Search Count: 53
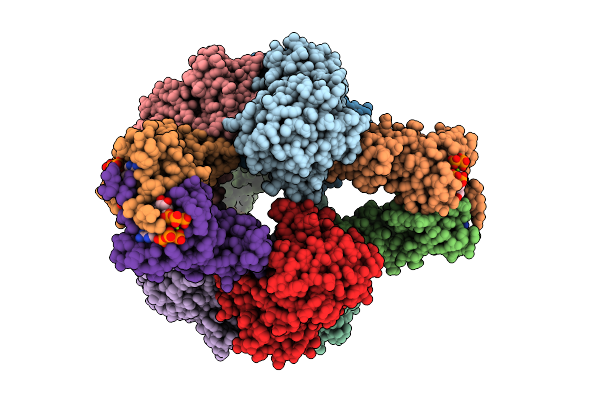 |
Crystal Structure Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Cp, Succinate, Atp, And Mg2+
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: CP, SIN, ZN, ATP, MG |
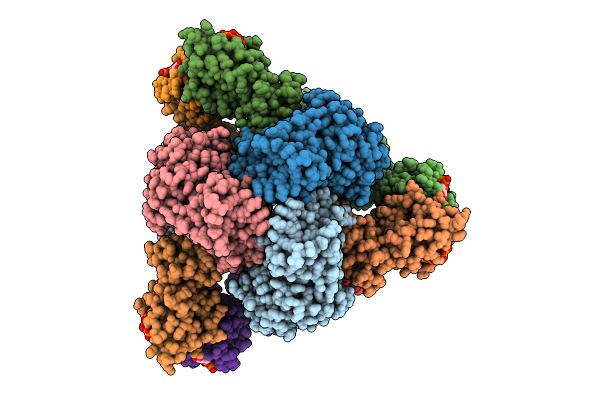 |
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The T-State Complexed With Cp, Ctp, Utp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: ZN, CTP, UTP, MG, CP |
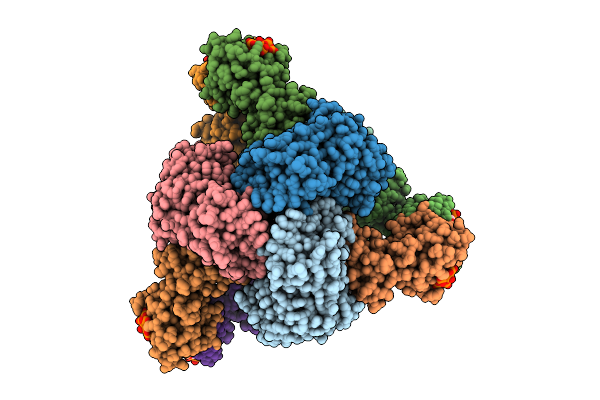 |
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Cp, Succinate, Ctp, Utp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: CP, SIN, ZN, CTP, UTP, MG |
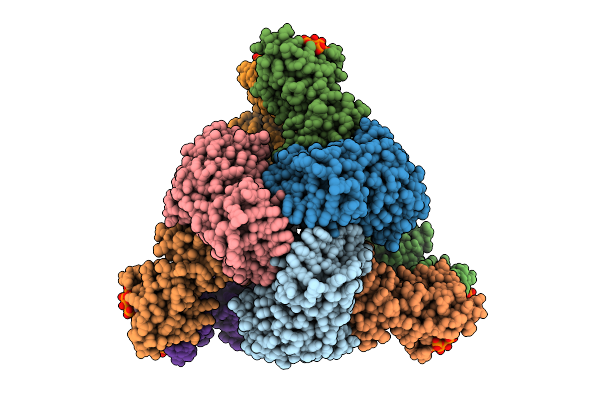 |
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Cp, Succinate, Ctp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: CP, SIN, ZN, CTP, MG |
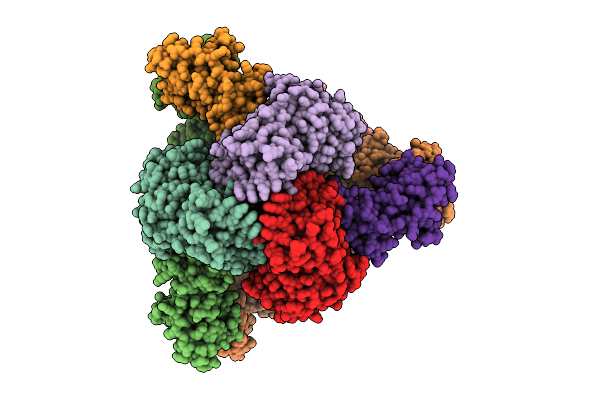 |
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Cp And Succinate
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: CP, SIN, ZN |
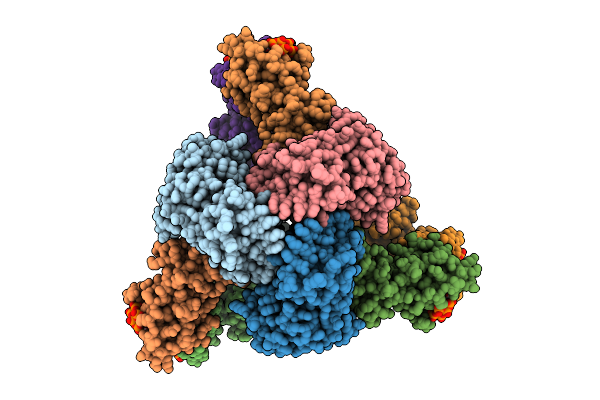 |
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Cp, Succinate, Atp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: CP, SIN, ZN, ATP, MG |
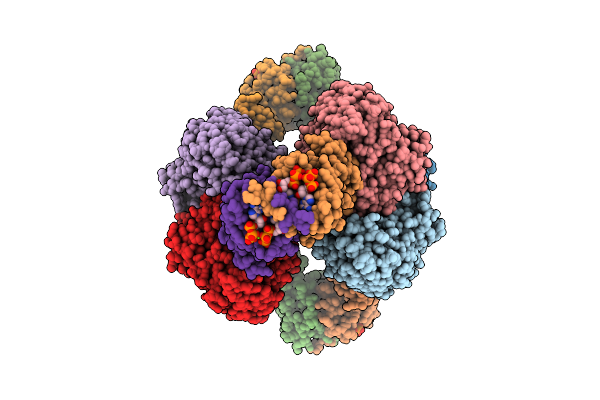 |
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The R-State Complexed With Cp, Succinate, Atp, Gtp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: CP, SIN, ZN, ATP, GTP, MG |
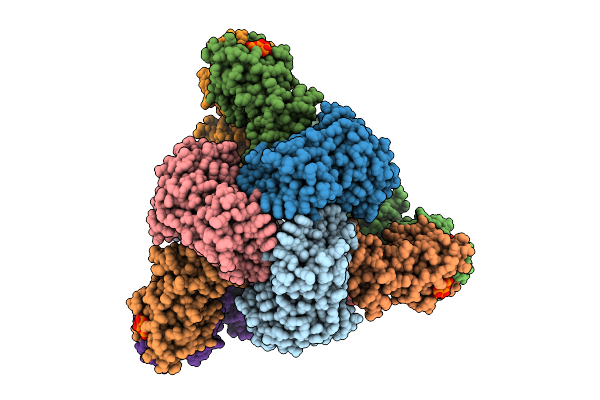 |
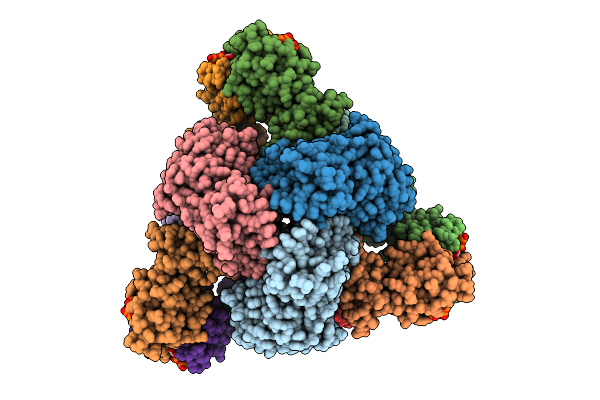
