Search Count: 232
All
Selected
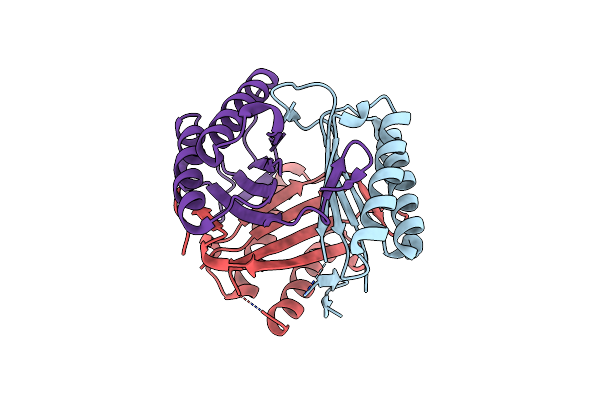 |
Crystal Structure Of Macrophage Migration Inhibitory Factor (Mif) From Trichomonas Vaginalis (I4122 Form)
Organism: Trichomonas vaginalis g3
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2024-03-20 Classification: CYTOKINE Ligands: CL |
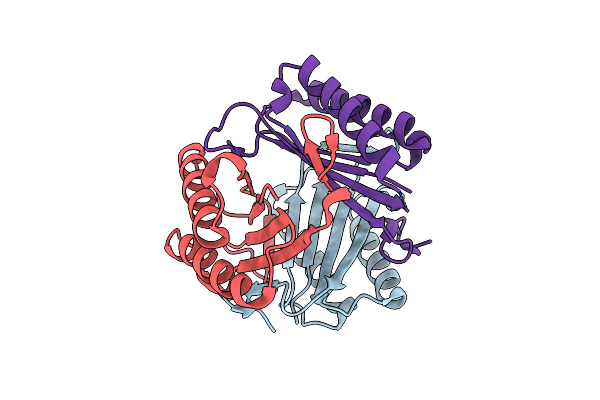 |
Crystal Structure Of Macrophage Migration Inhibitory Factor (Mif) From Trichomonas Vaginalis (Apo, P41212 Form)
Organism: Trichomonas vaginalis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2023-12-06 Classification: CYTOKINE |
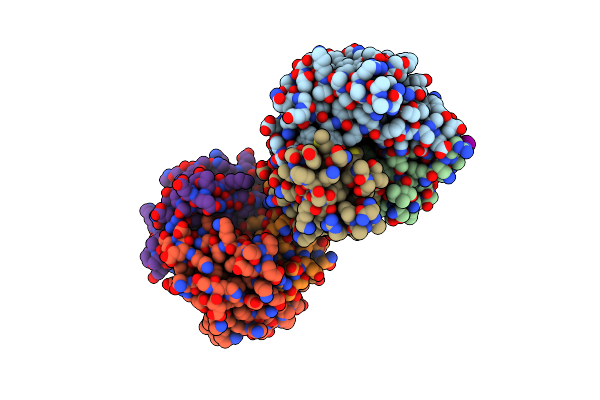 |
Crystal Structure Of Macrophage Migration Inhibitory Factor (Mif) From Trichomonas Vaginalis (I41 Form)
Organism: Trichomonas vaginalis g3
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2023-11-08 Classification: CYTOKINE Ligands: IOD, PYR |
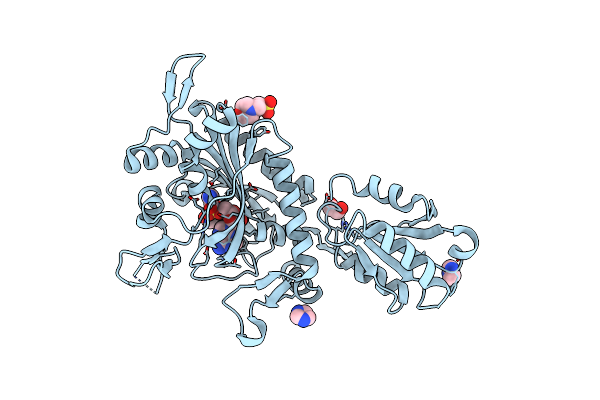 |
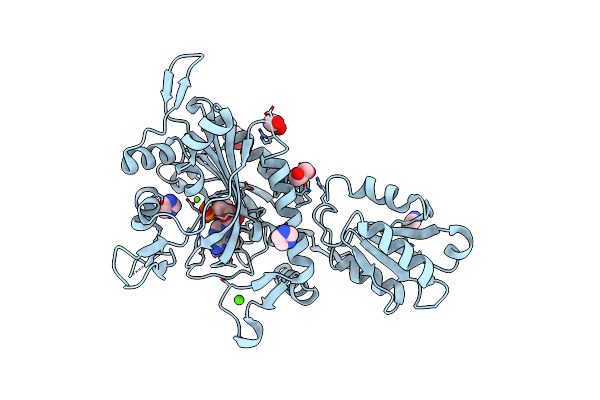
Crystal Structure Of Glycine--Trna Ligase From Mycobacterium Tuberculosis (G5A Bound)
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2023-09-13 Classification: TRANSFERASE Ligands: G5A, IMD, MES, EDO, MG, CL |
 |
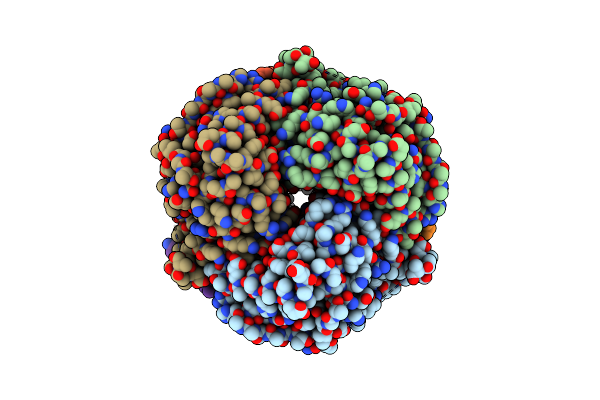
Crystal Structure Of Glycine--Trna Ligase From Mycobacterium Tuberculosis (Amp-Mg Bound)
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2023-06-28 Classification: LIGASE Ligands: MG, CA, EDO, GOL, IMD, AMP |
