Search Count: 334
All
Selected
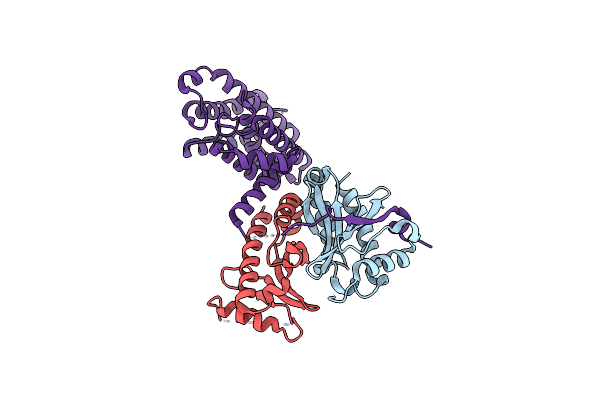 |
Crystal Structure Of The Salmonella Secretion Chaperone Invb In Complex With Sipa
Organism: Salmonella typhimurium
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2006-03-21 Classification: CHAPERONE/CELL INVASION |
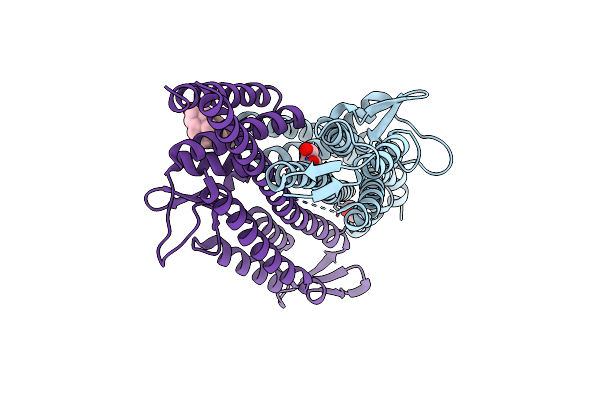 |
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2011-08-17 Classification: CELL INVASION Ligands: GOL, DXC |
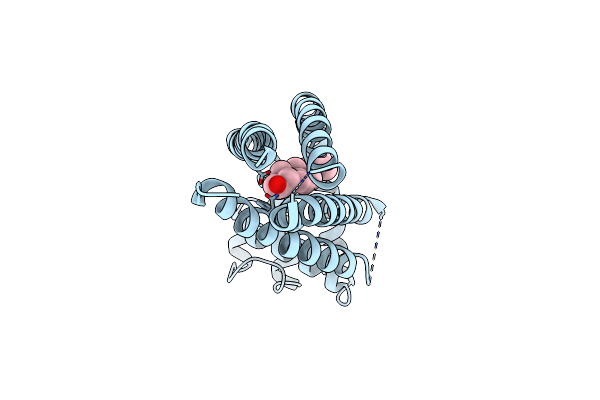 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.96 Å Release Date: 2016-11-09 Classification: CELL INVASION Ligands: CLR |
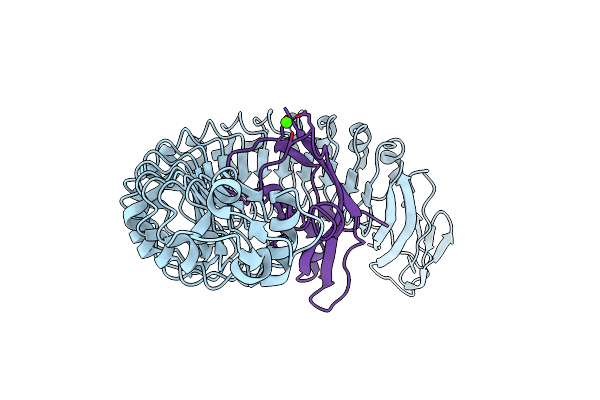 |
Organism: Listeria monocytogenes, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2007-06-05 Classification: CELL INVASION/CELL ADHESION Ligands: CA, CL |
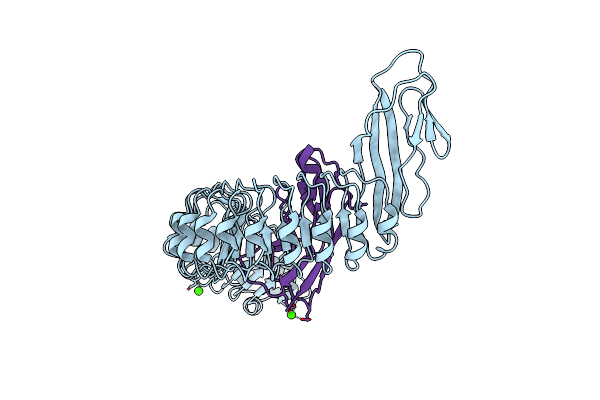 |
Organism: Listeria monocytogenes, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2007-06-05 Classification: CELL INVASION/CELL ADHESION Ligands: CA, CL |
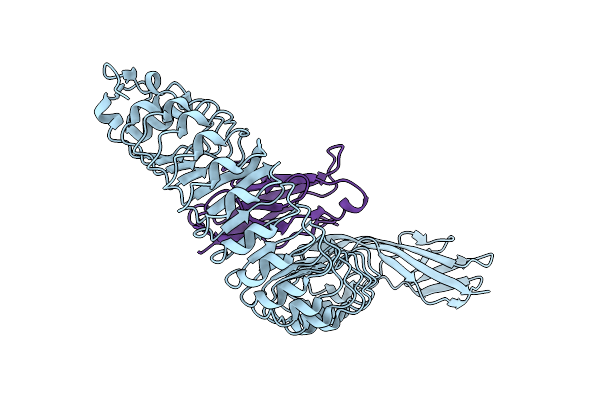 |