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In An Expanded State Complexed With Cp, Atp, Gtp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: ZN, ATP, GTP, MG, CP |
 |
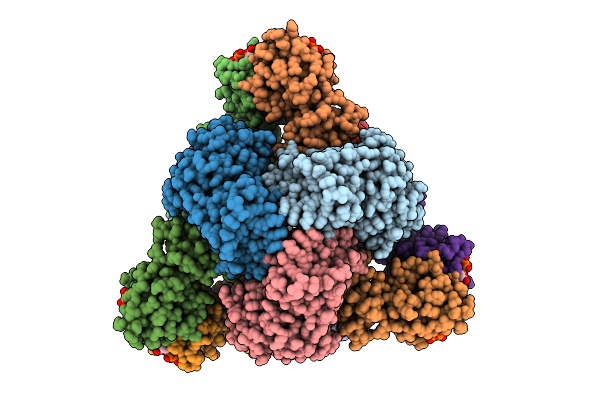
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The T-State Complexed With Cp, Atp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CYTOSOLIC PROTEIN Ligands: ZN, ATP, MG, CP |
 |
Cryo-Em Model Of E. Coli Aspartate Transcarbamoylase In The T-State Complexed With Cp, Ctp, And Mg2+
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: CYTOSOLIC PROTEIN Ligands: ZN, CTP, MG, CP |
 |
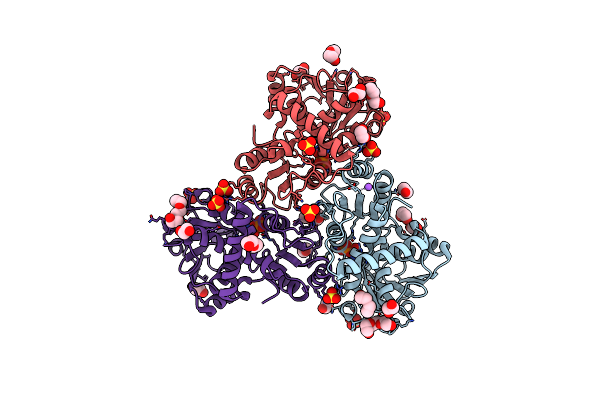
Crystal Structure Of Ornithine Carbamoyltransferase From Burkholderia Xenovorans In Complex With Phosphono Carbamate
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:2.67 Å Release Date: 2025-09-24 Classification: TRANSFERASE Ligands: CP, PO4, CL |
 |
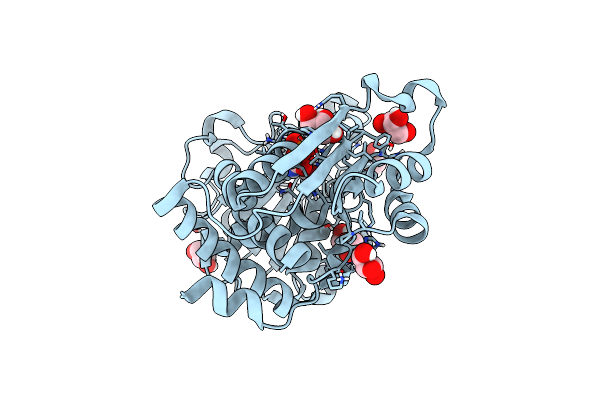
Crystal Structure Of Ornithine Transcarbamylase From Arabidopsis Thaliana (Atotc) In Complex With Carbamoyl Phosphate
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2023-12-06 Classification: TRANSFERASE Ligands: CP, EDO, PGE, PEG, SO4, NA |
 |
Aspartate Transcarbamoylase Mutant (N2045C, R2238C) From Chaetomium Thermophilum Cad-Like Bound To Carbamoyl Phosphate
Organism: Thermochaetoides thermophila
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2023-02-01 Classification: TRANSFERASE Ligands: CP, GOL, SIN |
 |