 |
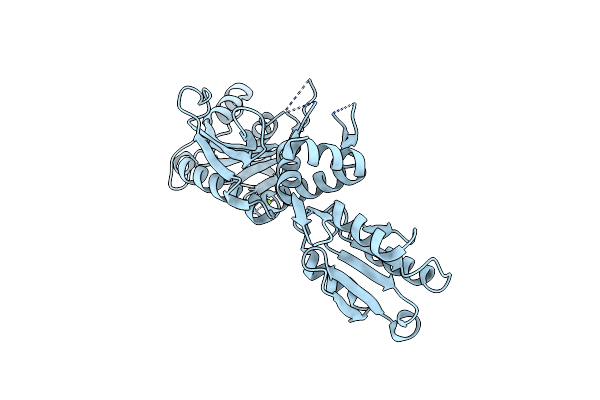
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2023-05-10 Classification: HYDROLASE |
 |
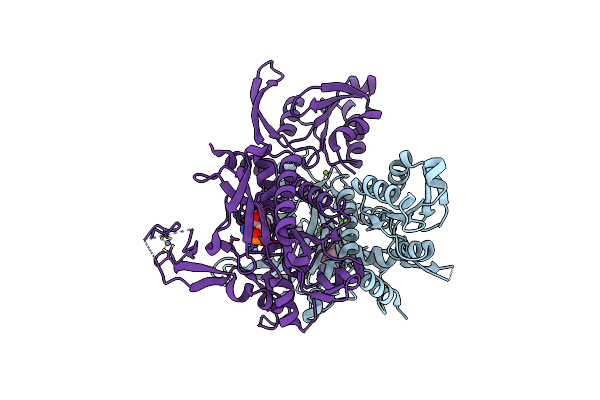
Crystal Structure Of Glycine Trna Ligase From Mycobacterium Thermoresistibile (Apo)
Organism: Mycolicibacterium thermoresistibile atcc 19527
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2023-05-03 Classification: LIGASE Ligands: MG |
 |
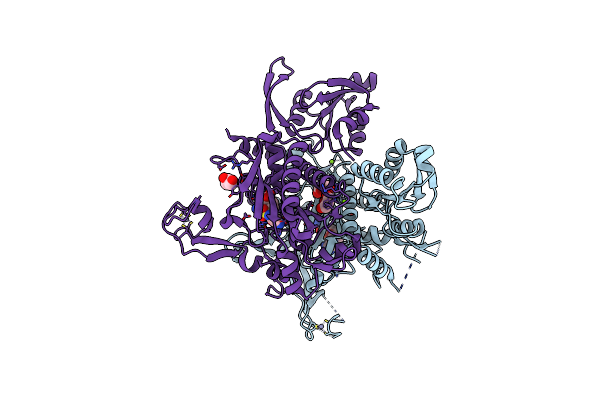
Crystal Structure Of Glycine Trna Ligase From Mycobacterium Thermoresistibile (Amp Bound)
Organism: Mycolicibacterium thermoresistibile atcc 19527
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2023-05-03 Classification: LIGASE Ligands: AMP, MG, ZN |
 |
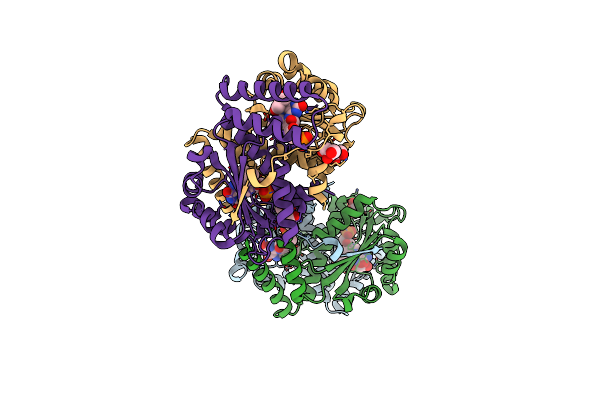
Crystal Structure Of Glycine Trna Ligase From Mycobacterium Thermoresistibile (Glycyl Adenylate Bound)
Organism: Mycolicibacterium thermoresistibile atcc 19527
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2023-05-03 Classification: LIGASE Ligands: G5A, ZN, MG, GOL |
 |
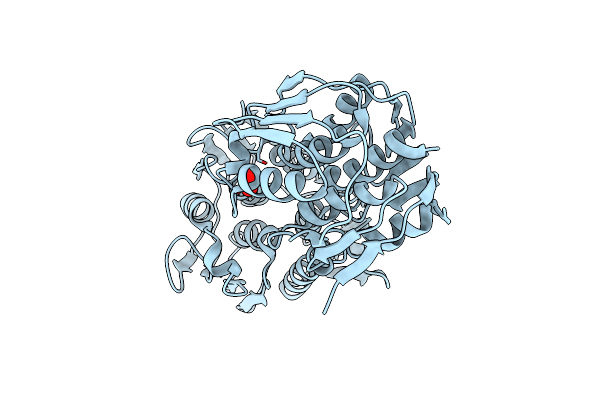
Crystal Structure Of Dihydropteridine Reductase/Oxygen-Insensitive Nad(P)H Nitroreductase From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2022-07-27 Classification: OXIDOREDUCTASE Ligands: FMN, PO4, GOL |
 |
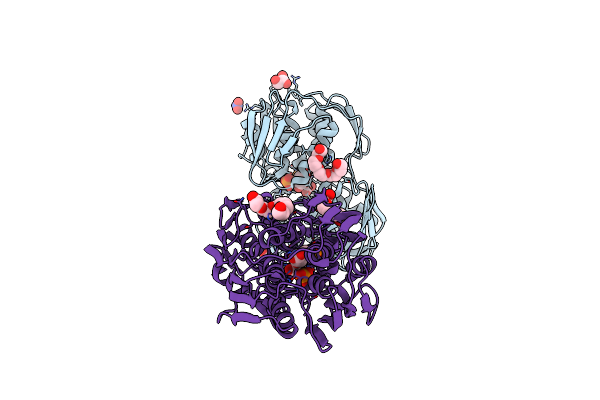
