Search Count: 74
All
Selected
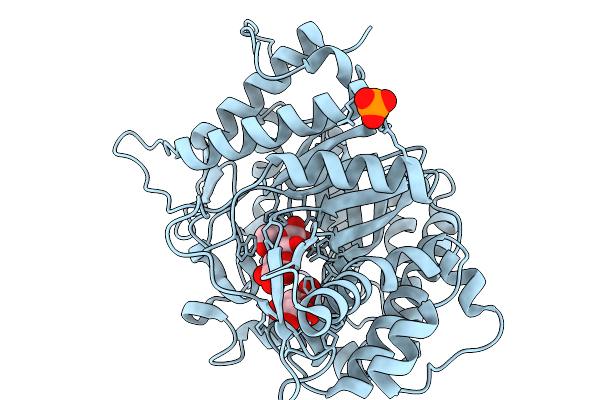 |
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: GLC, BGC, PO4 |
 |
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
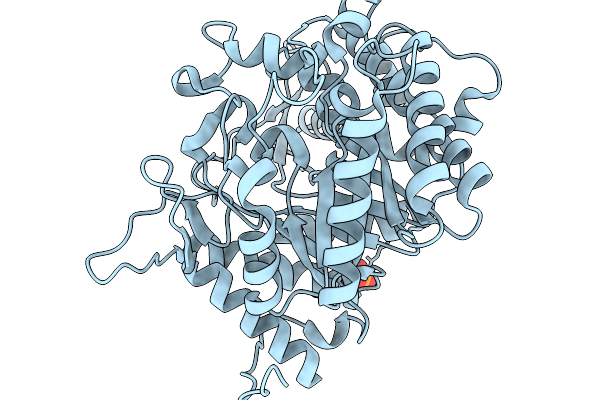 |
Crystal Structure Of Glucose Tolerant Beta-Glucosidase Unbgl1 Mutant (C188V)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: CL, PO4 |
 |
Crystal Structure Of Glucsoe Bound Glucose Tolerant Gh1 Beta-Glucosidase Mutant (Unbgl1_C188V)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
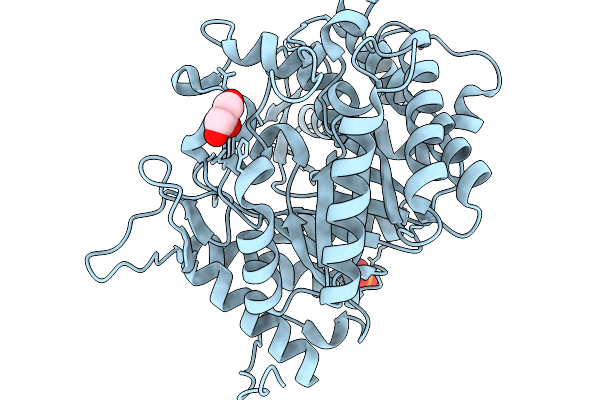 |
Crystal Structure Of Glucose Tolerant Gh1 Beta-Glucosidase Mutant (Unbgl1_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: PO4, GOL, CL |
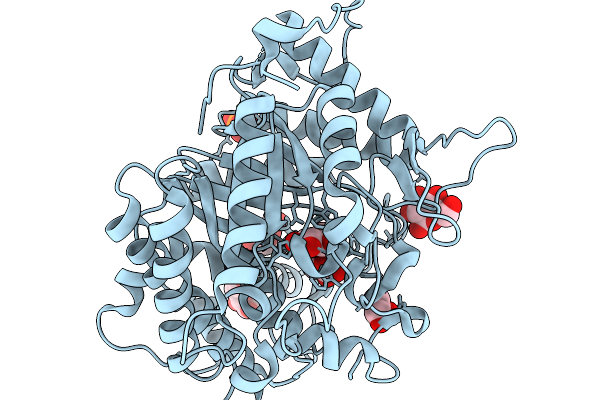 |
Crystal Structure Of Glucose Bound Gh1 Beta-Glucosidase Mutant (Unbgl1_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, GOL, PO4 |
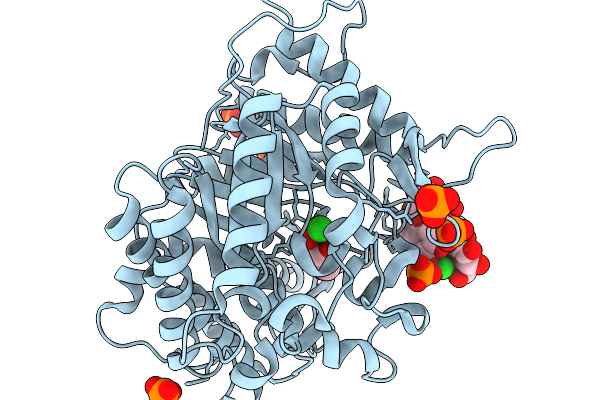 |
Crystal Structure Of Cellobiose And Glucose Bound Glucose Toleranant Beta-Glucosidase Mutant (Unbgl1_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4, CL |
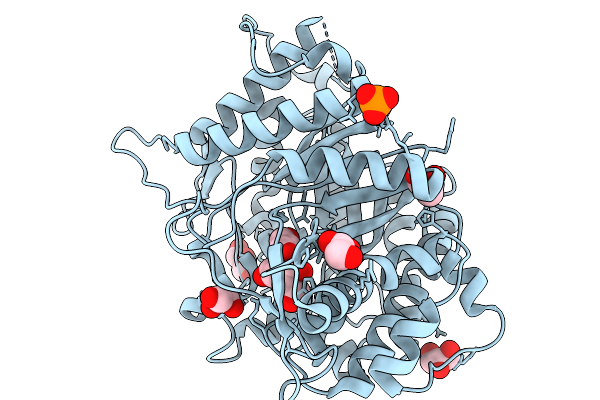 |
High Resolution Crystal Strucutre Of Highly Glucose Tolerant Gh1 Beta-Glucosidase (Unbgl1_C188V_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: PO4, GOL |
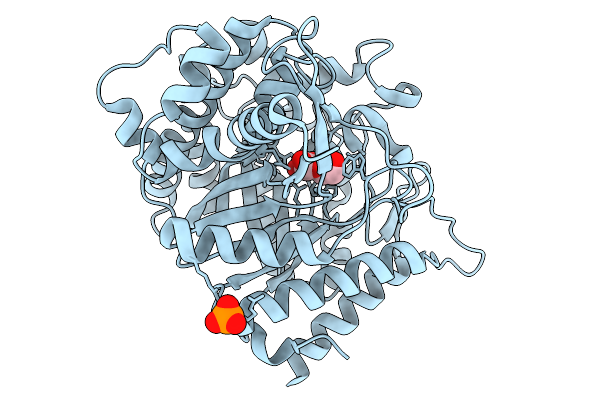 |