Organism: Streptomyces hygroscopicus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2022-11-16 Classification: TRANSFERASE Ligands: ATP, FE, GOL, SO4, EDO, CP, PO4 |
 |
Organism: Streptomyces hygroscopicus
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2022-11-16 Classification: TRANSFERASE Ligands: FE, CA0, SO4, CP, EDO, GOL, PEG |
 |
Organism: Streptomyces hygroscopicus
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2022-11-16 Classification: TRANSFERASE Ligands: SO4, FE, GOL, EDO, PEG, CP, CL |
 |
The Structure Of Gdmn Complex With The Natural Tetrahedral Intermediate, Carbamoylated Derivative, And Amp
Organism: Streptomyces hygroscopicus
Method: X-RAY DIFFRACTION Resolution:2.88 Å Release Date: 2022-11-16 Classification: TRANSFERASE Ligands: FE, SO4, CP, EDO, 82Z, AMP, 8CW |
 |
The Structure Of Gdmn In Complex With Carbamoyl Adenylate Intermediate And 20-O-Methyl-19-Chloroproansamitocin
Organism: Streptomyces hygroscopicus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2022-11-16 Classification: TRANSFERASE Ligands: FE, 83Z, CA0, PEG, EDO, GOL, CP, SO4 |
 |
The Structure Of Gdmn V24Y/G157A/R158A/G188R Mutant In Complex With Carbamoyl Adenylate Intermediate
Organism: Streptomyces hygroscopicus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2022-11-16 Classification: TRANSFERASE Ligands: CP, FE, EDO, CA0, SO4, CL |
 |
The Structure Of Gdmn Complex With Amp And 20-O-Methyl-19-Chloroproansamitocin
Organism: Streptomyces hygroscopicus
Method: X-RAY DIFFRACTION Resolution:2.81 Å Release Date: 2022-11-16 Classification: TRANSFERASE Ligands: 83Z, FE, AMP, GOL, SO4, CP, EDO, PEG |

