Search Count: 284
All
Selected
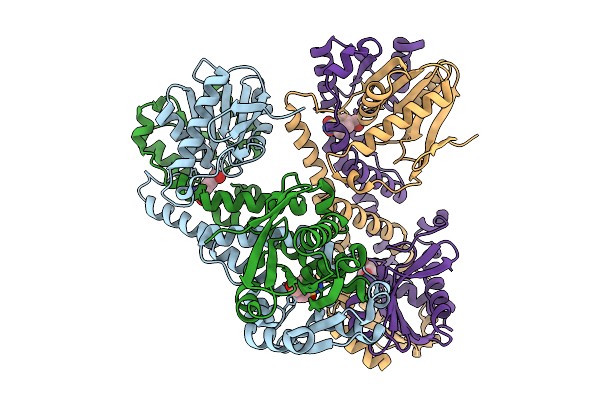 |
Fphe, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases E, Borolane-Based Compound Q41 Bound In Crystal Form 6
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: XPU |
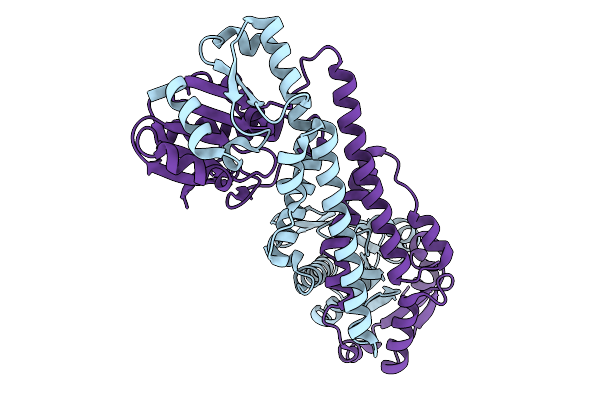 |
Fphe, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases E, Unbound Dimer Crystal Form 9
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-01-21 Classification: HYDROLASE |
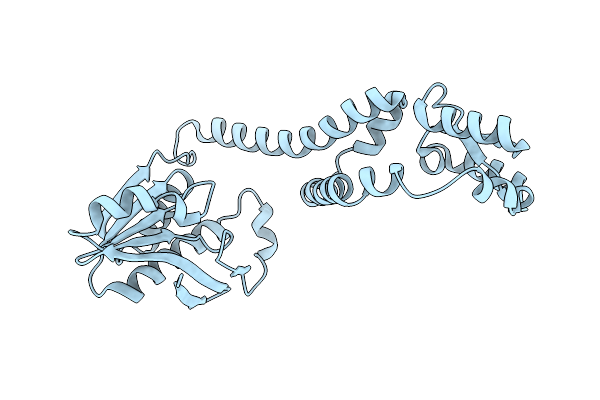 |
Fphe, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases E, Unbound Dimer Crystal Form 8
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-08-27 Classification: HYDROLASE |
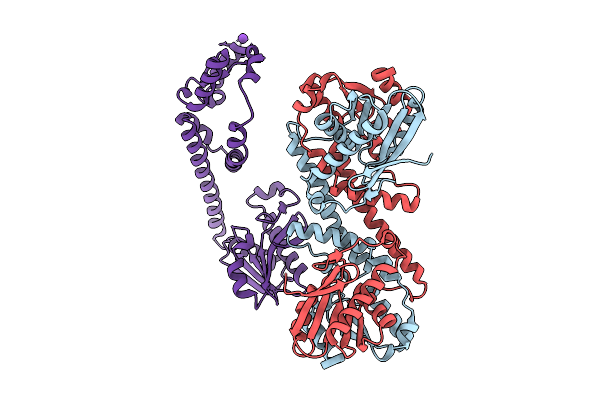 |
Fphe, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases E, Unbound Dimer Crystal Form 5
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2025-08-06 Classification: HYDROLASE Ligands: K |
 |
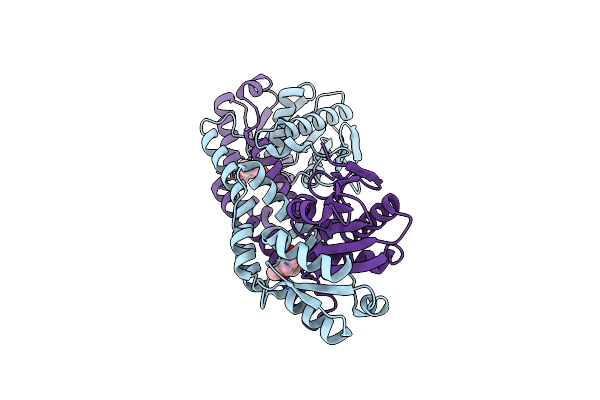
Fphe, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases E, Borolane-Based Compound Q41 Bound
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2024-11-20 Classification: HYDROLASE Ligands: XPU, CA |