Crystal Structure Of Glucose Bound Highly Glucose Tolerant Gh1 Beta-Glcosidase Mutant (Unbgl1_C188V_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
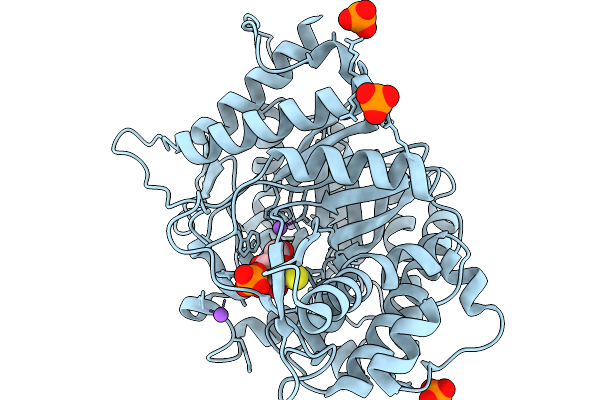 |
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: GS1, PO4, NA |
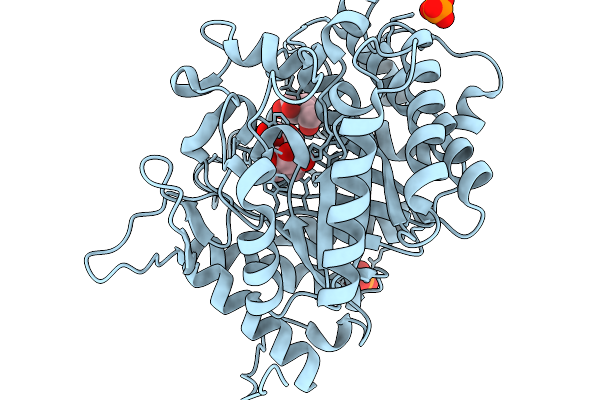 |
Crystal Structure Of Glucose Bound Covalent Intermediate Of Gh1 Beta-Glucosidase (Unbgl1)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
 |
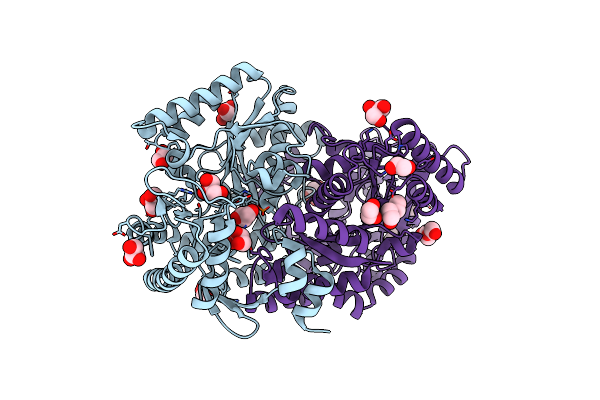
High Resolution Crystal Structure Of Gh1 Beta-Glucosidase From Soil Metagenome (Unbgl1)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: PO4, CL, GOL |
 |
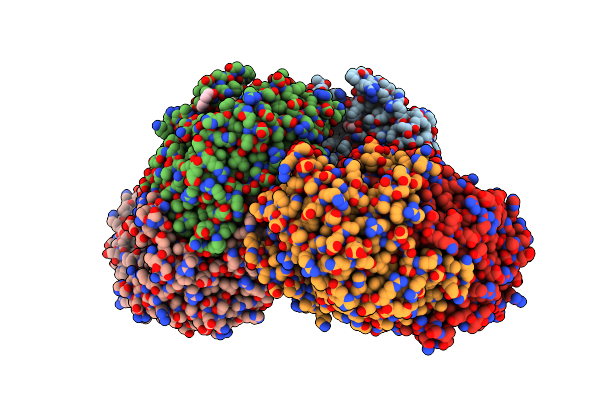
Crystal Structure Of Omega Transaminase Ta_2799 From Pseudomonas Putida Kt2440
Organism: Pseudomonas putida kt2440
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2025-06-04 Classification: TRANSFERASE Ligands: GOL, EDO, PEG |
 |
Crystal Structure Of The L322F Mutant Of Omega Transaminase Ta_2799 From Pseudomonas Putida Kt2440
Organism: Pseudomonas putida kt2440
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-06-04 Classification: TRANSFERASE Ligands: PEG, EDO, SO4, GOL |
 |
Crystal Structure Of The Open State Of Omega Transaminase Ta_5182 From Pseudomonas Putida Kt2440
Organism: Pseudomonas putida kt2440