Organism: Listeria monocytogenes, Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2007-06-05 Classification: CELL INVASION/CELL ADHESION Ligands: CL |
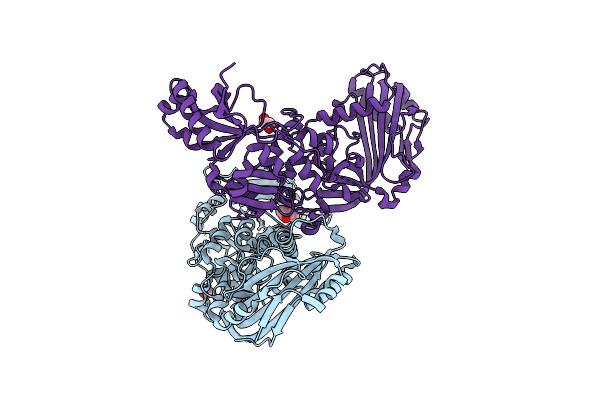 |
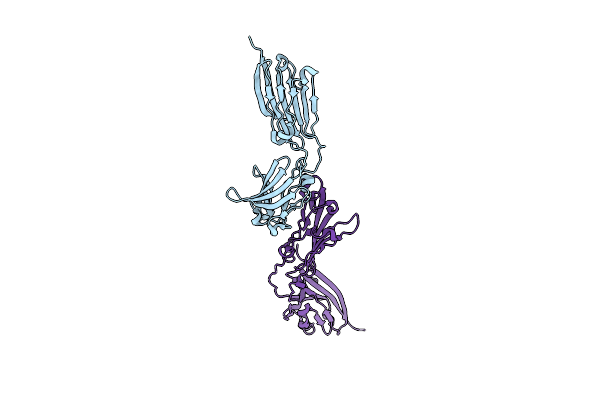
Crystal Structure Of Putative Cell Invasion Protein With Mac/Perforin Domain (Np_812351.1) From Bacteriodes Thetaiotaomicron Vpi-5482 At 2.46 A Resolution
Organism: Bacteroides thetaiotaomicron vpi-5482
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2009-11-24 Classification: CELL INVASION Ligands: CL, EDO, MPD |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2019-05-22 Classification: CELL INVASION |
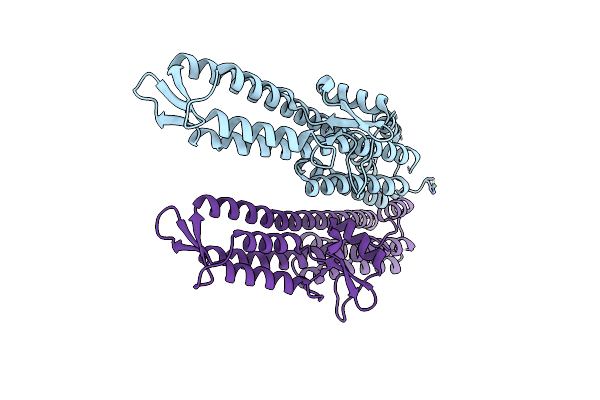 |
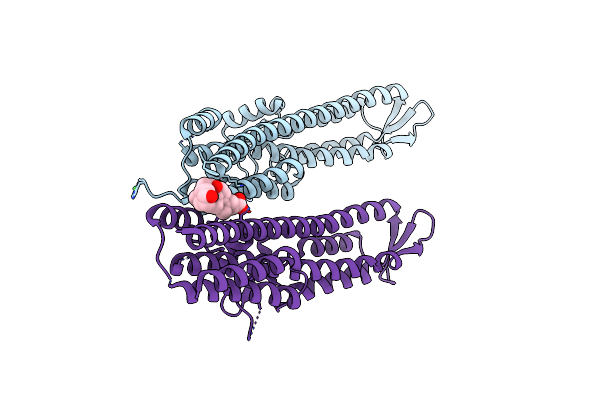
Crystal Structure Of The Salmonella Type Iii Secretion System Tip Protein Sipd
Organism: Salmonella enterica
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2010-11-17 Classification: CELL INVASION Ligands: NI |
 |
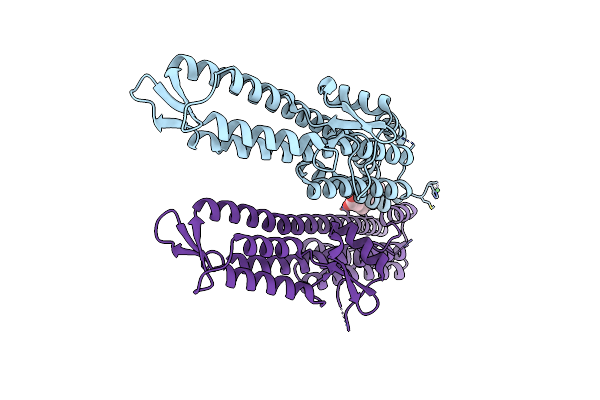