Crystal Structure Of Apo 3-Phosphoshikimate 1-Carboxyvinyltransferase From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae subsp. pneumoniae hs11286
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2022-02-02 Classification: TRANSFERASE Ligands: NO3 |
 |
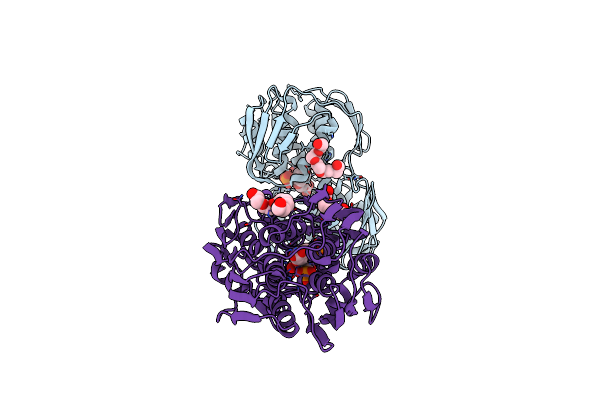
Crystal Structure Of Shikimate-3-Phosphate Bound 3-Phosphoshikimate 1-Carboxyvinyltransferase From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae subsp. pneumoniae hs11286
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2022-02-02 Classification: TRANSFERASE Ligands: S3P, FMT, PO4, 1PE, GOL, NO3 |
 |
Crystal Structure Of Shikimate-3-Phosphate And Glyphosate Bound 3-Phosphoshikimate 1-Carboxyvinyltransferase From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae subsp. pneumoniae hs11286
Method: X-RAY DIFFRACTION Resolution:1.26 Å Release Date: 2022-02-02 Classification: TRANSFERASE Ligands: S3P, GPJ, 1PE |
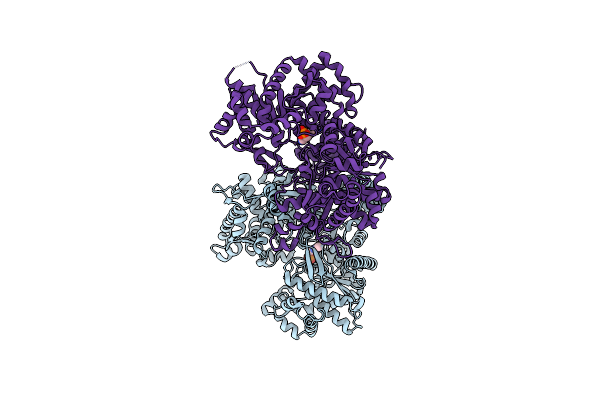 |
Crystal Structure Of Alpha-1,4 Glucan Phosphorylase From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae subsp. pneumoniae hs11286
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2022-02-02 Classification: TRANSFERASE Ligands: PLP |
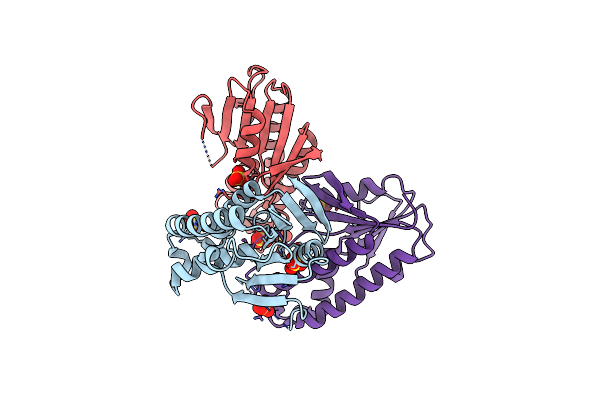 |
Crystal Structure Of Bifunctional Adenosylcobalamin Biosynthesis Protein From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae subsp. pneumoniae hs11286
