Search Count: 118
All
Selected
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: Y01, XKP, C14, D12, D10, A1EOT, NAG |
 |
Unspecific Peroxygenase From Psathyrella Aberdarensis, Grogu Variant, In Complex With Tetradecane
Organism: Candolleomyces aberdarensis
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: NAG, MES, GOL, HEM, ZN, MG, C14 |
 |
Structure Of A Membrane-Bound Inositol Phosphorylceramide Synthase And Ceramide Complex
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-03-11 Classification: LIPID BINDING PROTEIN Ligands: UJO, C14, 46E |
 |
Structure Of A Membrane-Bound Inositol Phosphorylceramide Synthase And Aureobasidin A Complex
Organism: Saccharomyces cerevisiae s288c, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.53 Å Release Date: 2026-03-11 Classification: TRANSFERASE Ligands: C14, 46E |
 |
The Structure Of Tmd With 2 Tarps And 2 Cnihs From All Native Ampa Receptor Subtypes
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: MEMBRANE PROTEIN Ligands: POV, C14, XVD, D12 |
 |
The Structure Of Tmd With 3 Tarps And 1 Cnih From All Native Ampa Receptor Subtypes
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: MEMBRANE PROTEIN Ligands: POV, C14, XVD |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: MEMBRANE PROTEIN Ligands: POV, C14, OCT, XVD |
 |
Hemichannel Sub-Structure Of Cx43/Gja1 Gap Junction Intercellular Channel, Treated With A 5-Molar Excess Of Carbenoxolone
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: C14 |
 |
Hemichannel Sub-Structure Of Cx36/Gjd2 Gap Junction Intercellular Channel (Fn Conformation) In Brain Polar Lipid Nanodiscs, Treated With A 14-Fold Molar Excess Of Carbenoxolone
Organism: Homo sapiens, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: C14 |
 |
Hemichannel Sub-Structure Of Cx36/Gjd2 Gap Junction Intercellular Channel (Fn Conformation) In Soybean Polar Lipid Nanodiscs, Treated With A 20-Fold Molar Excess Of Carbenoxolone
Organism: Homo sapiens, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: MC3, C14 |
 |
Hemichannel Sub-Structure Of Cx36/Gjd2 Gap Junction Intercellular Channel (Fn Conformation) In Soybean Polar Lipid Nanodiscs, Treated With A 10-Fold Molar Excess Of Carbenoxolone And Incubated Shortly
Organism: Homo sapiens, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: MC3, C14 |
 |
Hemichannel Sub-Structure Of Cx36/Gjd2 Gap Junction Intercellular Channel (Fn Conformation) In Soybean Polar Lipid Nanodiscs, Treated With A 10-Fold Molar Excess Of Carbenoxolone
Organism: Homo sapiens, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: MC3, C14 |
 |
Structure Of The Double Cys-Substituted Cross-Linked Acrb Variant S562C_T837C
Organism: Escherichia coli k-12, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-11-19 Classification: TRANSPORT PROTEIN Ligands: LMT, D10, GOL, D12, HEX, C14, OCT, DD9, SO4 |
 |
Organism: Escherichia coli k-12, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2025-11-12 Classification: TRANSPORT PROTEIN Ligands: LMT, OCT, C14, EDO, GOL, D12, HEX, XE9, D10, DDR, DDQ, SO4, LPX |
 |
Cryo-Em Structure Of The Myxol-Bound Light-Driven Chloride Ion-Pumping Rhodopsin, Nm-R3
Organism: Nonlabens marinus s1-08
Method: ELECTRON MICROSCOPY Resolution:2.48 Å Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN Ligands: RET, A1L4O, CL, PC1, PLC, R16, 8K6, D12, C14, D10 |
 |
Cryo-Em Structure Of The Light-Driven Chloride Ion-Pumping Rhodopsin, Nm-R3
Organism: Nonlabens marinus s1-08
Method: ELECTRON MICROSCOPY Resolution:2.50 Å Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN Ligands: RET, CL, PC1, PLC, D12, R16, 8K6, C14 |
 |
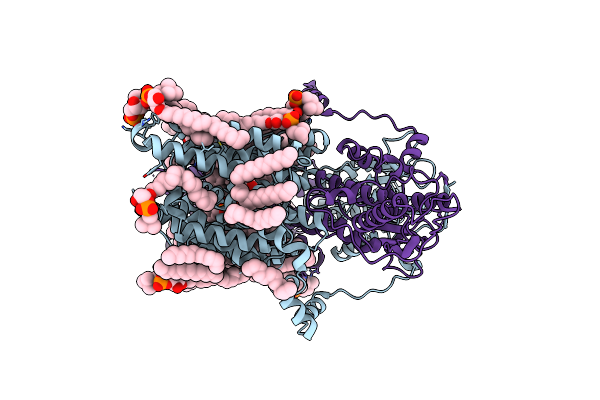
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-16 Classification: MEMBRANE PROTEIN Ligands: TAU, D12, OCT, C14, DD9, HP6, D10, HEX, CL, NA, NAG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: TRANSPORT PROTEIN Ligands: OXL, POV, C14, D12, LPE, D10, CLR |
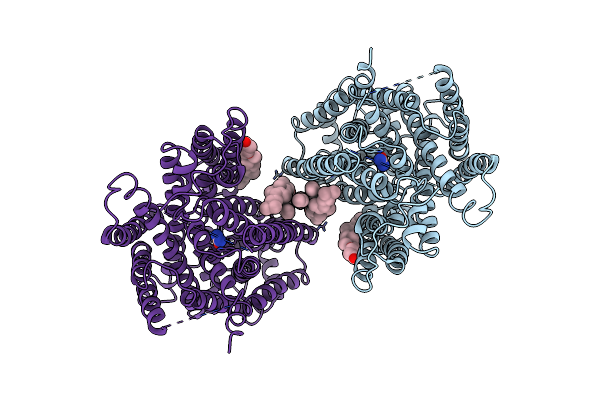 |
Arabidopsis High-Affinity Urea Transport Dur3 In The Urea-Bound Occluded Conformation, Dimeric State
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2025-05-07 Classification: MEMBRANE PROTEIN Ligands: Y01, URE, R16, C14 |
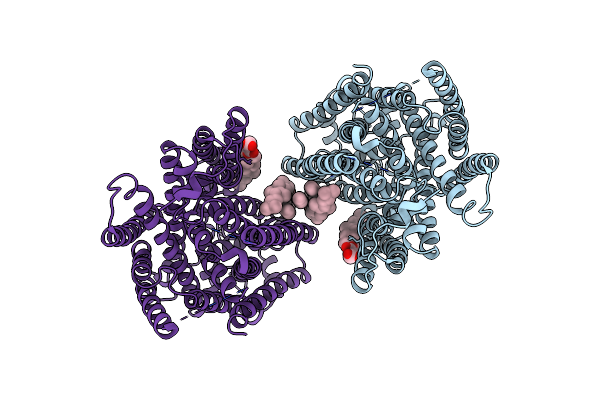 |
Arabidopsis High-Affinity Urea Transport Dur3 In The Inward-Facing Open Conformation, Dimeric State
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2025-05-07 Classification: MEMBRANE PROTEIN Ligands: Y01, C14, R16 |

