Search Count: 53
All
Selected
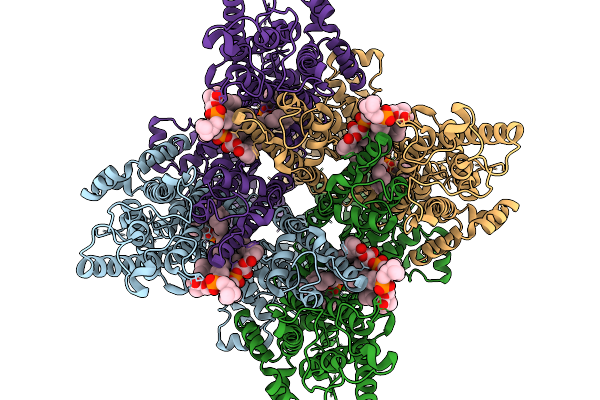 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: CPL |
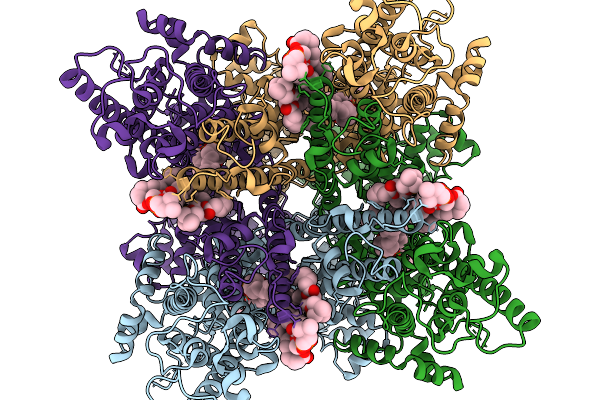 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: CPL |
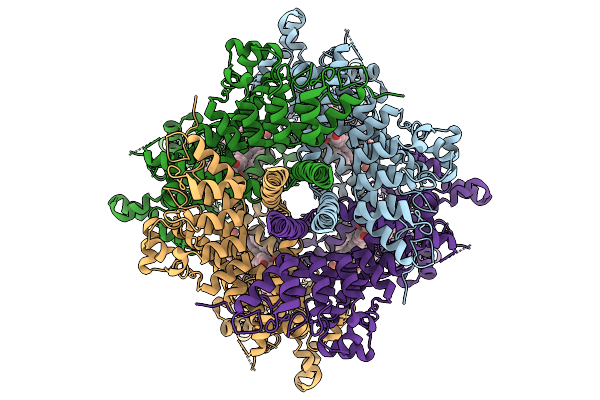 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: UNL |
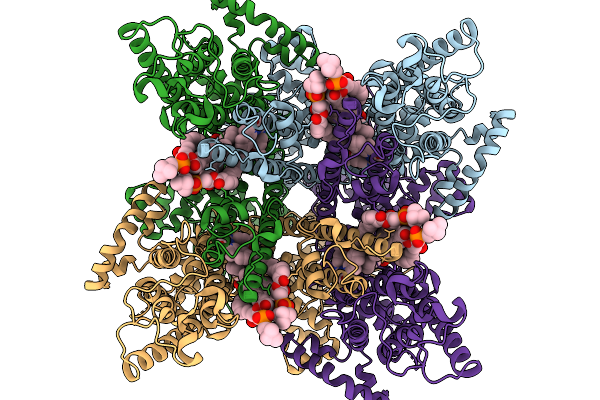 |
Structure Of A Constitutively Open Human Trpc3 Mutant In The Inhibited State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: UNL, A1CCX |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: UNL |
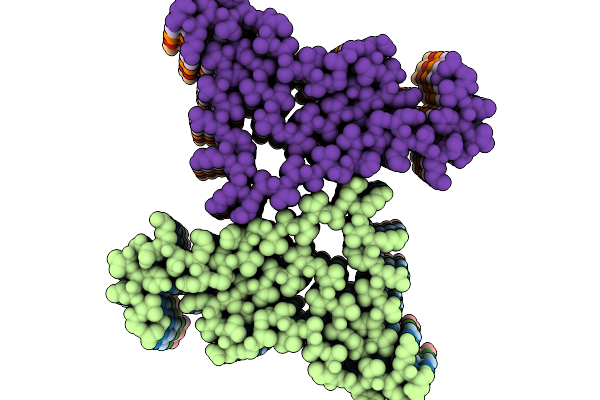 |
Structure Of Amplified Asyn Filament By Using Seed Amplification Assay (Saa) From Msa Patient Csf.
