Search Count: 232
All
Selected
 |
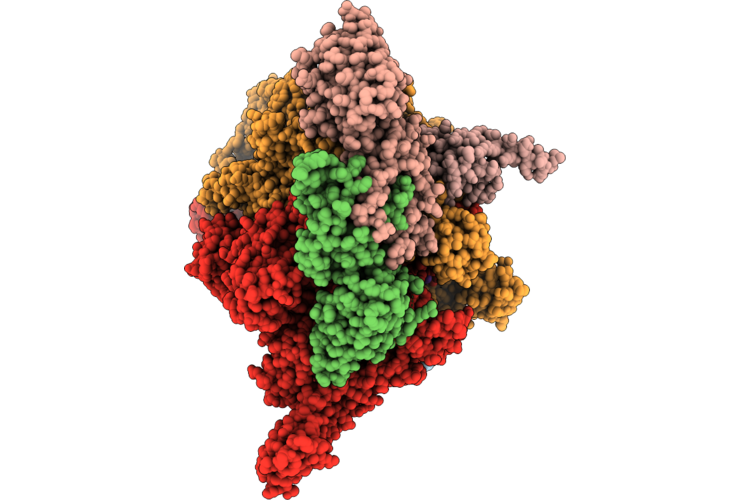
Closed Eco-Epec: Cryo-Em Structure Of Eco Rnap His-Elemental Paused Elongation Complex With A Closed Active Site (Closed Tl, Si3 And Rh-Fl)
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-04-29 Classification: Transcription/DNA/RNA Ligands: 1N7, MG, ZN |
 |
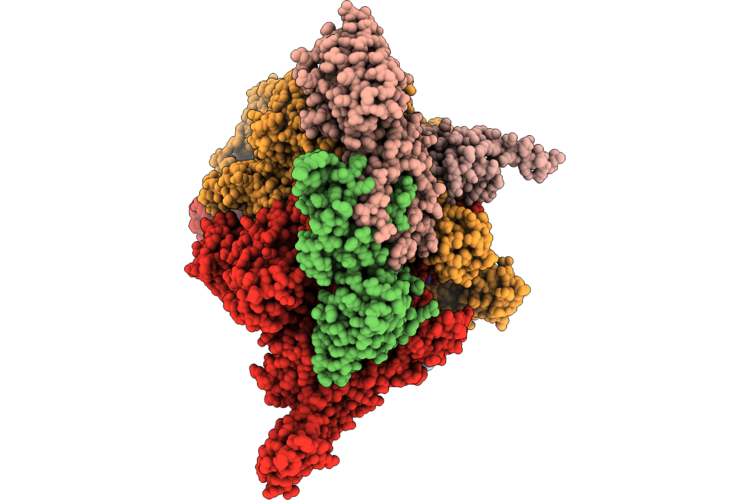
Open1 Eco-Epec: Cryo-Em Structure Of Eco Rnap His-Elemental Paused Elongation Complex With An Open Active Site (Open Tl, Si3 And Rh-Fl)
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-04-29 Classification: TRANSCRIPTION/DNA/RNA Ligands: 1N7, MG, ZN |
 |
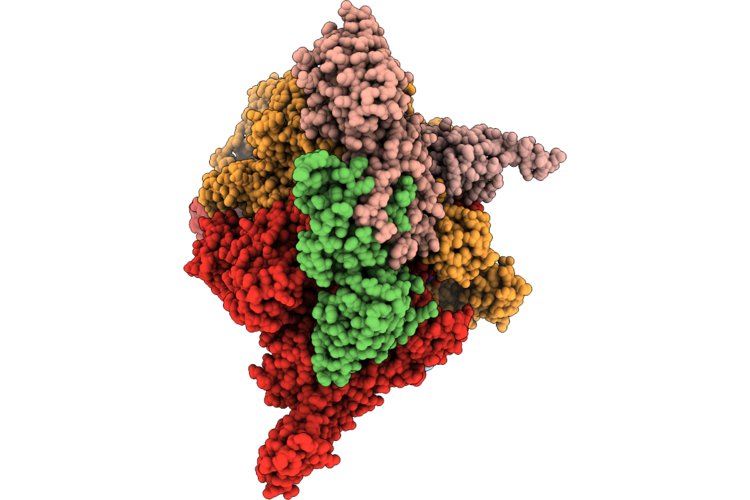
Open2 Eco-Epec: Cryo-Em Structure Of Eco Rnap His-Elemental Paused Elongation Complex With An Open Active Site (Open Tl, Si3 And Rh-Fl)
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-04-29 Classification: Transcription/DNA/RNA Ligands: 1N7, MG, ZN |
 |
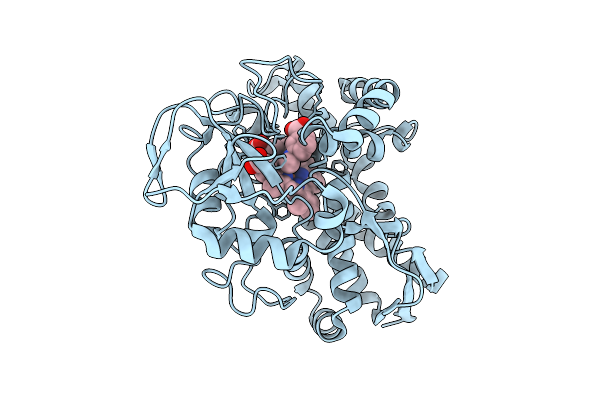
Open3 Eco-Epec: Cryo-Em Structure Of Eco Rnap His-Elemental Paused Elongation Complex With An Open Active Site (Open Tl, Si3 And Rh-Fl)
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-04-29 Classification: TRANSCRIPTION Ligands: 1N7, MG, ZN |
 |
The 100-K Crystal Structure Of Cyp199A4 Bound To 3-Methylaminobenzoic Acid (Dataset 1, Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: OW4, HEM, CL |