The Crystal Structure Of The Salmonella Type Iii Secretion System Tip Protein Sipd In Complex With Chenodeoxycholate
Organism: Salmonella enterica
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2010-11-17 Classification: CELL INVASION Ligands: NI, JN3 |
 |
The Crystal Structure Of The Salmonella Type Iii Secretion System Tip Protein Sipd In Complex With Deoxycholate
Organism: Salmonella enterica
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2010-11-17 Classification: CELL INVASION Ligands: NI, DXC |
 |
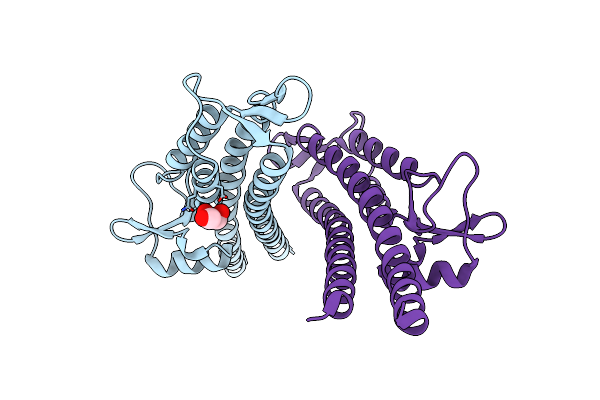
Crystal Structure Of The Salmonella Type Iii Secretion System Tip Protein Sipd-C244S
Organism: Salmonella enterica
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2010-11-17 Classification: CELL INVASION Ligands: NI |
 |
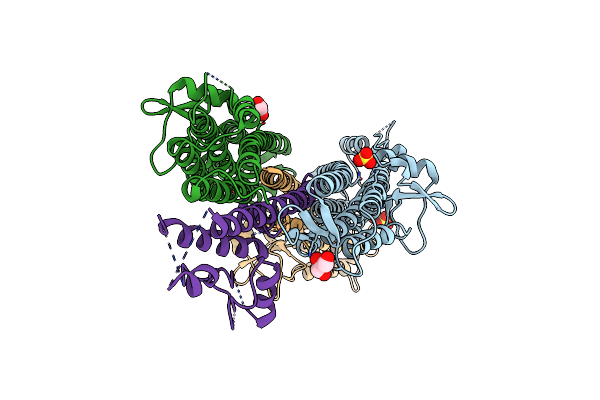
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2011-08-17 Classification: CELL INVASION Ligands: GOL |
 |
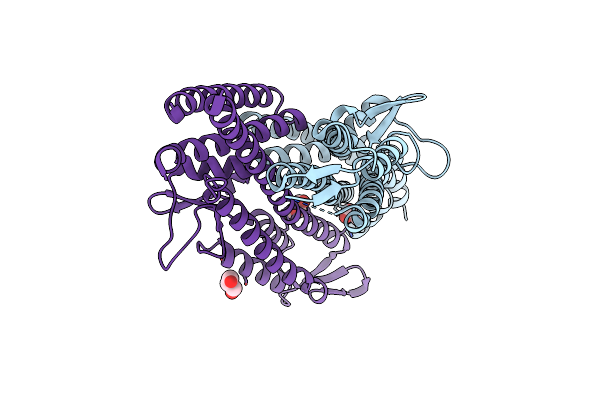
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2011-08-17 Classification: CELL INVASION Ligands: SO4, GOL |