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.18 Å Release Date: 2026-02-25 Classification: PROTEIN FIBRIL |
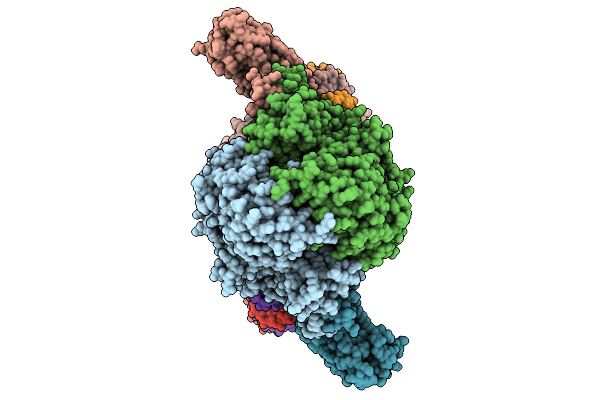 |
Cryoem Structure Of M. Mazei Topoisomerase Vi(A-E342Q)-Minicircle Dna Complex In Cleavage State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: CA, ANP, K |
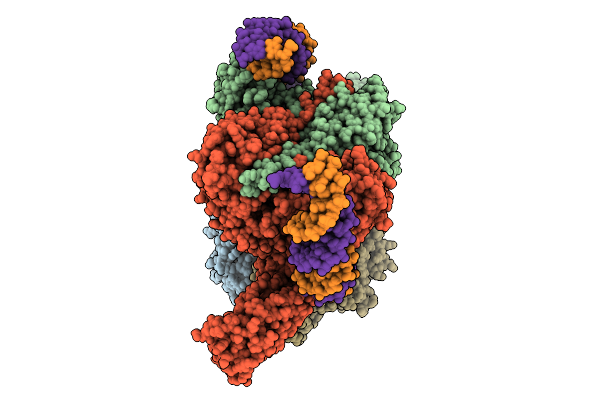 |
Cryoem Structure Of M. Mazei Topoisomerase Vi(A-E342Q)-Minicircle Dna Complex In Asymmetric State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, CA, K |
 |
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, MG, K |
 |
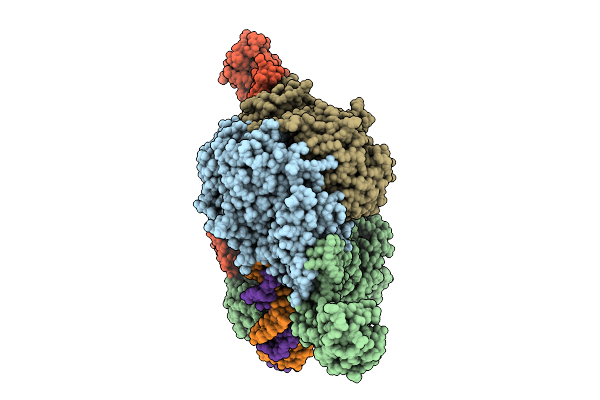
Cryoem Structure Of M. Mazei Topoisomerase Vi-Minicircle Dna Complex In Partially Unfolded Transducer State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: MG, ANP, K |
 |
Cryoem Structure Of M. Mazei Topoisomerase Vi-Minicircle Dna Complex In Asymmetric State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, MG, K |
 |
Organism: Caenorhabditis elegans
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN |
 |
Organism: Caenorhabditis elegans
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN |
 |
Cryo-Em Structure Of The Major Capsid Protein Vp39 Of Autographa Californica Multiple Nucleopolyhedrovirus (Acmnpv)
Organism: Autographa californica multiple nucleopolyhedrovirus
Method: ELECTRON MICROSCOPY Release Date: 2023-12-13 Classification: VIRAL PROTEIN |
 |
Outer Shell And Inner Layer Structures Of Autographa Californica Multiple Nucleopolyhedrovirus (Acmnpv)
Organism: Autographa californica multiple nucleopolyhedrovirus
Method: ELECTRON MICROSCOPY Release Date: 2023-12-13 Classification: VIRAL PROTEIN |
 |
Plug Structure Of The Autographa Californica Multiple Nucleopolyhedrovirus (Acmnpv)
Organism: Autographa californica multiple nucleopolyhedrovirus
Method: ELECTRON MICROSCOPY Release Date: 2023-12-13 Classification: VIRAL PROTEIN |
 |
Iii2Iv2 Respiratory Supercomplex From Saccharomyces Cerevisiae With 4 Bound Uq6
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2023-05-24 Classification: OXIDOREDUCTASE Ligands: PEF, FES, CDL, PCF, CN5, HEM, UQ6, HEA, CU, CUA |
 |
Iii2Iv Respiratory Supercomplex From Saccharomyces Cerevisiae Cardiolipin-Lacking Mutant
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2023-05-24 Classification: OXIDOREDUCTASE Ligands: PEF, PGT, FES, PCF, HEM, UQ6, CU, HEA, CUA |
 |
Organism: Scylla serrata reovirus sz-2007
Method: ELECTRON MICROSCOPY Release Date: 2023-04-19 Classification: VIRUS |
 |
Organism: Scylla serrata reovirus sz-2007
Method: ELECTRON MICROSCOPY Release Date: 2023-04-19 Classification: VIRUS |