 |
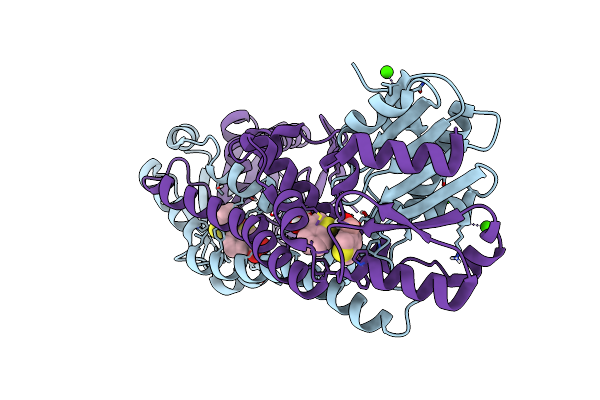
Fphe, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases E, Boronic Acid-Based Compound Y43 Bound
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2024-10-23 Classification: HYDROLASE Ligands: WS8 |
 |
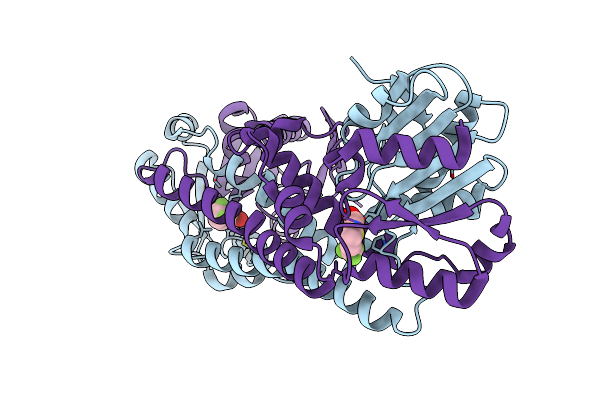
Fphe, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases E, Boronic Acid-Based Compound Z27 Bound
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2024-10-16 Classification: HYDROLASE Ligands: WKK, CA |
 |
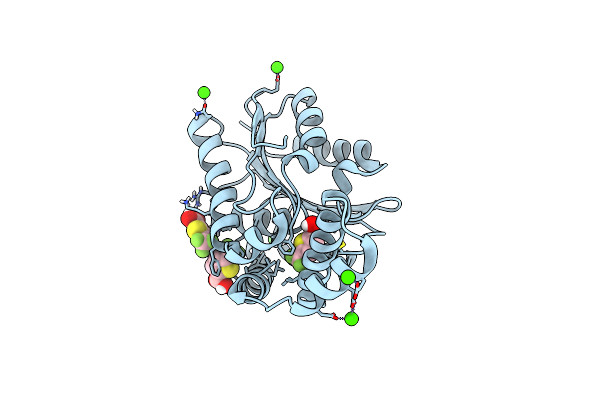
Fphe, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases E, Boronic Acid-Based Compound N34 Bound
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2024-07-24 Classification: HYDROLASE Ligands: ZKR, MG |
 |
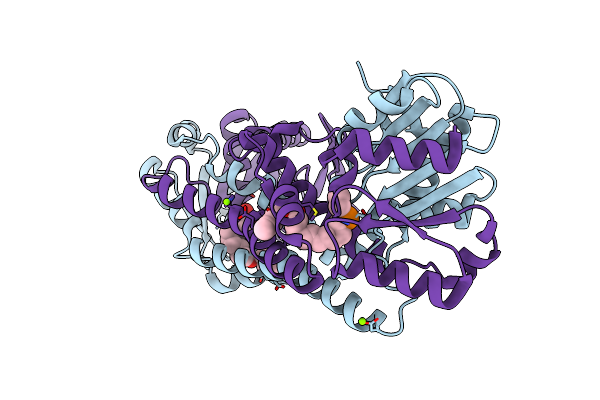
Fphh, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases H, Boronic Acid-Based Compound N34 Bound
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2024-07-17 Classification: HYDROLASE Ligands: ZKR, CA |
 |
Fphe, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases E, Fluorophosphonate Jb101 Bound, Dimer Crystal Form 1
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2024-04-17 Classification: HYDROLASE Ligands: ZUV, MG |