 |
The 150-K Crystal Structure Of Cyp199A4 Bound To 3-Methylaminobenzoic Acid (Dataset 2, Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: OW4, HEM, CL |
 |
The 200-K Crystal Structure Of Cyp199A4 Bound To 3-Methylaminobenzoic Acid (Dataset 3, Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: OW4, HEM, CL |
 |
The 150-K Crystal Structure Of Cyp199A4 Bound To 4-Methoxybenzoic Acid (Dataset 2, Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: ANN, HEM, CL |
 |
The 200-K Crystal Structure Of Cyp199A4 Bound To 4-Methoxybenzoic Acid (Dataset 3, Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: ANN, HEM, CL |
 |
The 300-K Crystal Structure Of Cyp199A4 Bound To 4-Methoxybenzoic Acid (Dataset 4, Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: HEM, ANN |
 |
The 200-K Crystal Structure Of Cyp199A4 Bound To 3-Methylaminobenzoic Acid (Dataset 1; Decreasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: OW4, HEM, CL |
 |
The 150-K Crystal Structure Of Cyp199A4 Bound To 3-Methylaminobenzoic Acid (Dataset 2; Decreasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: OW4, HEM, CL |
 |
The 100-K Crystal Structure Of Cyp199A4 Bound To 3-Methylaminobenzoic Acid (Dataset 3; Decreasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: OW4, HEM, CL |
 |