Method: X-RAY DIFFRACTION Resolution:3.40 Å Release Date: 2025-06-04 Classification: TRANSFERASE Ligands: SO4 |
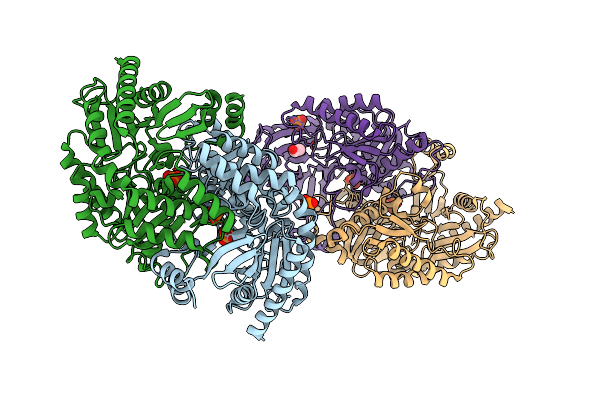 |
Crystal Structure Of The Closed State Of The Omega Transaminase Ta_5182 From Pseudomonas Putida Kt2440
Organism: Pseudomonas putida kt2440
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-06-04 Classification: TRANSFERASE Ligands: PO4, EDO |
 |
Cryo-Em Structure Of Apo Aspergillus Terreus Glutamate Dehydrogenase (Atgdh) In The Closed Conformation (Form 1)
Organism: Aspergillus terreus
Method: ELECTRON MICROSCOPY Resolution:3.15 Å Release Date: 2025-03-05 Classification: OXIDOREDUCTASE |
 |
Cryo-Em Structure Of Apo Aspergillus Terreus Glutamate Dehydrogenase (Atgdh) In The Partially Closed Conformation (Form 2)
Organism: Aspergillus terreus
Method: ELECTRON MICROSCOPY Resolution:3.25 Å Release Date: 2025-03-05 Classification: OXIDOREDUCTASE |
 |
Cryo-Em Structure Of Apo Aspergillus Terreus Glutamate Dehydrogenase (Atgdh) In The Closed Conformation (Form 2)
Organism: Aspergillus terreus
Method: ELECTRON MICROSCOPY Resolution:3.18 Å Release Date: 2025-03-05 Classification: OXIDOREDUCTASE |
 |
Cryo-Em Structure Of Apo Aspergillus Niger Glutamate Dehydrogenase (Angdh) In The Closed Conformation
Organism: Aspergillus niger
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: OXIDOREDUCTASE |