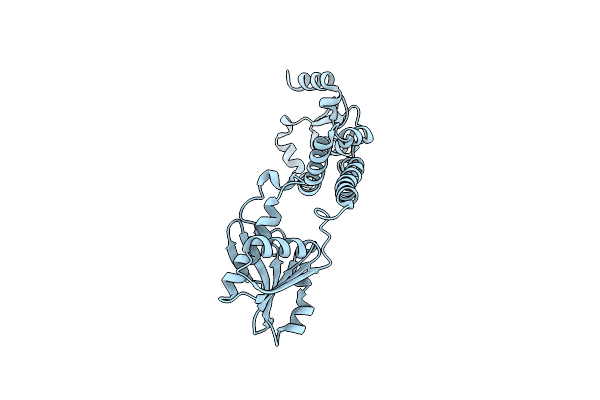 |
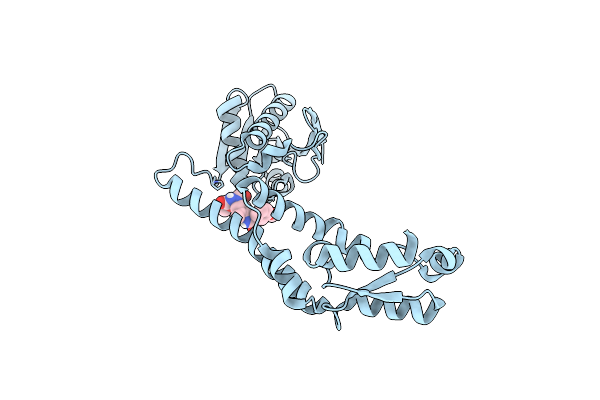
Fphe, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases E, Unbound, Dimer Crystal Form 3
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2024-02-21 Classification: HYDROLASE |
 |
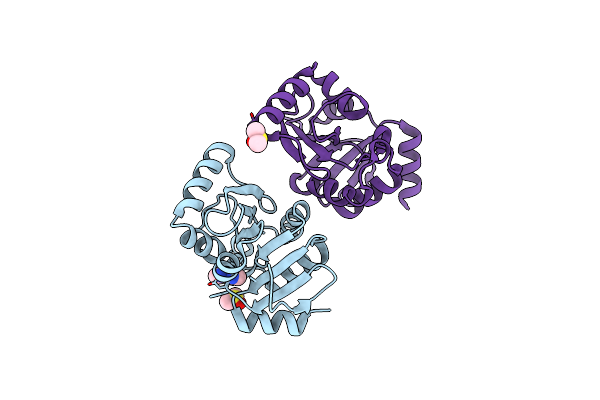
Fphe, Staphylococcus Aureus Fluorophosphonate-Binding Serine Hydrolases E, Oxadiazolone Compound 3 Bound, Dimer Crystal Form 2
Organism: Staphylococcus aureus subsp. aureus usa300
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2024-02-21 Classification: HYDROLASE Ligands: YKF |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With En300-321461
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.13 Å Release Date: 2021-01-13 Classification: VIRAL PROTEIN Ligands: WOY, DMS |
 |
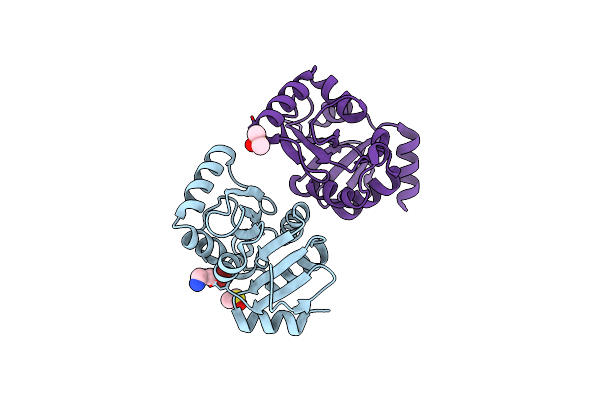
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With En300-43406
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.08 Å Release Date: 2021-01-13 Classification: VIRAL PROTEIN Ligands: WPS |
 |
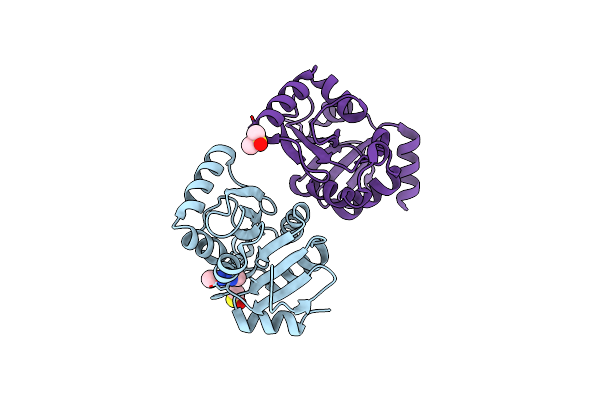
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With Z3034471507
