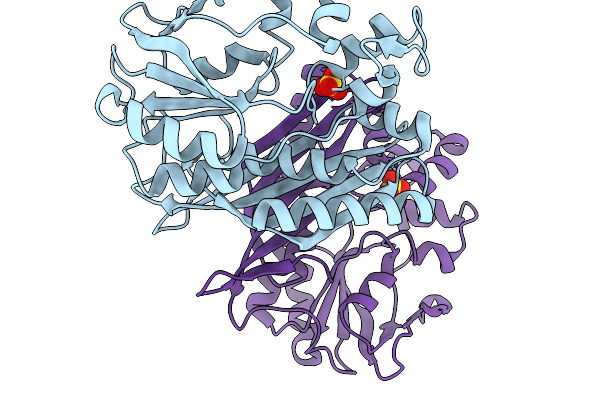 |
Crystal Structure Of Pyrococcus Abyssi Air Synthetase With An N-Terminal 43-Residues Deletion
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-11 Classification: LIGASE Ligands: SO4 |
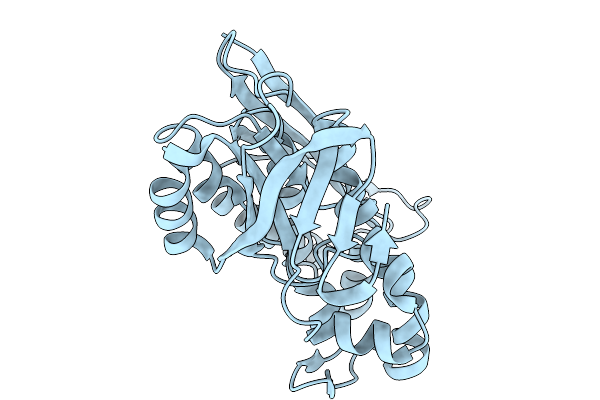 |
Crystal Structure Of Pyrococcus Abyssi Air Synthetase With An N-Terminal 54-Residues Deletion
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-11 Classification: LIGASE |
 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: SO4 |
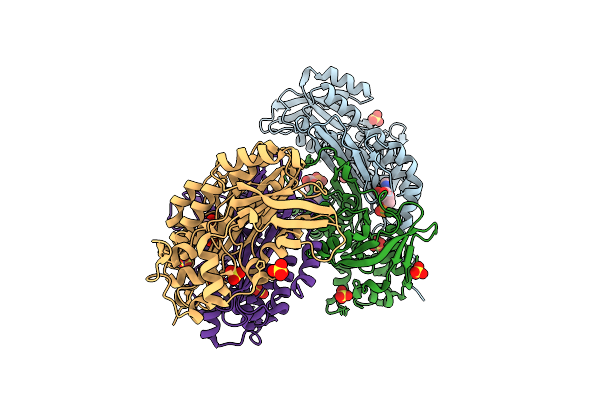 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: SO4, AMP |
 |
Organism: Pyrococcus horikoshii ot3
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: ZN, CL |
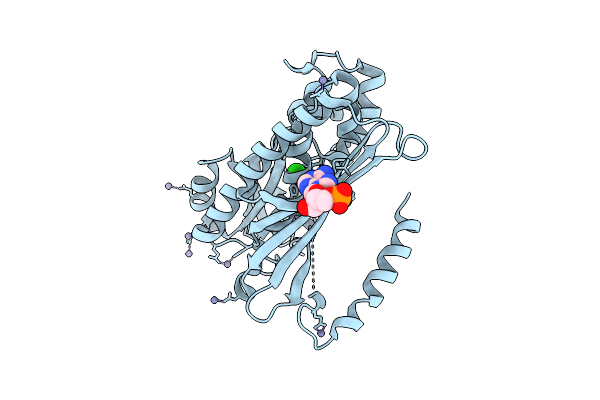 |
Organism: Pyrococcus horikoshii ot3
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: AMP, ZN, CL |
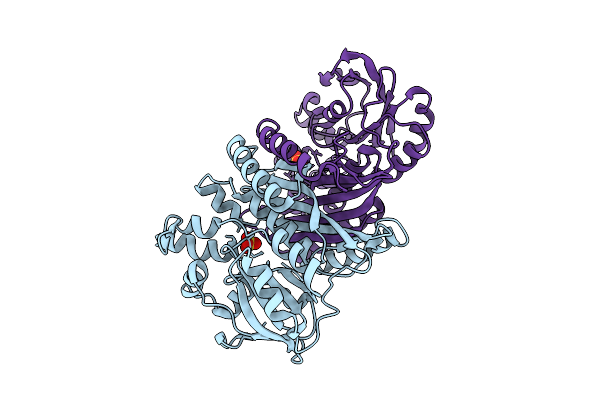 |
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: SO4 |
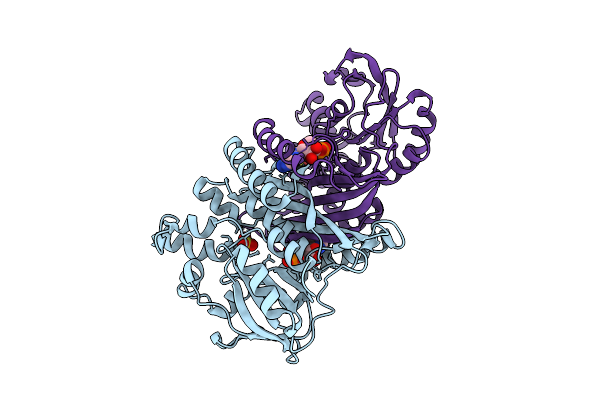 |
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: SO4, ADP |
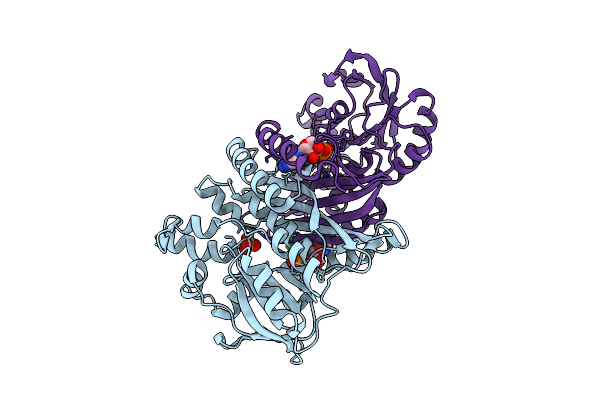 |
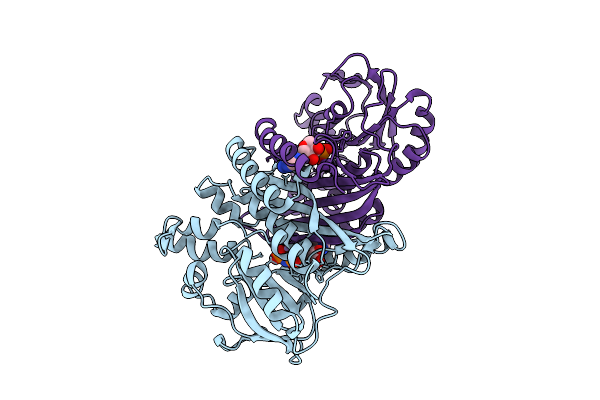
Crystal Structure Of Pyrococcus Abyssi Air Synthetase Bound To Adp And Mg(Ii)
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: SO4, ADP, MG |
 |
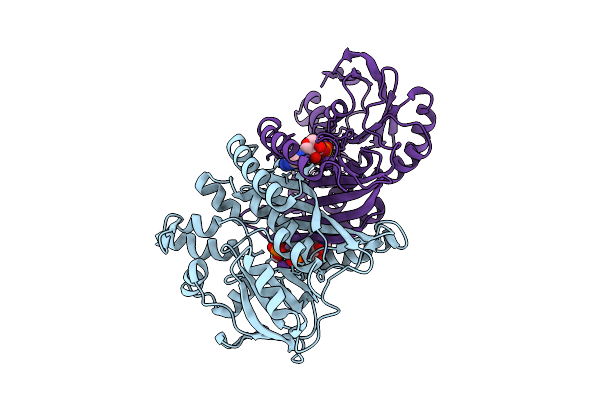