The 100-K Crystal Structure Of Cyp199A4 Bound To 4-Phenoxybenzoic Acid (Dataset 1; Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.53 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: HEM, JO4, CL |
 |
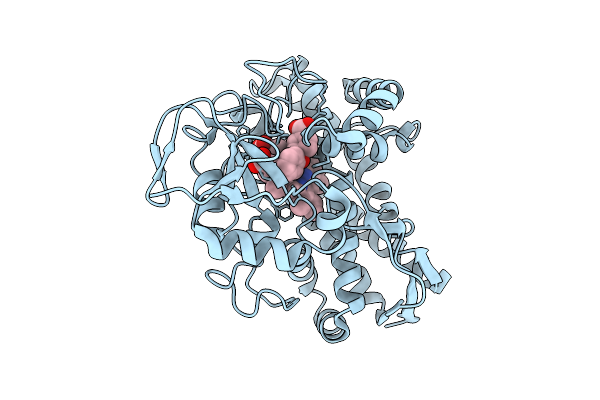
The 150-K Crystal Structure Of Cyp199A4 Bound To 4-Phenoxybenzoic Acid (Dataset 2; Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: JO4, HEM, CL |
 |
The 200-K Crystal Structure Of Cyp199A4 Bound To 4-Phenoxybenzoic Acid (Dataset 3; Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: JO4, HEM, CL |
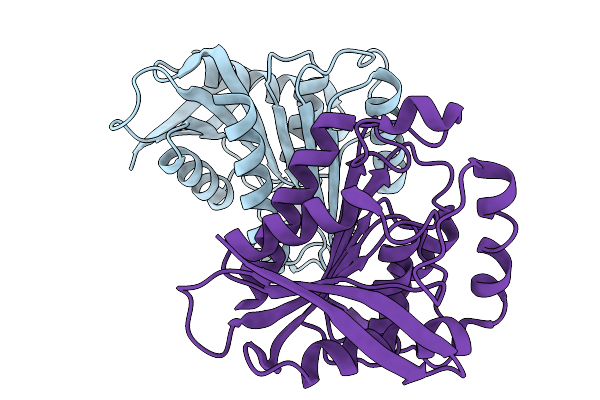 |
Organism: Solimonas aquatica
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-02-18 Classification: HYDROLASE |
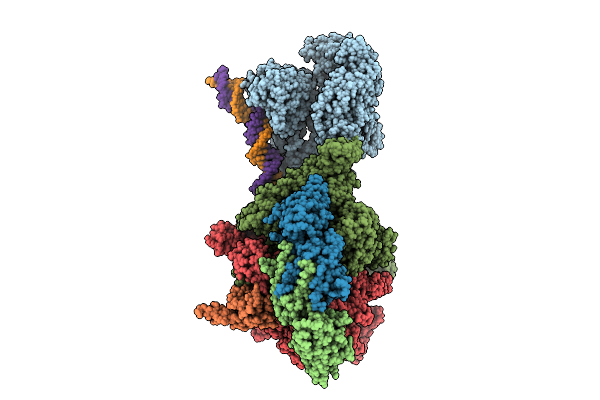 |
Organism: Escherichia coli, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-10-15 Classification: TRANSCRIPTION Ligands: ADP, BEF, MG, ZN |
 |
Organism: Escherichia coli, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-10-15 Classification: TRANSCRIPTION Ligands: MG, ZN |
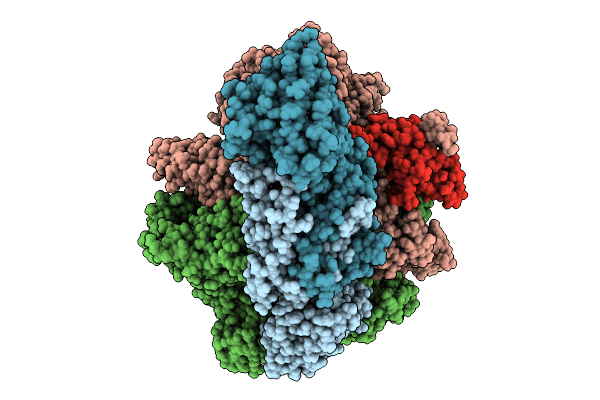 |
Mycobacterium Tuberculosis Rna Polymerase Elongation Complex Bound To Inhibitor N-Aroyl-N-Aryl-Phenylalanine Amide (Aap)-So2
Organism: Mycobacterium tuberculosis h37rv, Mycobacterium tuberculosis
Method: ELECTRON MICROSCOPY Resolution:2.97 Å Release Date: 2025-10-08 Classification: TRANSCRIPTION, TRANSFERASE/INHIBITOR Ligands: A1BNV, ZN, MG |