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2022-02-02 Classification: TRANSFERASE Ligands: SO4 |
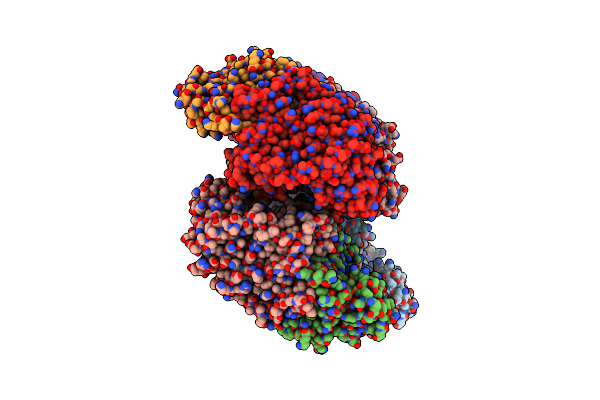 |
Crystal Structure Of Bacterial Alkaline Phosphatase From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae subsp. pneumoniae hs11286
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2022-02-02 Classification: HYDROLASE Ligands: ZN, MG |
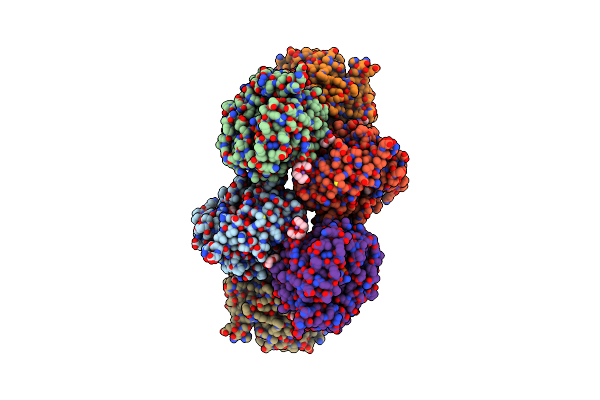 |
Organism: Klebsiella pneumoniae subsp. pneumoniae hs11286
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2022-02-02 Classification: OXIDOREDUCTASE |
 |
Crystal Structure Of Ornithine Carbamoyltransferase From Klebsiella Pneumoniae
Organism: Klebsiella pneumoniae subsp. pneumoniae hs11286
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2022-02-02 Classification: TRANSFERASE Ligands: IOD, BR |
 |
Crystal Structure Of Nad(P)H Nitroreductase From Klebsiella Pneumoniae (Short B-Axis)
Organism: Klebsiella pneumoniae subsp. pneumoniae hs11286
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2022-02-02 Classification: OXIDOREDUCTASE Ligands: FMN, BTB, GOL, PO4, 1PE |
 |
Crystal Structure Of Nad(P)H Nitroreductase From Klebsiella Pneumoniae (Long B-Axis)
Organism: Klebsiella pneumoniae subsp. pneumoniae hs11286
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2022-02-02 Classification: OXIDOREDUCTASE Ligands: FMN, PO4 |