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.17 Å Release Date: 2021-01-13 Classification: VIRAL PROTEIN Ligands: WPV, DMS |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With Ab-601_30915014
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.17 Å Release Date: 2021-01-13 Classification: VIRAL PROTEIN Ligands: WPY, DMS |
 |
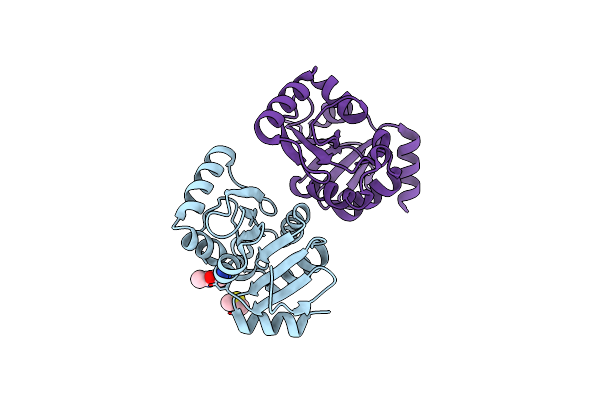
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With En300-108952
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.11 Å Release Date: 2021-01-13 Classification: VIRAL PROTEIN Ligands: WQ1 |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With En300-301084
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.07 Å Release Date: 2021-01-13 Classification: VIRAL PROTEIN Ligands: WQ4, DMS |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With En300-105873
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.08 Å Release Date: 2021-01-13 Classification: VIRAL PROTEIN Ligands: WQ7 |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With Stk497968
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.18 Å Release Date: 2021-01-13 Classification: VIRAL PROTEIN Ligands: DE5 |

