Search Count: 48
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-04-01 Classification: CELL CYCLE Ligands: A1MBI |
 |
Crystal Structure Of Multidrug Efflux Transporter Oqxb From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2025-06-18 Classification: MEMBRANE PROTEIN Ligands: LMT, GOL |
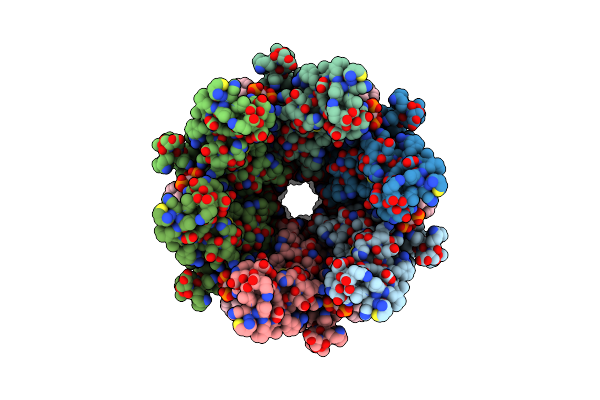 |
Structure Of Cx43/Gja1 Gap Junction Intercellular Channel In Complex With Dic8-Pip2
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-06-11 Classification: MEMBRANE PROTEIN Ligands: PIO |
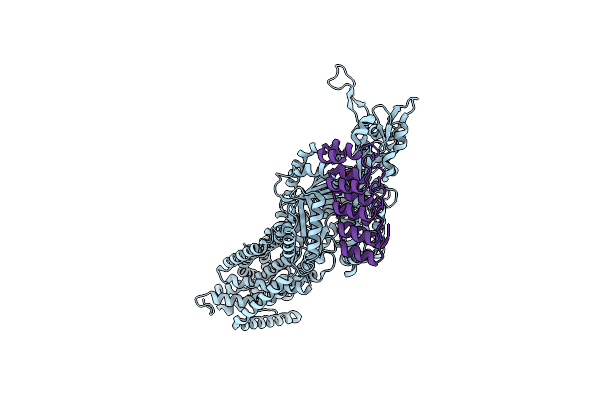 |
Organism: Escherichia coli k-12, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-06-11 Classification: TRANSPORT PROTEIN Ligands: MIY |
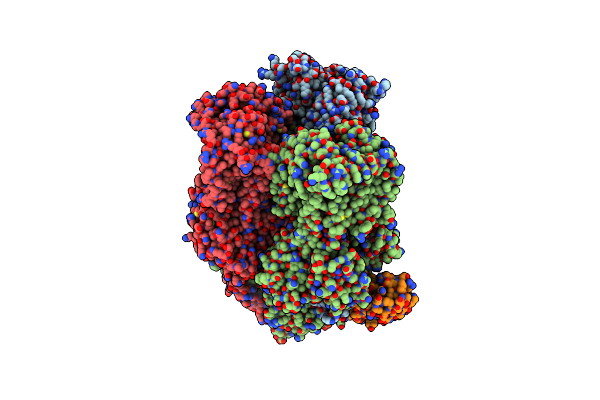 |
Organism: Escherichia coli k-12, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2025-06-11 Classification: TRANSPORT PROTEIN |
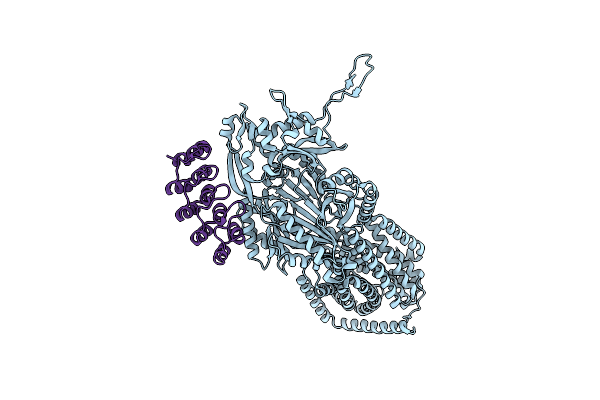 |
Organism: Escherichia coli k-12, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.55 Å Release Date: 2025-06-11 Classification: TRANSPORT PROTEIN |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-05-28 Classification: TRANSPORT PROTEIN |
 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2025-05-28 Classification: TRANSPORT PROTEIN |
 |
Single Particle Cryo-Em Structure Of The Multidrug Efflux Pump Oqxb From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae
Method: ELECTRON MICROSCOPY Release Date: 2025-05-28 Classification: TRANSPORT PROTEIN |
 |
Organism: Synthetic construct, Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2025-05-28 Classification: TRANSPORT PROTEIN Ligands: MIY |
 |
Organism: Synthetic construct, Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-05-28 Classification: TRANSPORT PROTEIN |
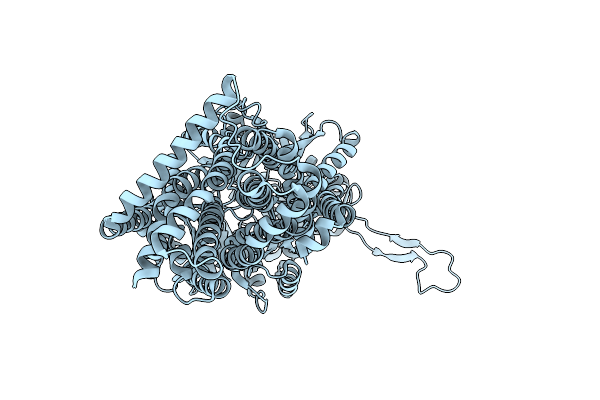 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-05-28 Classification: TRANSPORT PROTEIN |
 |
Organism: Enhygromyxa salina
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2024-11-27 Classification: OXIDOREDUCTASE Ligands: CD |
 |
Structure Of C-Terminal Truncated Connexin43/Cx43/Gja1 Gap Junction Intercellular Channel In Pope/Chs Nanodiscs (C1 Symmetry)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-02-22 Classification: MEMBRANE PROTEIN Ligands: C14, PTY, Y01 |
 |
Structure Of C-Terminal Truncated Connexin43/Cx43/Gja1 Gap Junction Intercellular Channel In Pope/Chs Nanodiscs
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-02-22 Classification: MEMBRANE PROTEIN Ligands: C14, PTY, Y01 |
 |
Structure Of Connexin43/Cx43/Gja1 Gap Junction Intercellular Channel In Pope/Chs Nanodiscs At Ph ~8.0
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-02-01 Classification: MEMBRANE PROTEIN Ligands: C14, Y01, PTY |
 |
Structure Of Connexin43/Cx43/Gja1 Gap Junction Intercellular Channel In Gdn Detergents At Ph ~8.0
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-01-25 Classification: MEMBRANE PROTEIN |
 |
Hemichannel-Focused Structure Of C-Terminal Truncated Connexin43/Cx43/Gja1 Gap Junction Intercellular Channel In Pope Nanodiscs (Gcn Conformation)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-01-25 Classification: MEMBRANE PROTEIN Ligands: PTY, Y01 |
 |
Hemichannel-Focused Structure Of C-Terminal Truncated Connexin43/Cx43/Gja1 Gap Junction Intercellular Channel In Pope Nanodiscs (Gcn-Tm1I Conformation)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-01-25 Classification: MEMBRANE PROTEIN |
 |
Hemichannel-Focused Structure Of C-Terminal Truncated Connexin43/Cx43/Gja1 Gap Junction Intercellular Channel In Pope Nanodiscs (Fin Conformation)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2023-01-25 Classification: MEMBRANE PROTEIN |