 |
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2011-08-17 Classification: CELL INVASION Ligands: GOL |
 |
Organism: Homo sapiens, Mus musculus
Method: X-RAY DIFFRACTION Resolution:3.40 Å Release Date: 2023-06-07 Classification: CELL INVASION Ligands: SO4, GOL |
 |
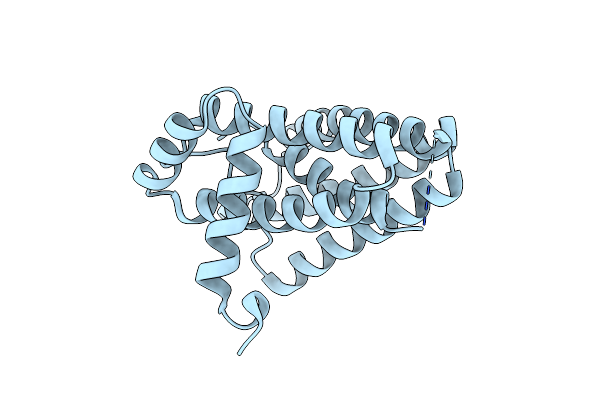
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.60 Å Release Date: 2024-01-24 Classification: CELL INVASION/INHIBITOR Ligands: NJ6 |
 |
Organism: Salmonella typhimurium
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2006-03-21 Classification: CELL INVASION |
 |
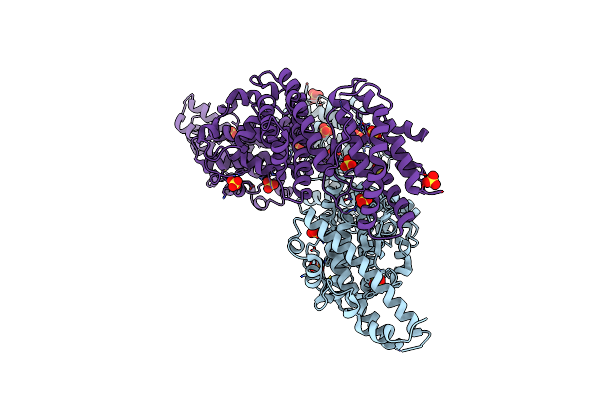
Organism: Plasmodium falciparum
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2005-08-09 Classification: CELL INVASION Ligands: SO4, CL |
 |
Crystal Structure Of Eba-175 Region Ii (Rii) Crystallized In The Presence Of (Alpha)2,3-Sialyllactose
Organism: Plasmodium falciparum
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2005-08-09 Classification: CELL INVASION Ligands: SO4, CL |