Crystal Structure Of Pyrococcus Abyssi Air Synthetase H239A Mutant Bound To Amppnp
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: ANP, MN, K |
 |
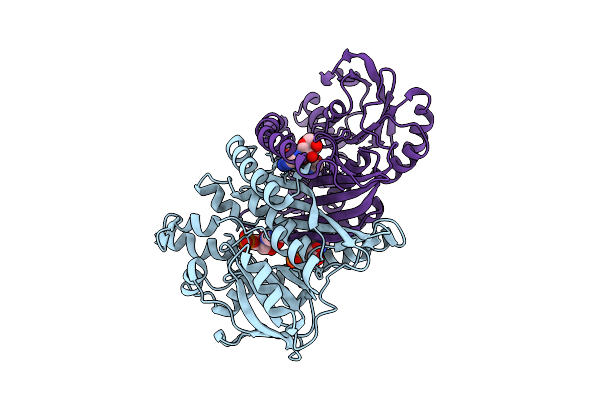
Crystal Structure Of Pyrococcus Abyssi Air Synthetase H239A Mutant Bound To Adp And Pi
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: ADP, PO4, MN, K |
 |
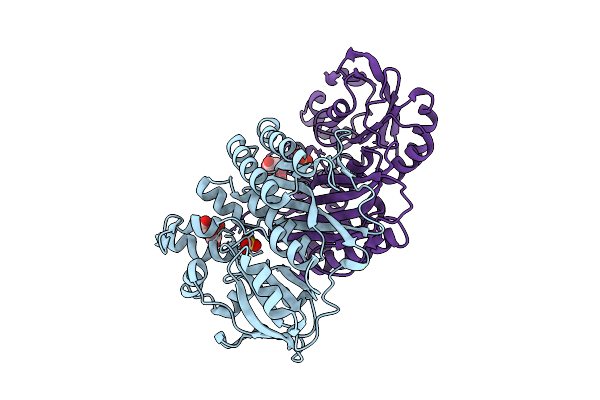
Crystal Structure Of Pyrococcus Abyssi Air Synthetase N186A Mutant Bound To Fgar And Amp
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: FGR, AMP |
 |
Crystal Structure Of Pyrococcus Abyssi Air Synthetase With Residues 51-54 Replaced By Neisseria Gonorrhoeae Residues 49-56
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: SO4, FLC |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-01-17 Classification: PROTEIN FIBRIL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2024-01-17 Classification: STRUCTURAL PROTEIN Ligands: MPD |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2005-12-27 Classification: IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Sars coronavirus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2005-11-22 Classification: HYDROLASE |
 |
Organism: Sars coronavirus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2005-11-22 Classification: HYDROLASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2005-03-01 Classification: CHAPERONE |

